Electrode Layouts
EegFun.jl includes a comprehensive collection of electrode layout files that define the spatial positions of EEG channels for various montage systems. These layouts are essential for topographic plotting, channel interpolation, and spatial analyses.
Overview
Layout files are stored in CSV format with three required columns:
label: Channel name (Symbol)inc: Incidence angle in degrees (vertical angle from vertex)azi: Azimuth angle in degrees (horizontal angle)
The polar coordinates (inc, azi) serve as the Source of Truth for electrode positions. Cartesian coordinates (2D and 3D) are computed on-demand from these polar coordinates.
Sources
The electrode layout files included in EegFun.jl were obtained from the following sources:
BioSemi: Downloaded from BioSemi Head Cap Specifications
BrainProducts actiCap: Downloaded from BrainProducts Cap Montages
EasyCap: Generated with reference to:
FieldTrip template layouts (
fieldtrip/template/layout/)MNE-Python channel montages (
mne-python/mne/channels/data/montages)EasyCap electrode layout documentation (PDF) - Converted to polar coordinates (label, inc, azi) for consistency with BioSemi format
Usage
# Load a layout file
layout = EegFun.read_layout("./resources/layouts/biosemi/biosemi72.csv")
# Convert to 2D Cartesian coordinates (preserving radial distances)
EegFun.polar_to_cartesian_xy!(layout)
# Calculate neighbors
EegFun.get_neighbours_xy!(layout, 0.3)
# Plot the layout with neighbours; hover over electrodes to see neighbours
EegFun.plot_layout_2d(layout; neighbours=true)
EegFun.get_neighbours_xyz!(layout, 0.5)
EegFun.plot_layout_3d(layout; neighbours=true)Available Layouts
BioSemi
Standard BioSemi ActiveTwo electrode caps and extended montages:
biosemi16 (16 channels) - Standard 16-channel montage
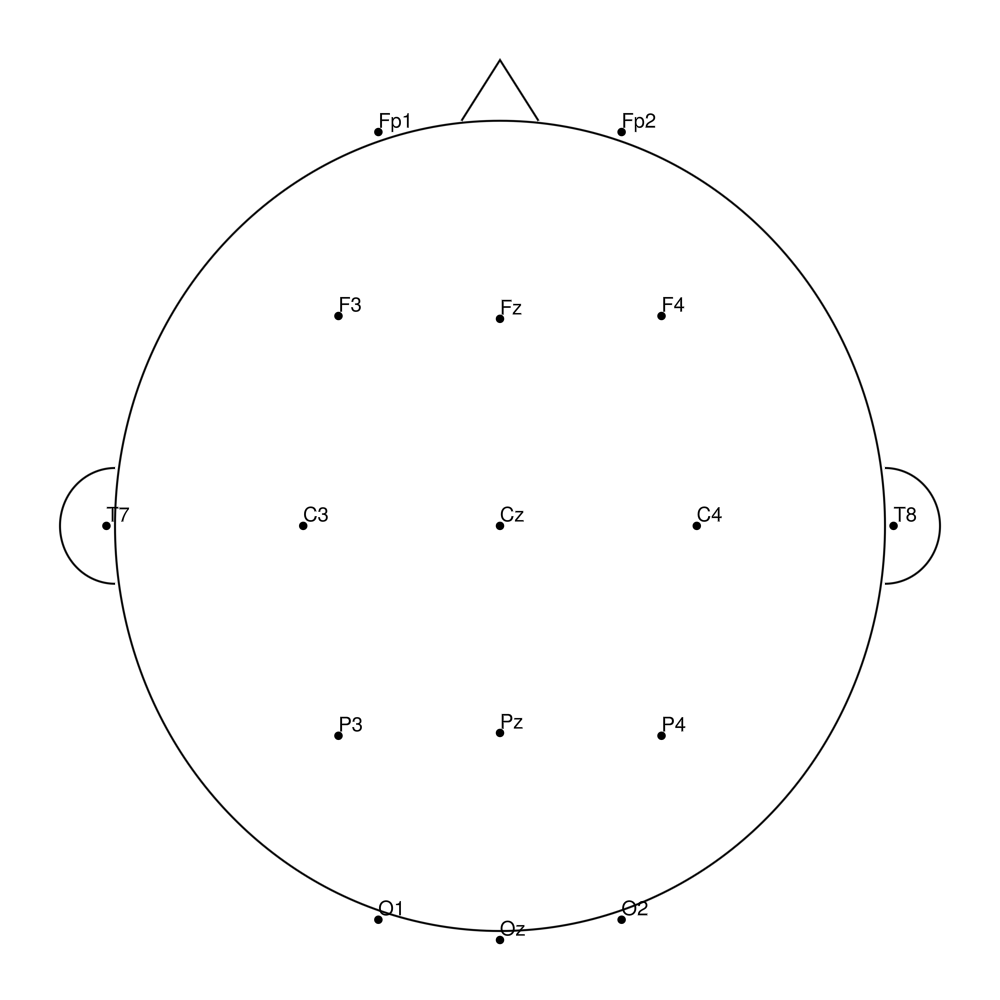
biosemi32 (32 channels) - Standard 32-channel montage
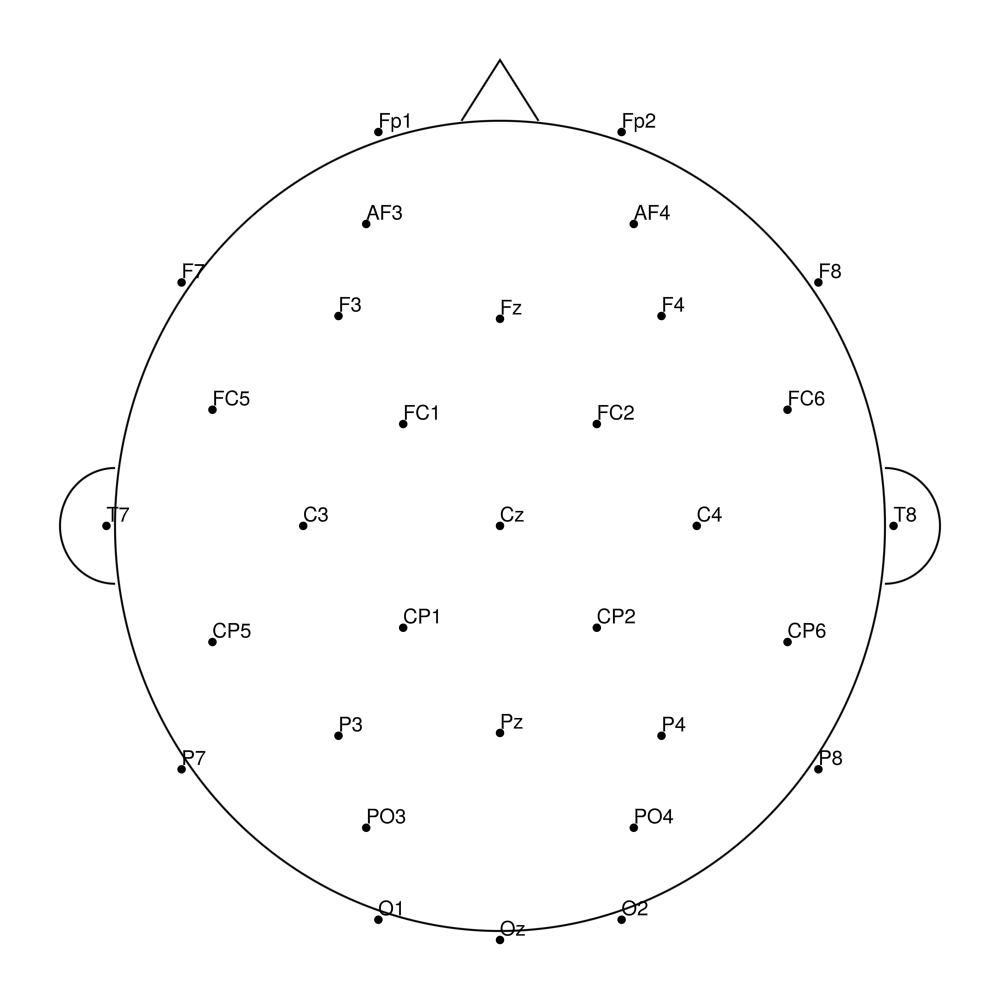
biosemi64 (64 channels) - Standard 64-channel montage
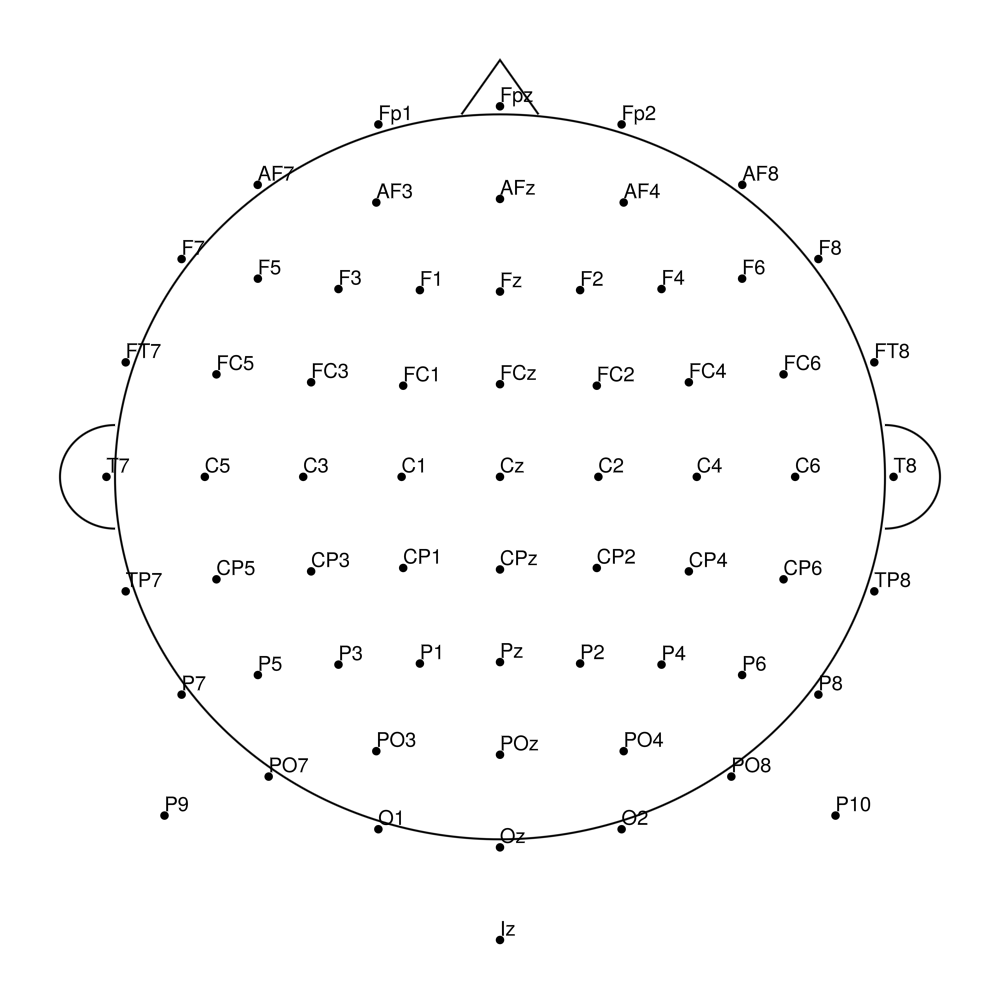
biosemi70 (70 channels) - 64 + 6 external channels
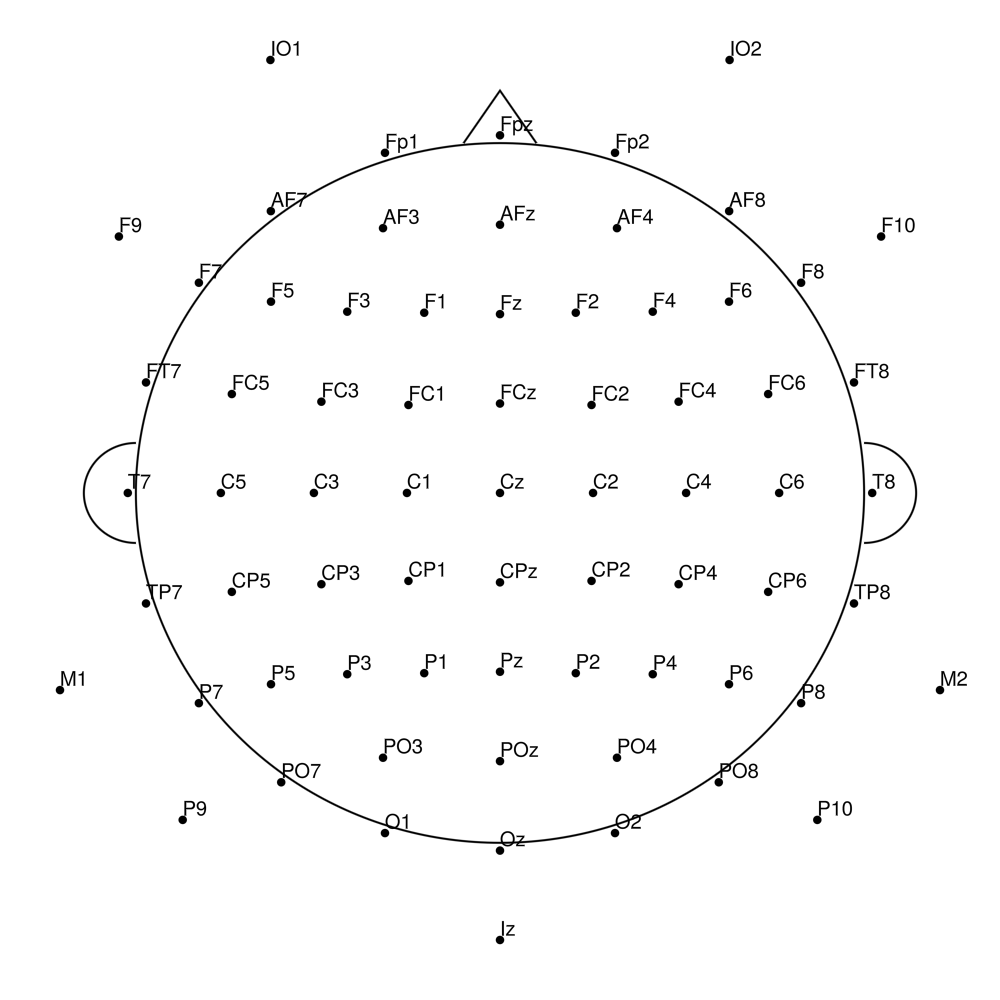
biosemi72 (72 channels) - 64 + 8 external channels
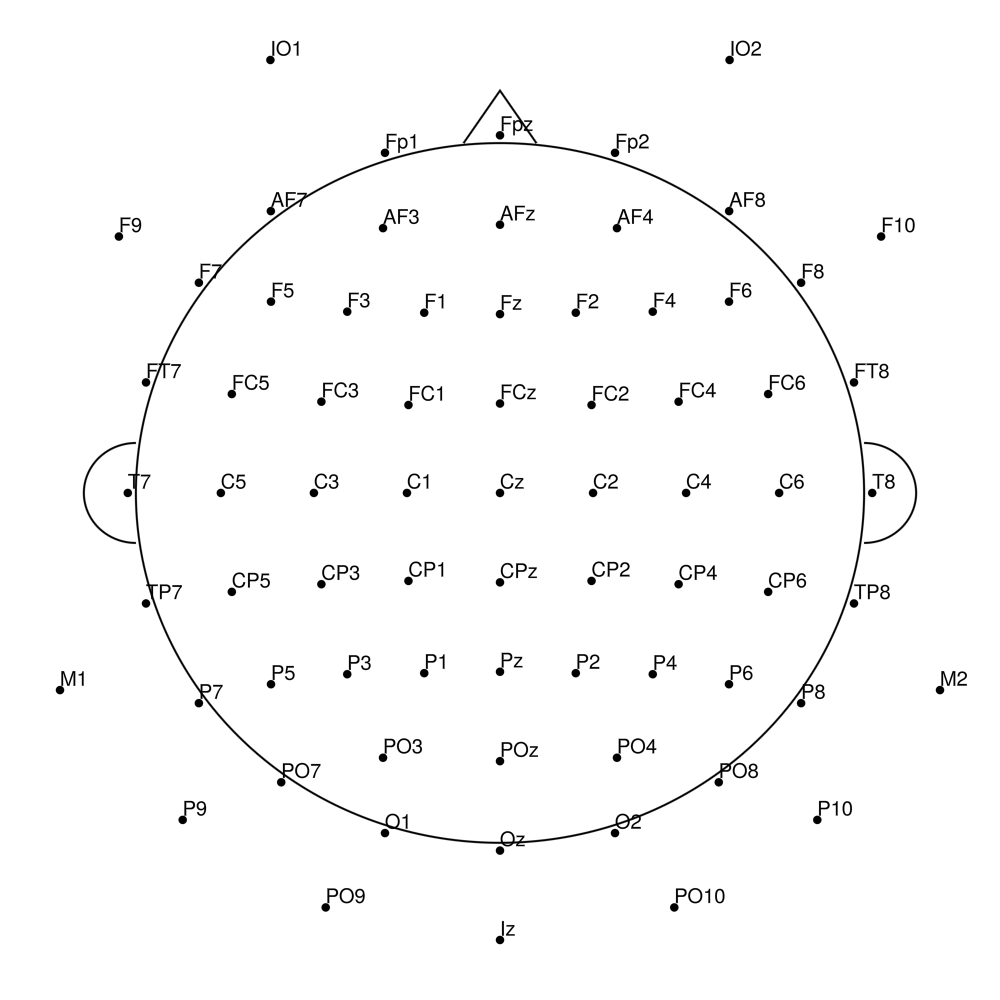
biosemi128 (128 channels) - Standard 128-channel montage
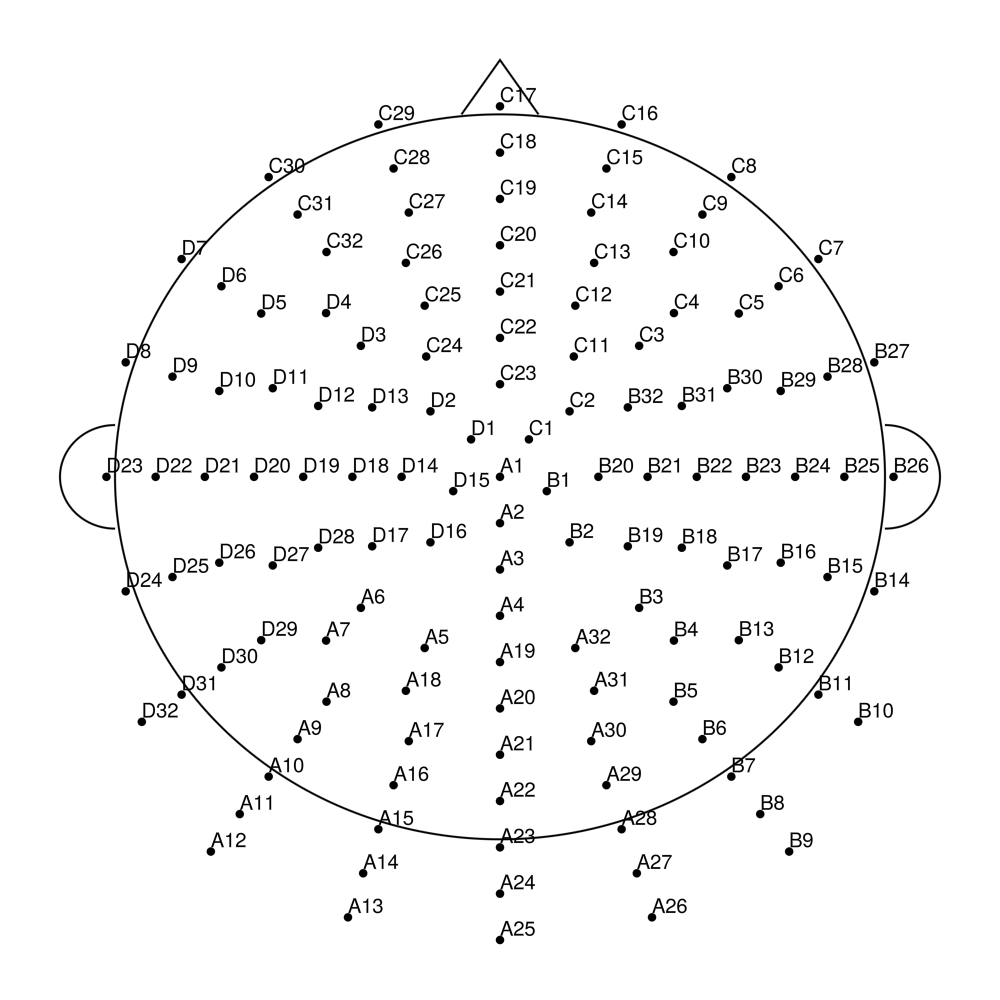
biosemi160 (160 channels) - Extended high-density montage
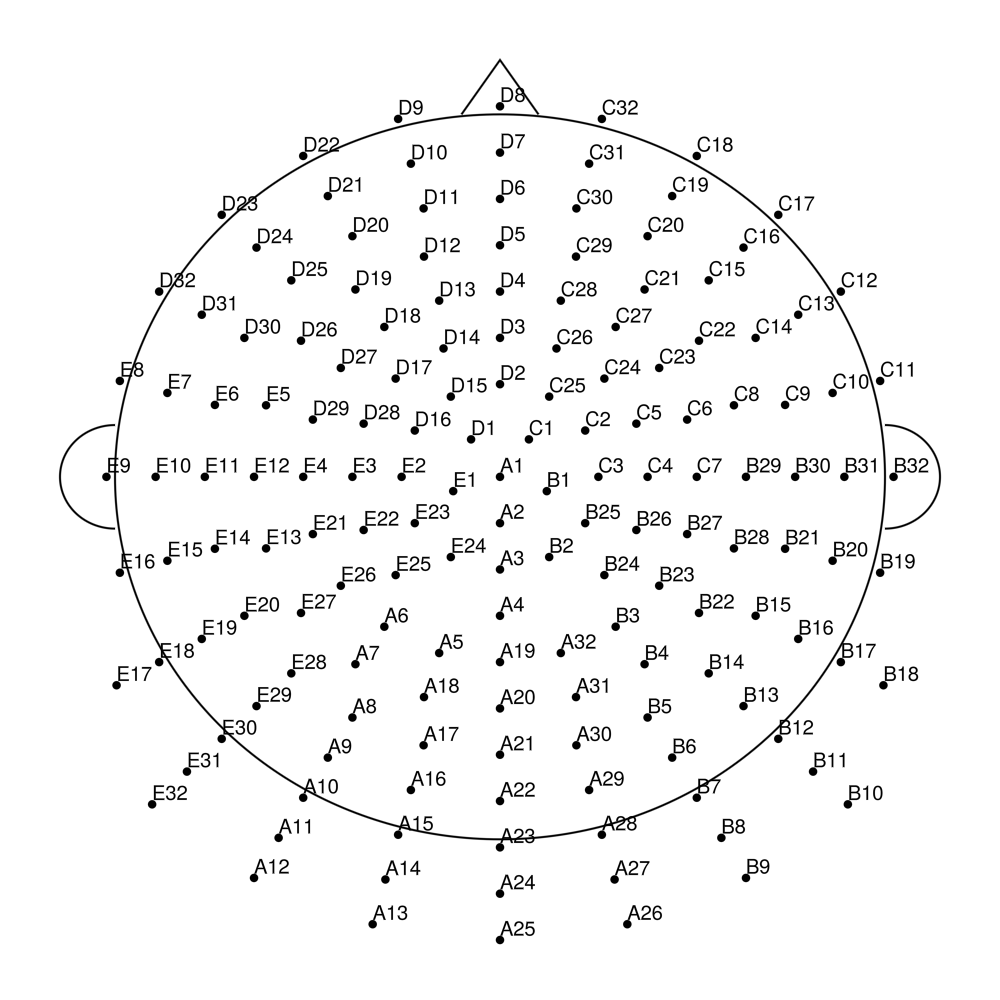
biosemi256 (256 channels) - Ultra high-density montage
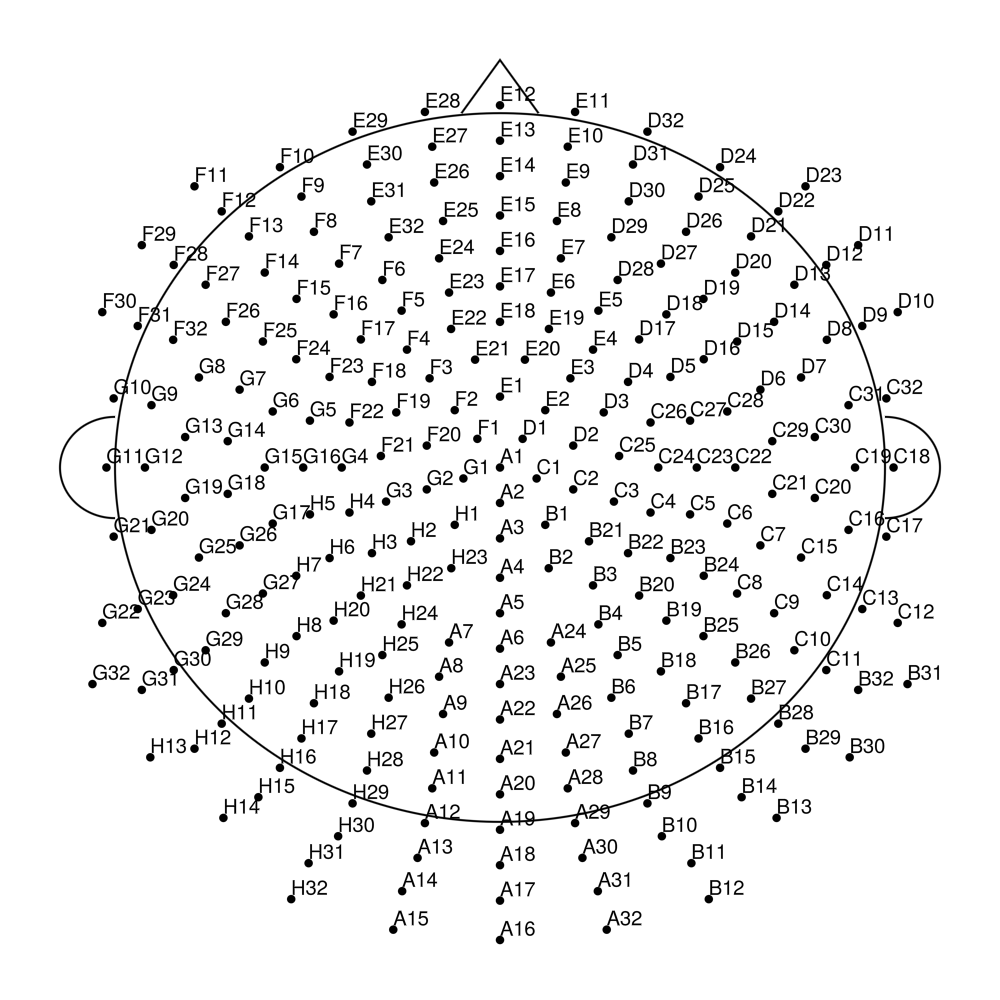
BrainProducts actiCap
BrainProducts electrode caps organized by product line:
Standard BrainCap
BC-22 (22 channels) - 22-channel clinical montage
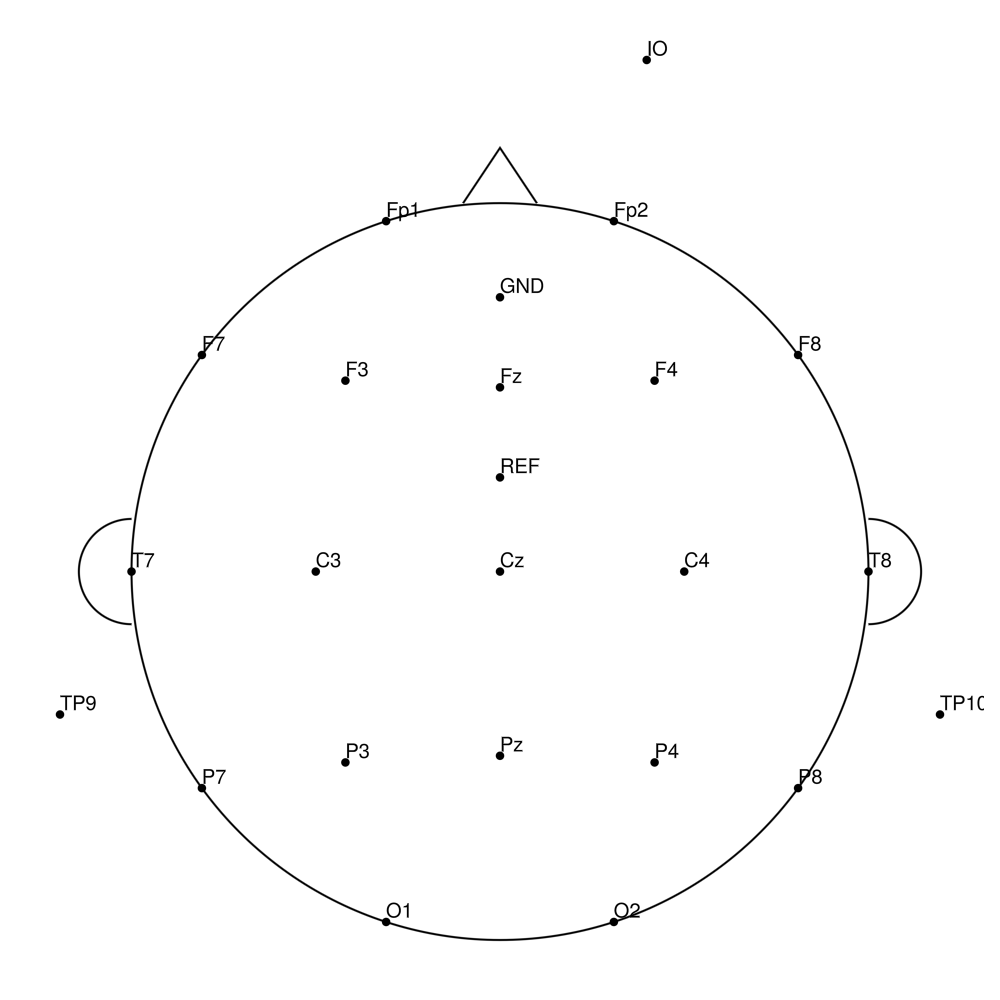
BC-32 (32 channels) - Standard 32-channel montage
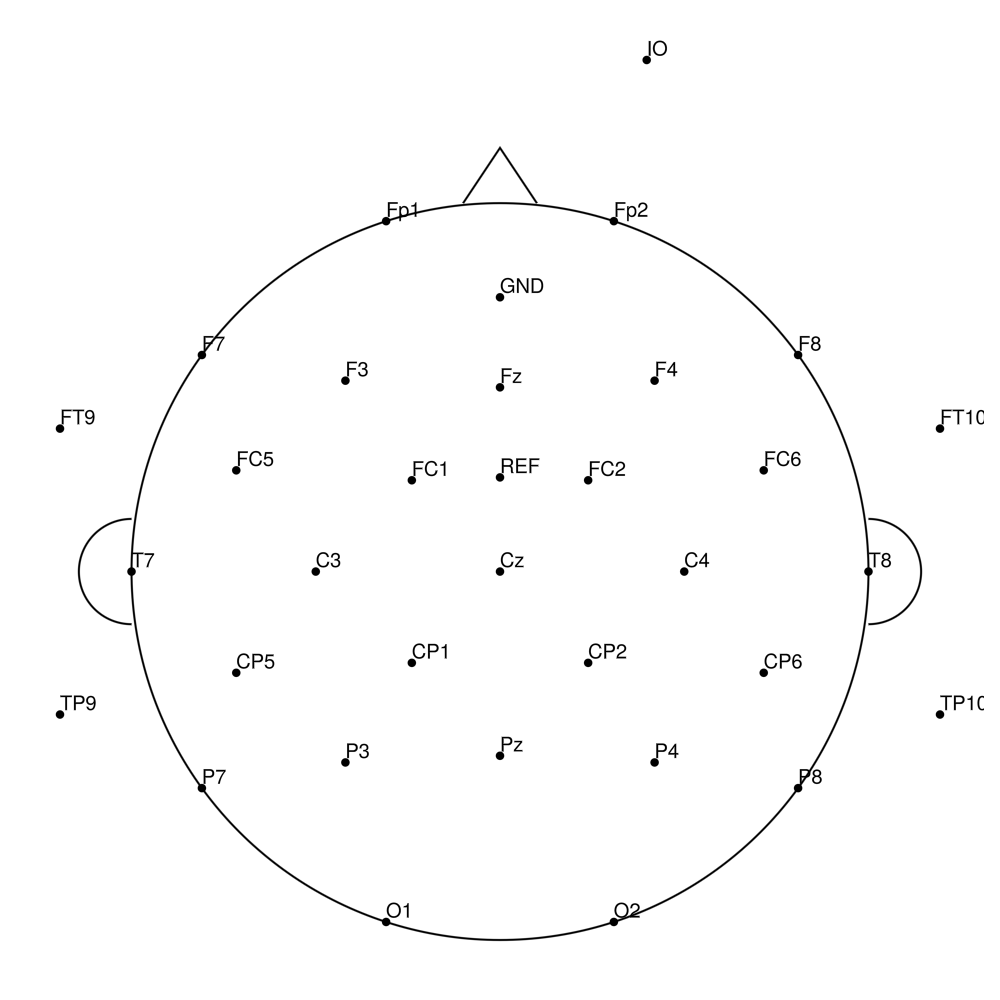
BC-64 (64 channels) - Standard 64-channel montage
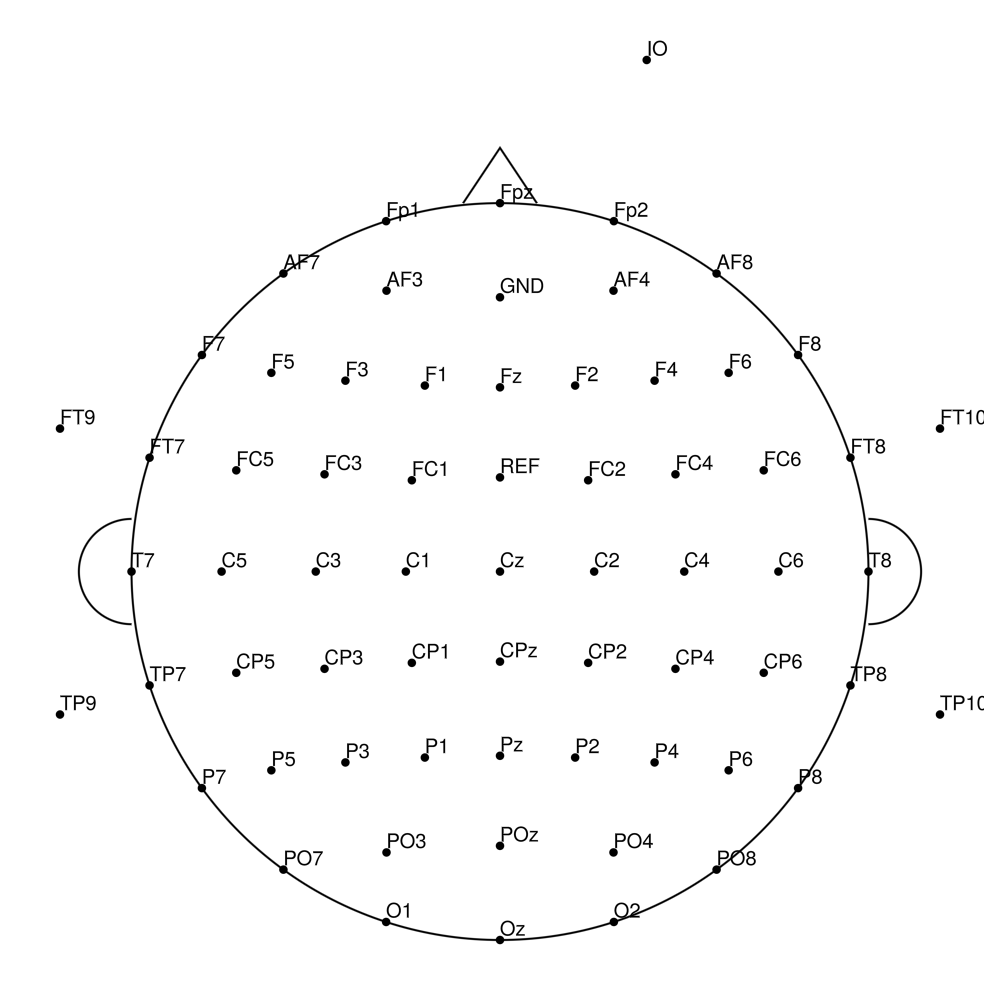
BC-96 (96 channels) - High-density 96-channel montage
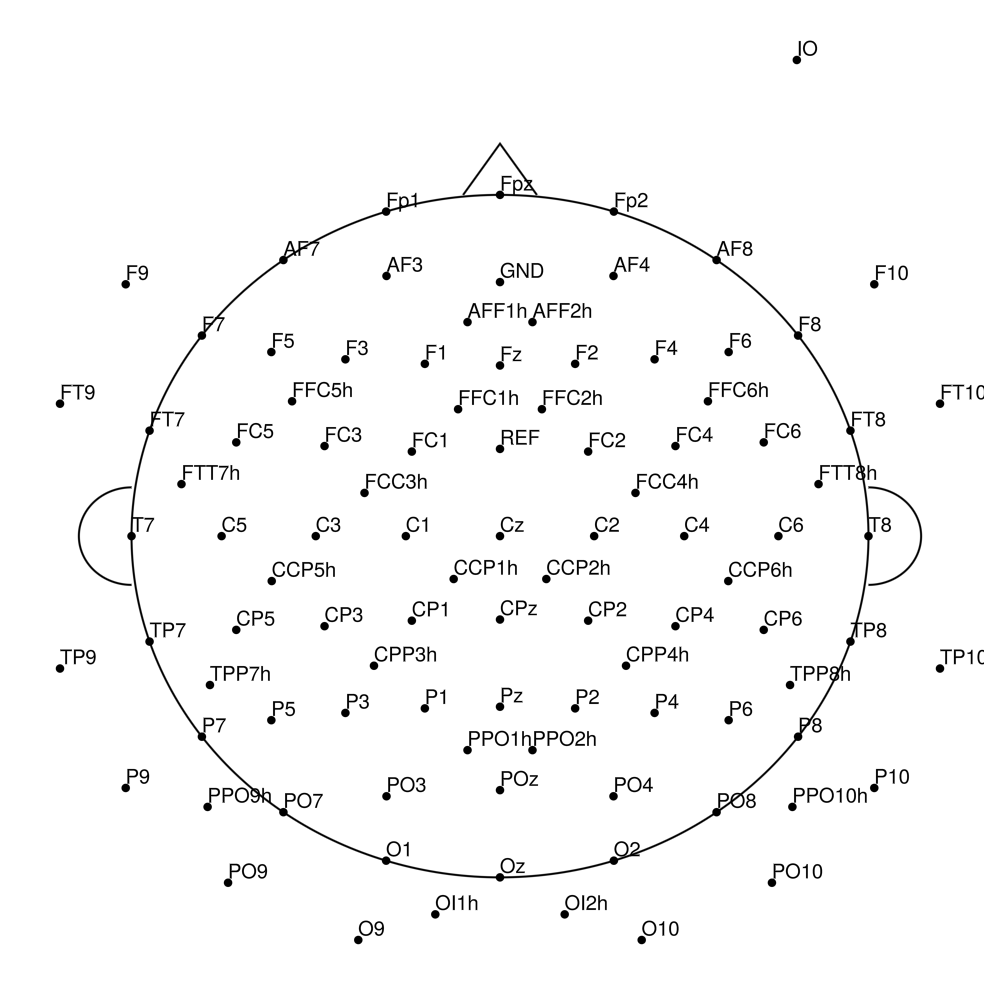
BC-128 (128 channels) - High-density 128-channel montage
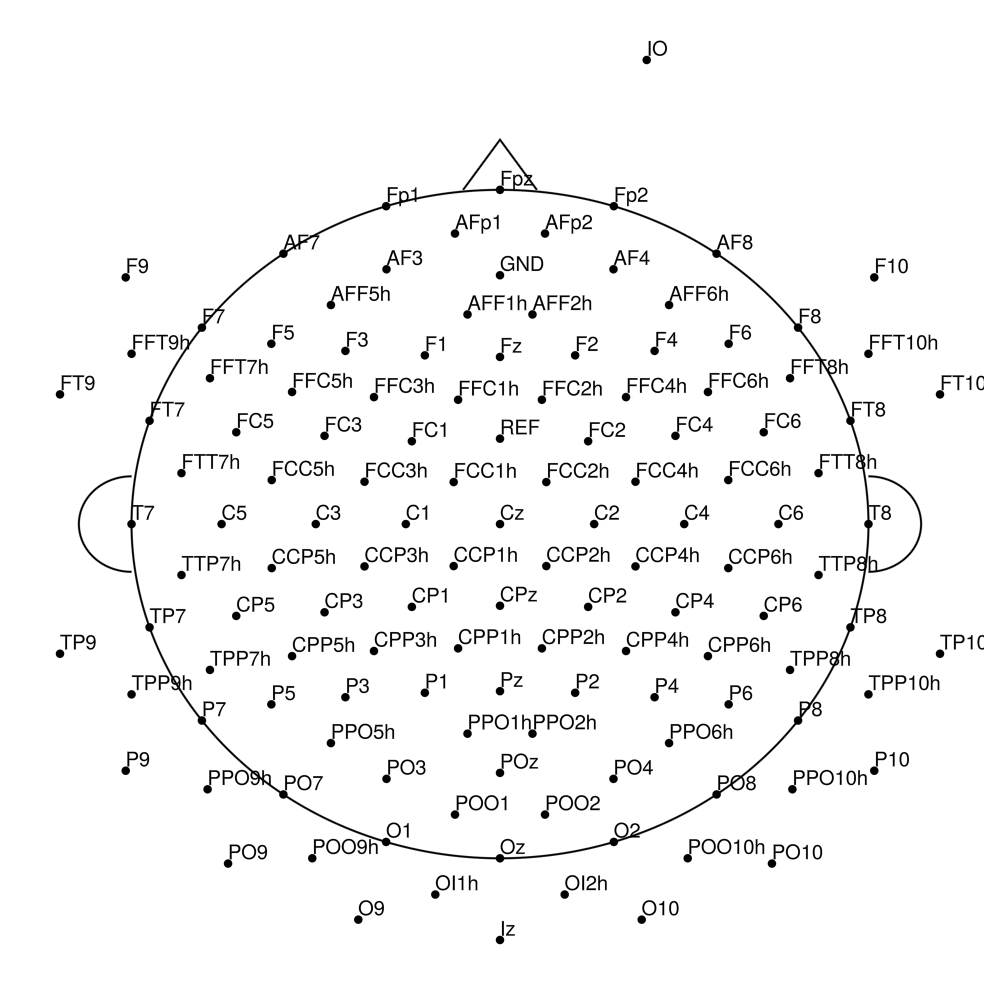
BC-256 (256 channels) - Ultra high-density montage
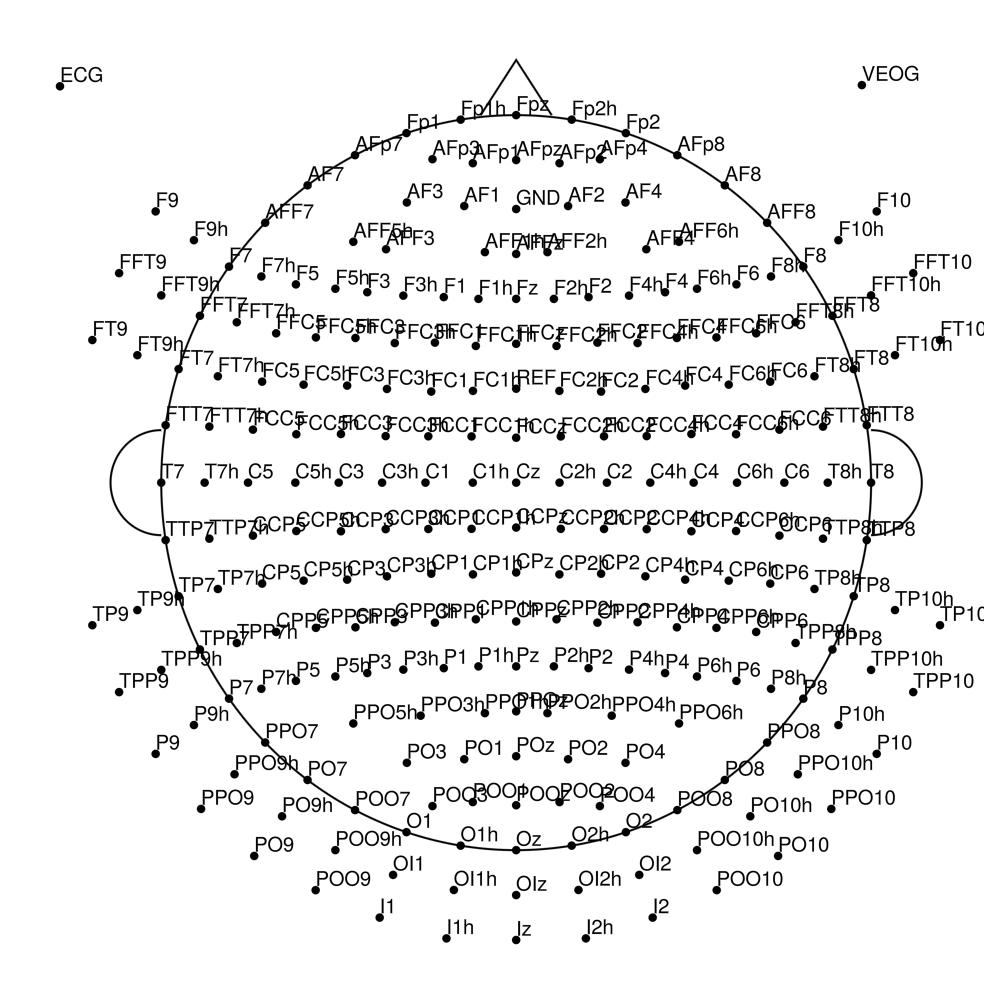
BrainCap MR (MRI-compatible)
BC-MR-32 (32 channels)
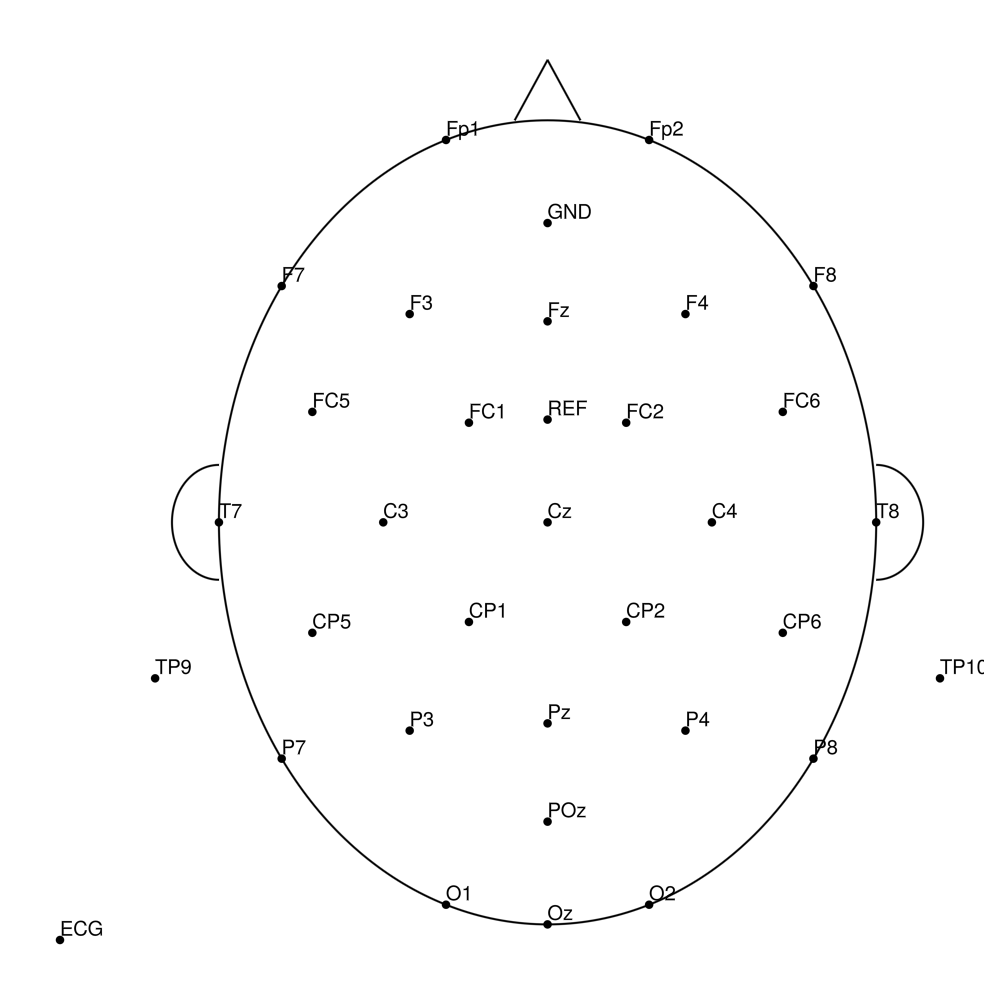
BC-MR-64 (64 channels)
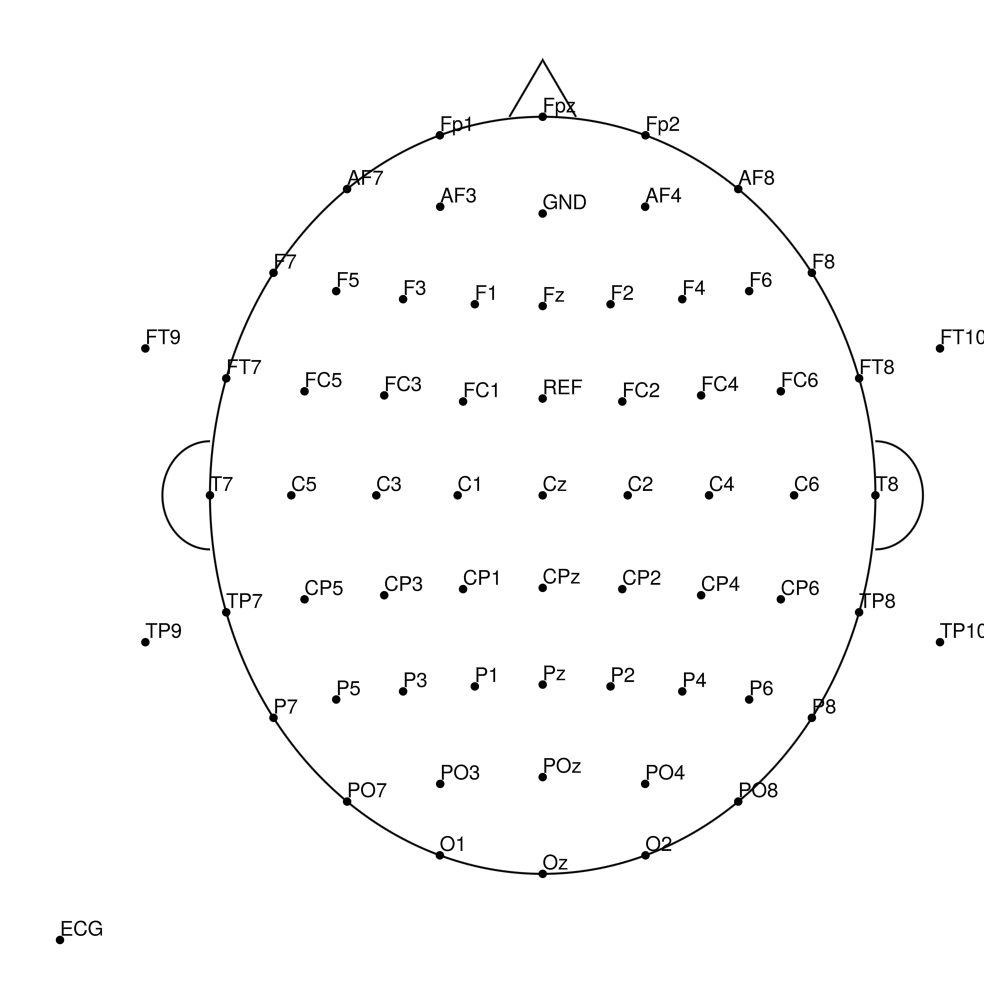
BC-MR-96 (96 channels)
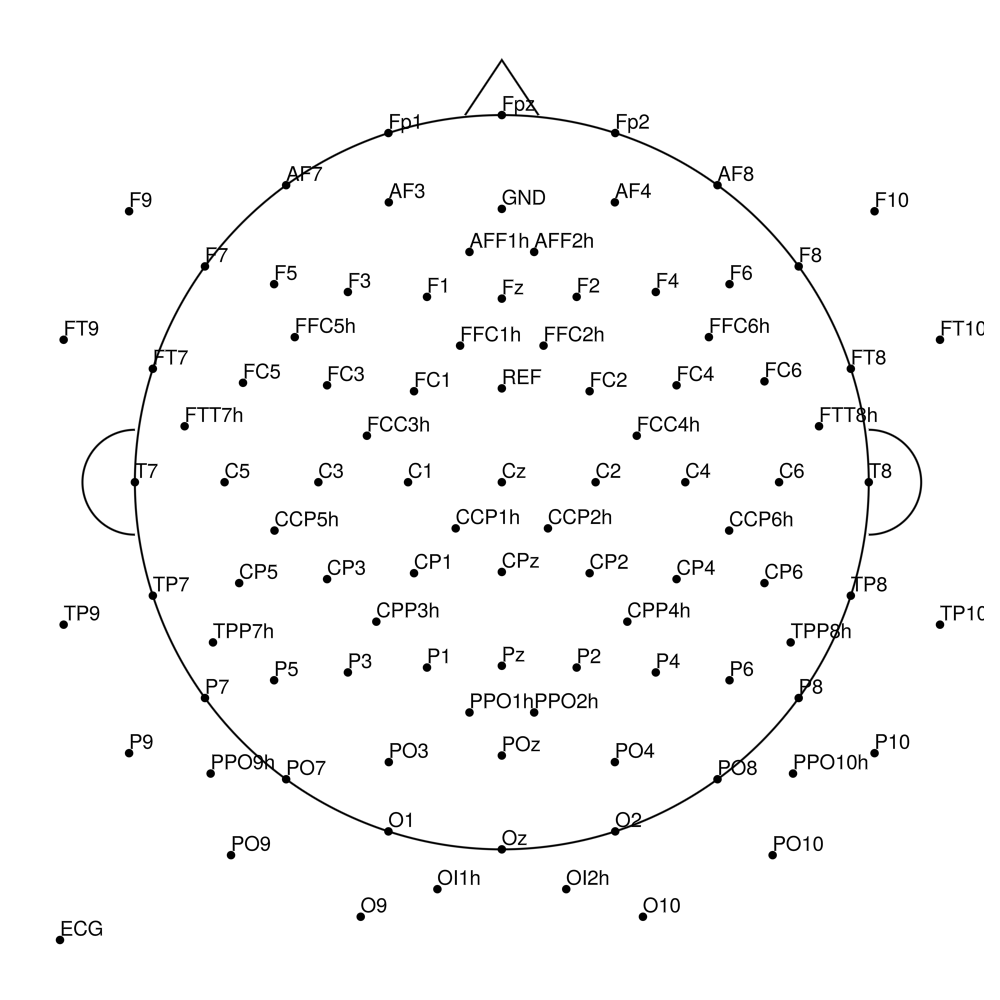
BC-MR-128 (128 channels)
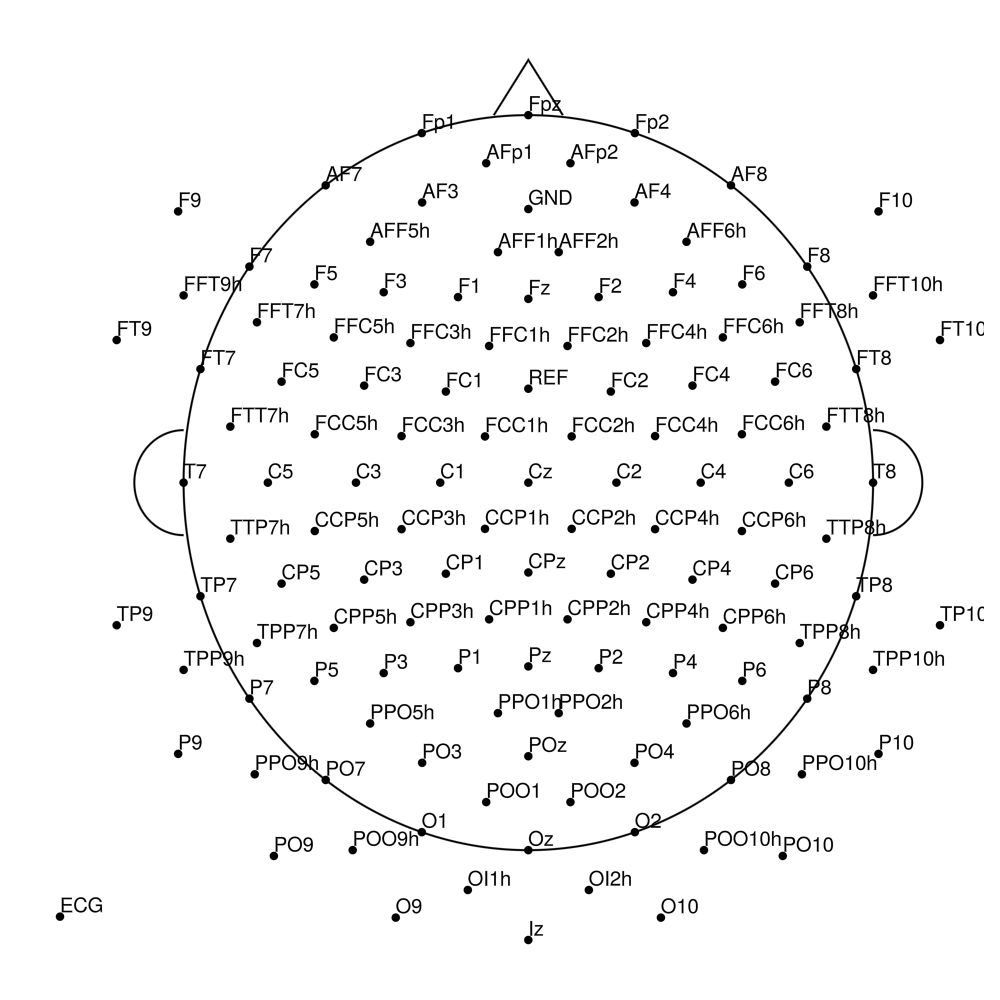
BrainCap MR3 (3T MRI-compatible)
BC-MR3-32 (32 channels)
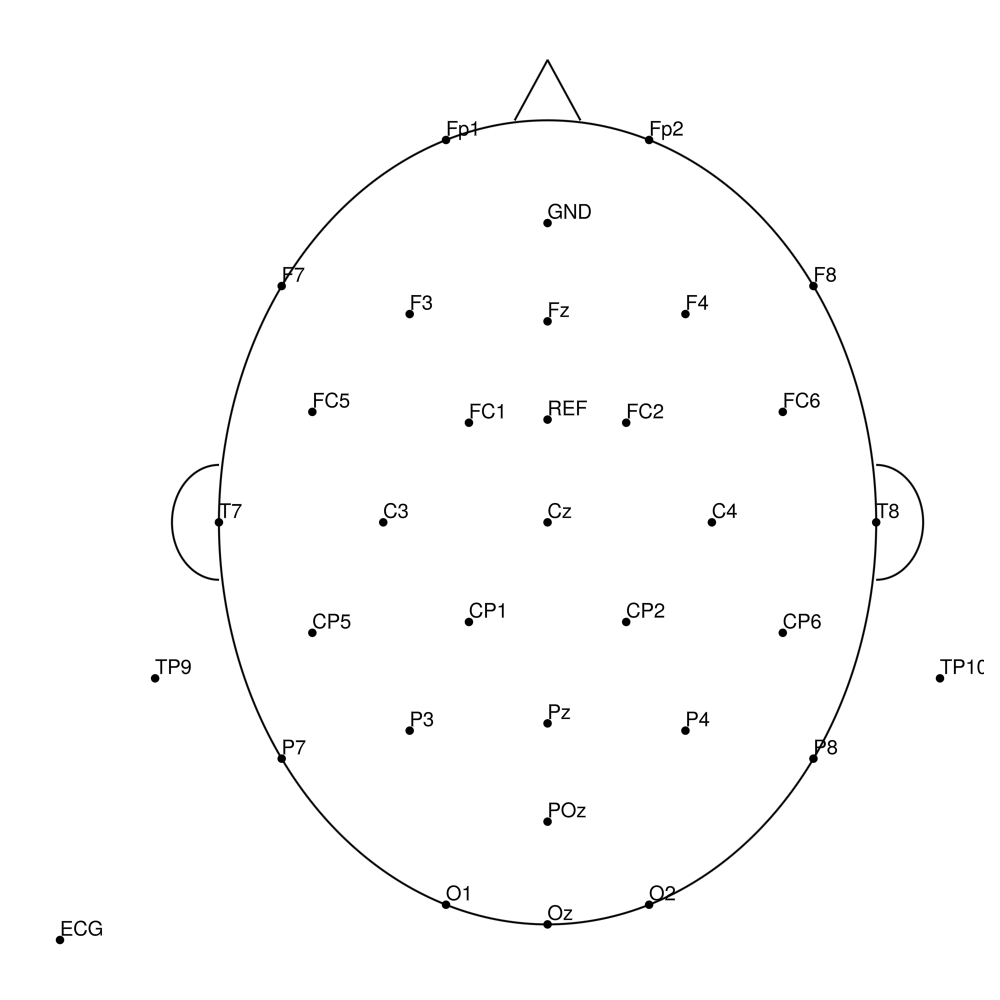
BC-MR3-64 (64 channels)
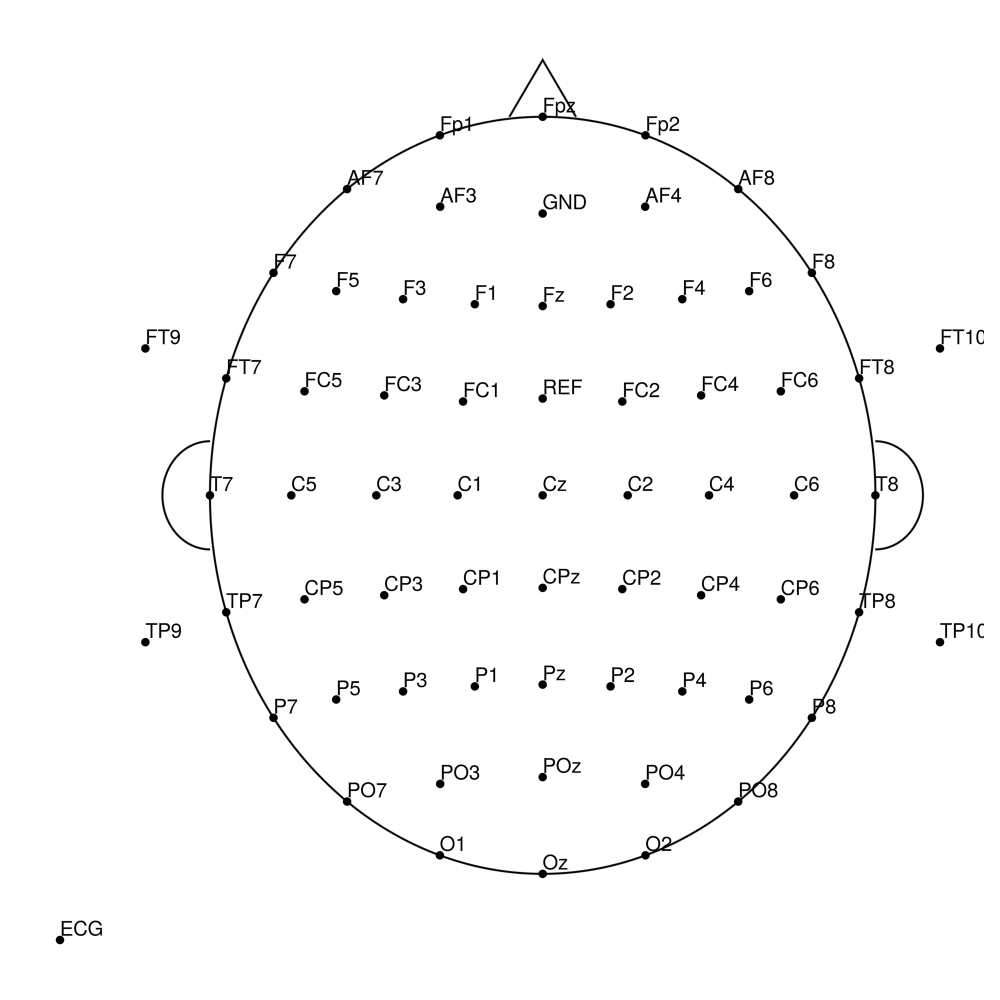
BC-MR3-96 (96 channels)
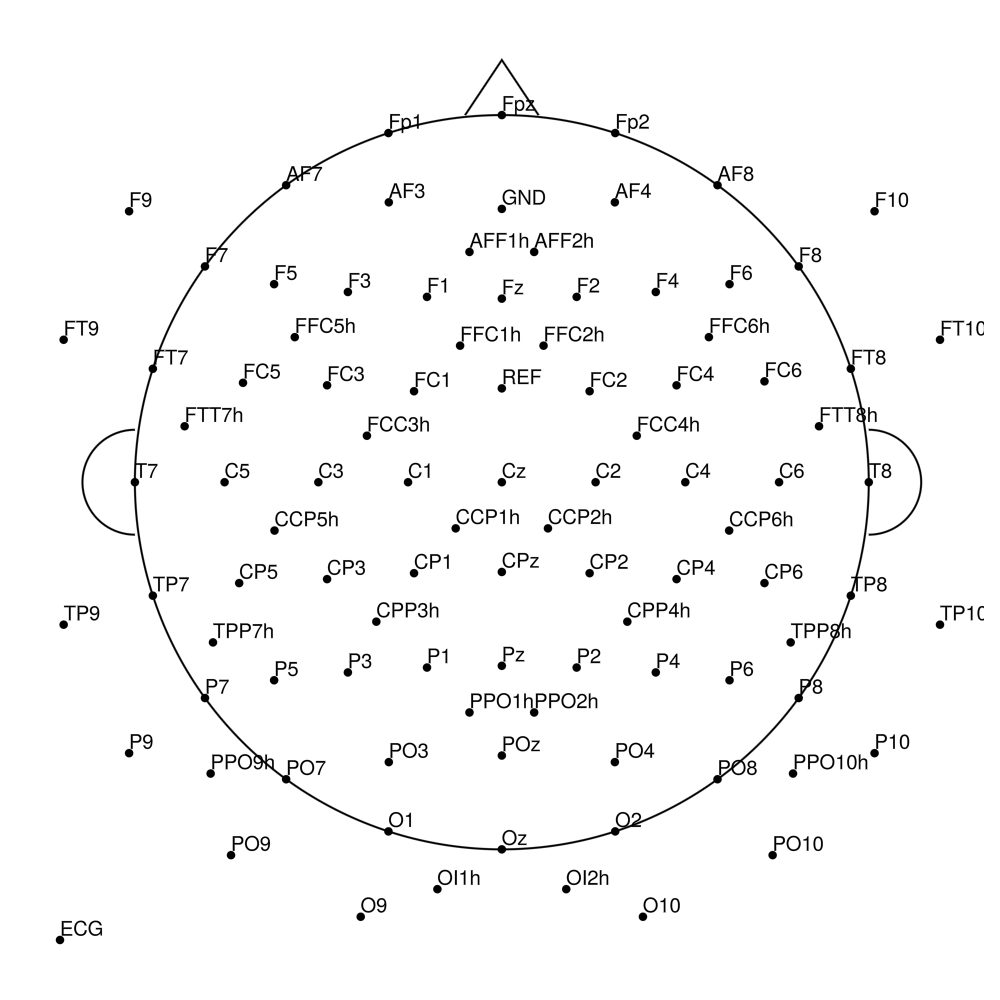
BC-MR3-128 (128 channels)
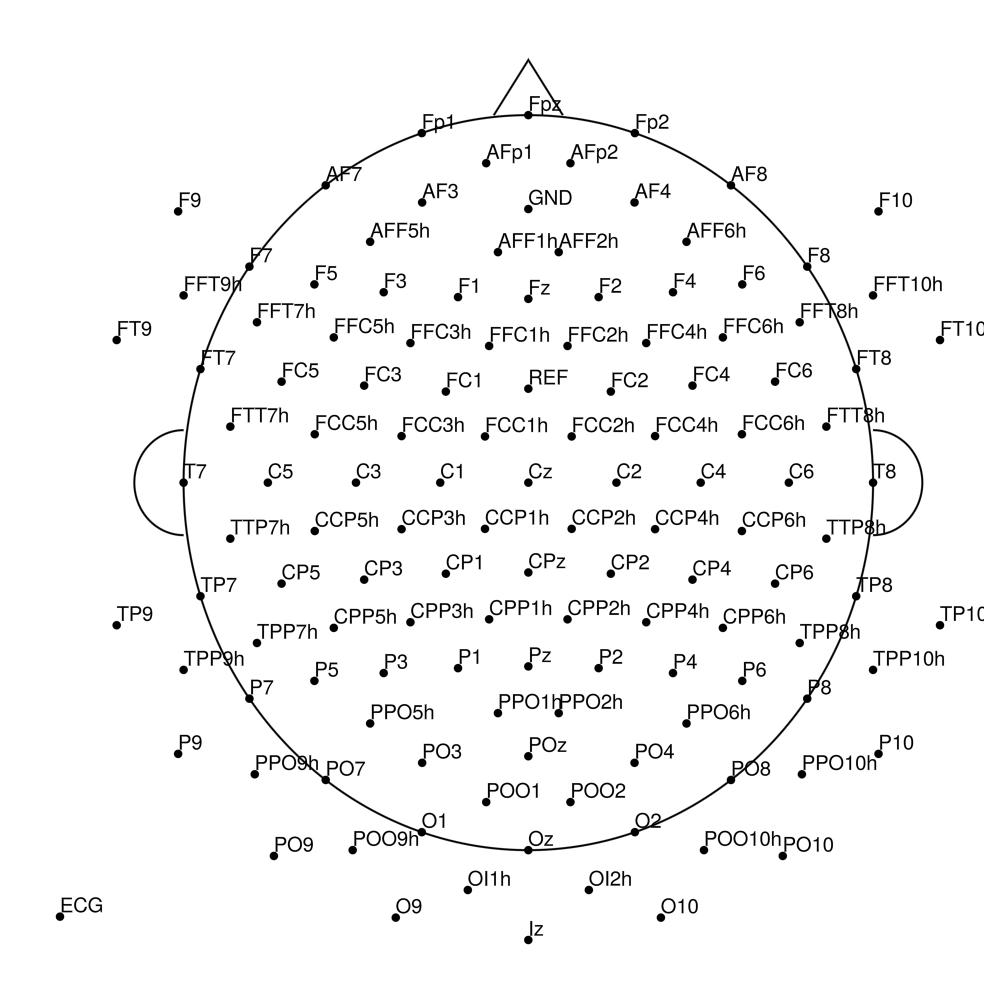
BrainCap MEG (MEG-compatible)
BC-MEG-32 (32 channels)
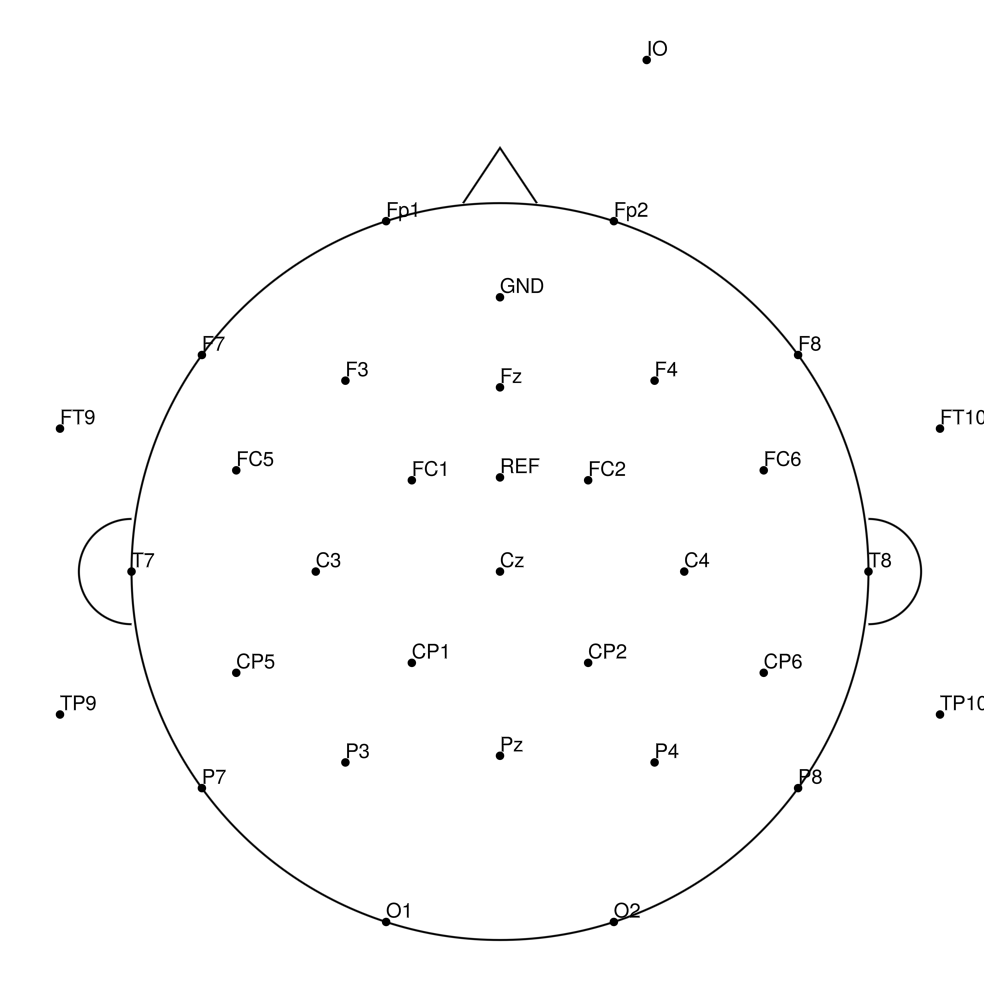
BC-MEG-64 (64 channels)
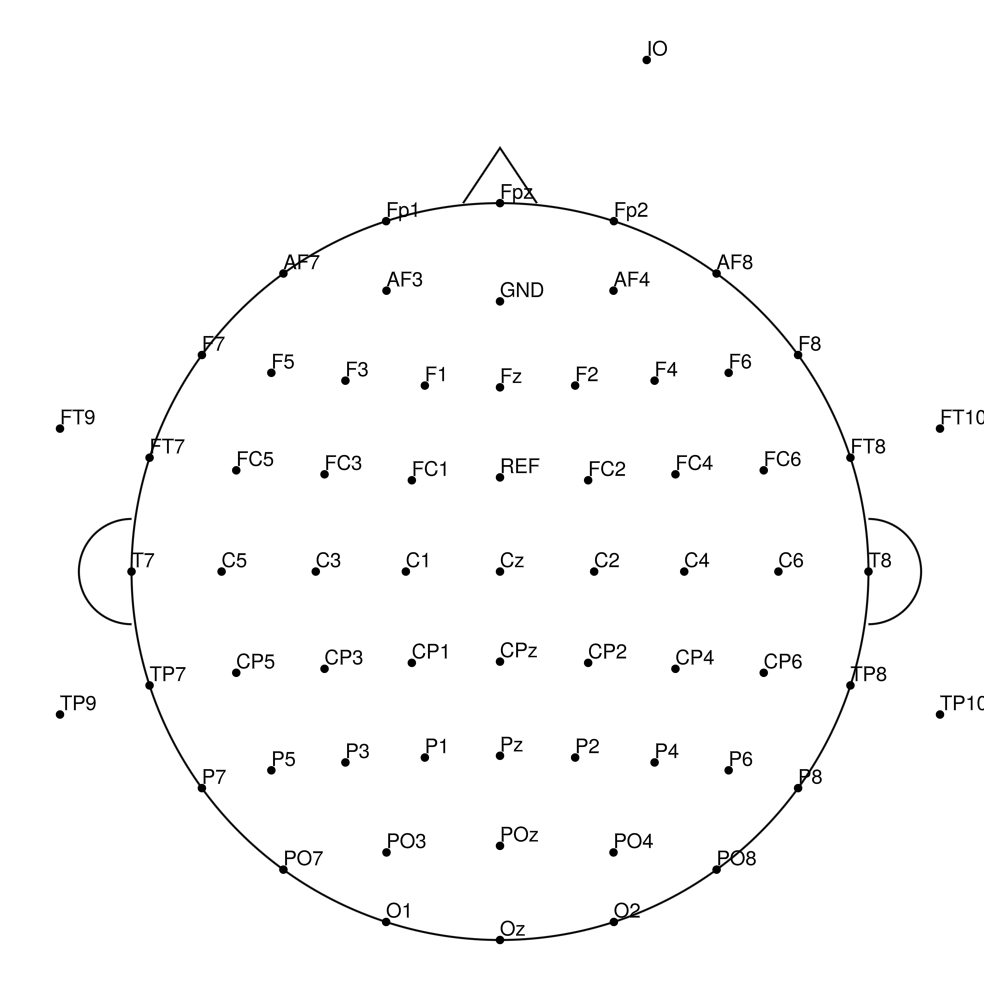
BC-MEG-128 (128 channels)
BrainCap TMS (TMS-compatible)
BC-TMS-32 (32 channels)
BC-TMS-64 (64 channels)
BC-TMS-96 (96 channels)
BC-TMS-128 (128 channels)
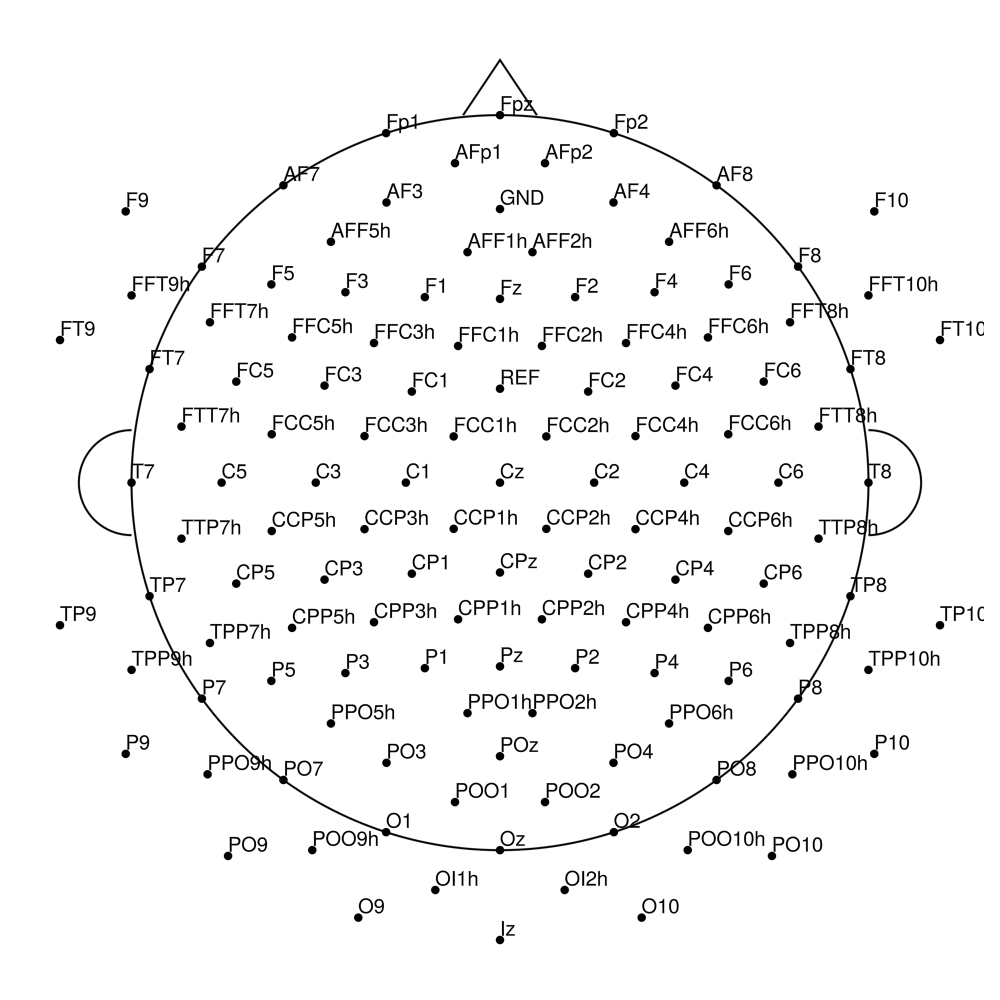
BrainCap Sleep
BC-MRS-15 (15 channels) - MR sleep - AASM standard
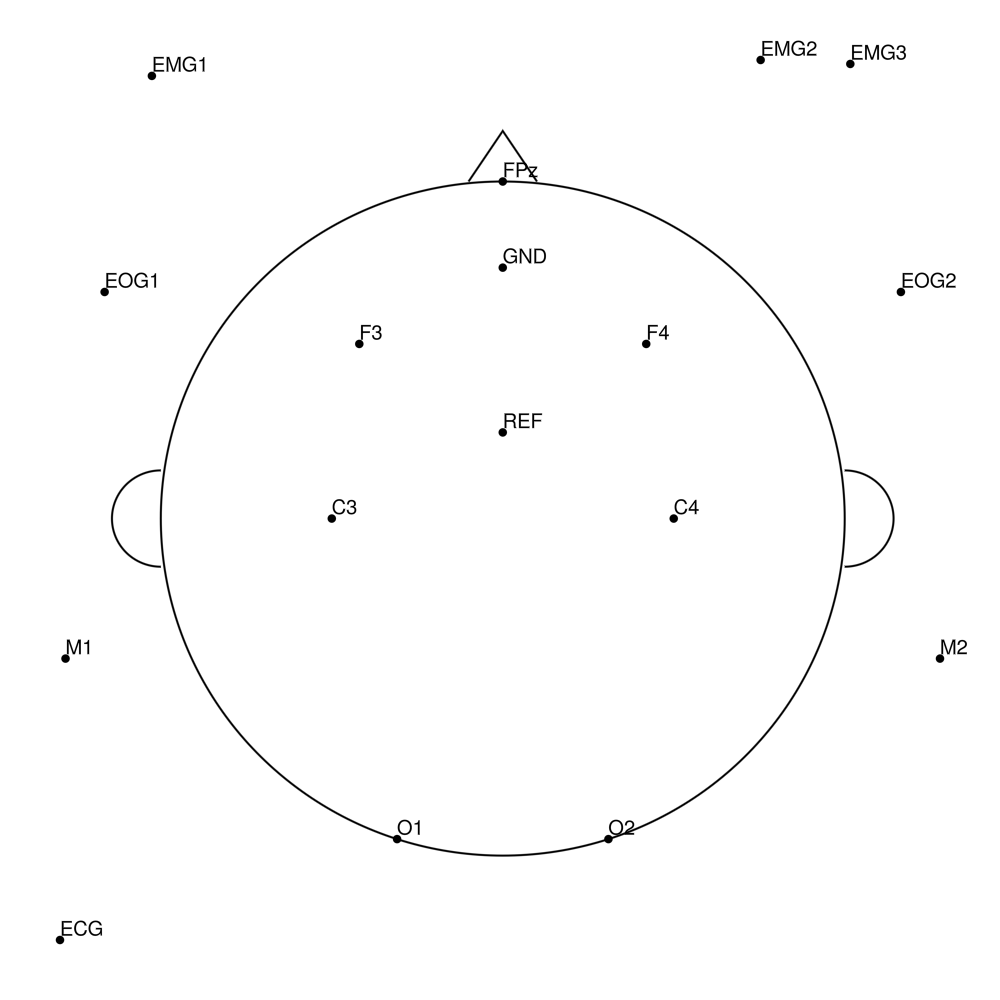
BC-MRS-32 (32 channels) - MR sleep montage
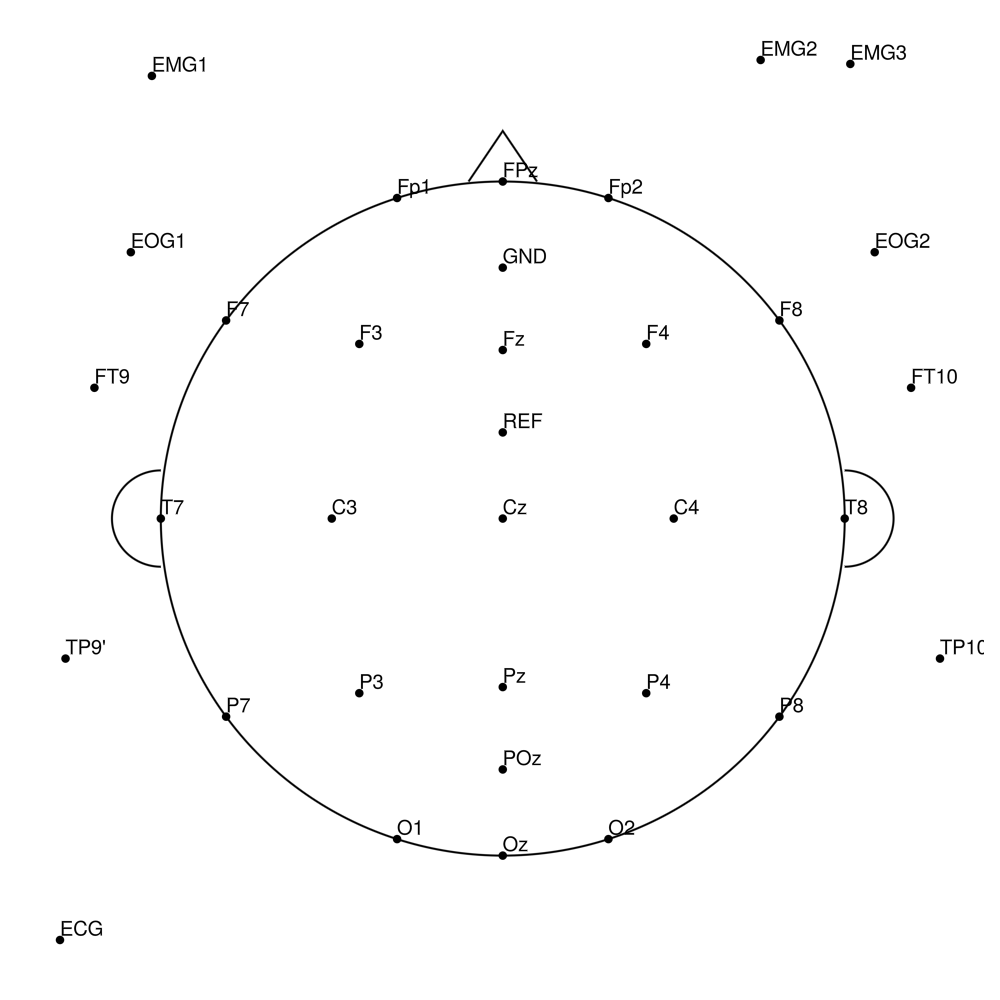
BC-MRS-64 (64 channels) - MR sleep montage
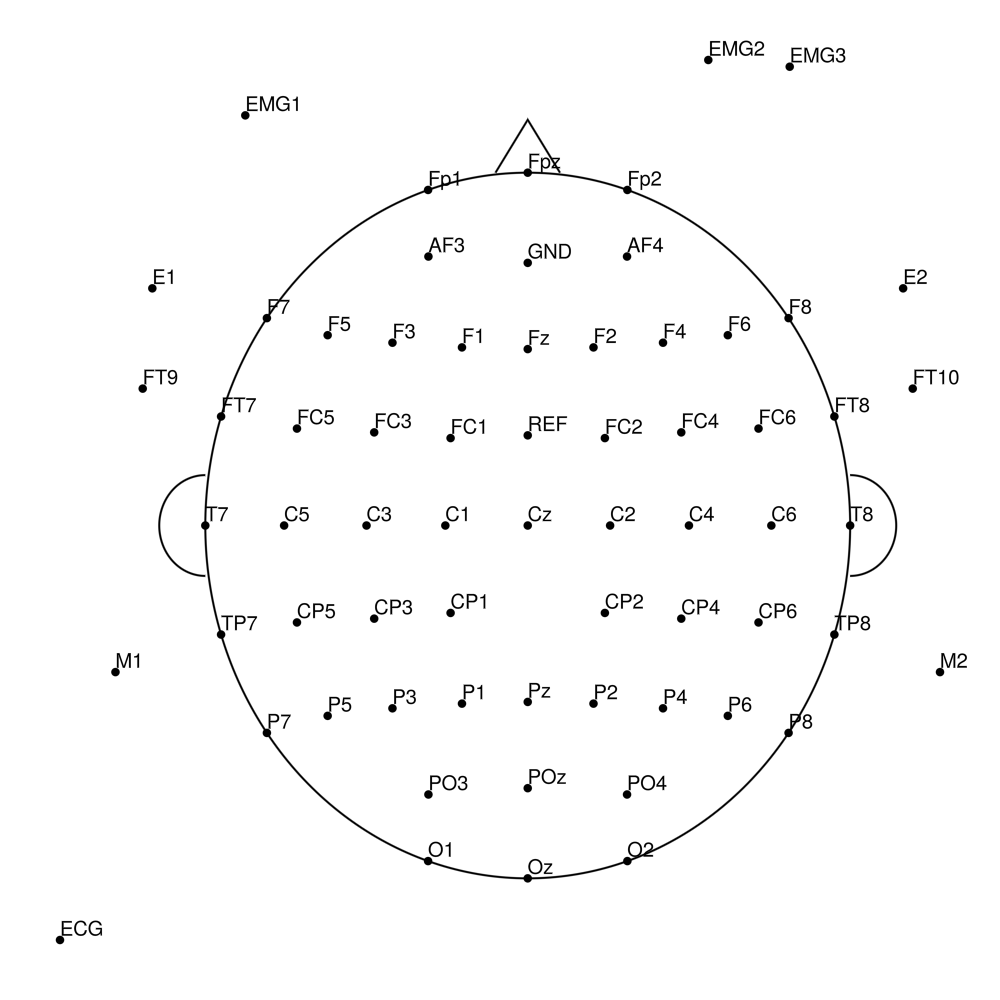
BC-MRS3-15 (15 channels) - MR3 sleep - AASM standard
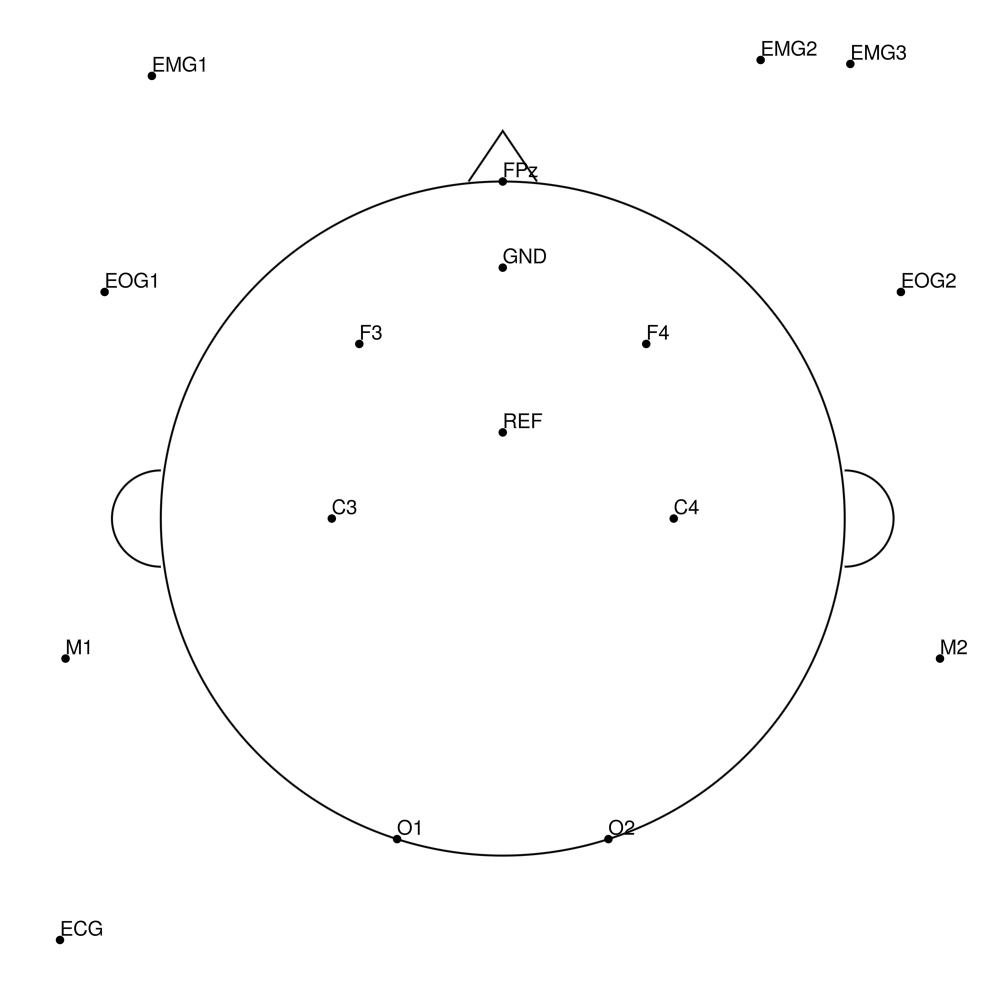
BC-MRS3-32 (32 channels) - MR3 sleep montage
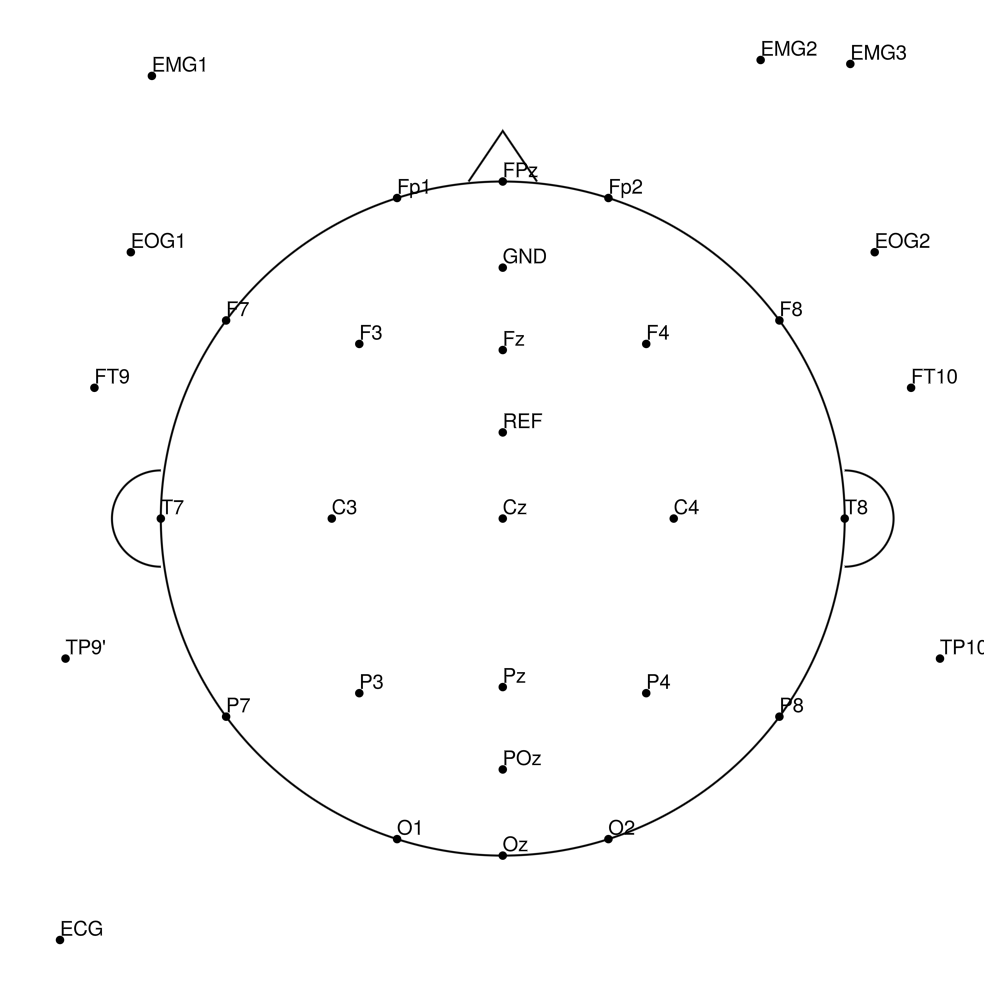
BC-MRS3-64 (64 channels) - MR3 sleep montage
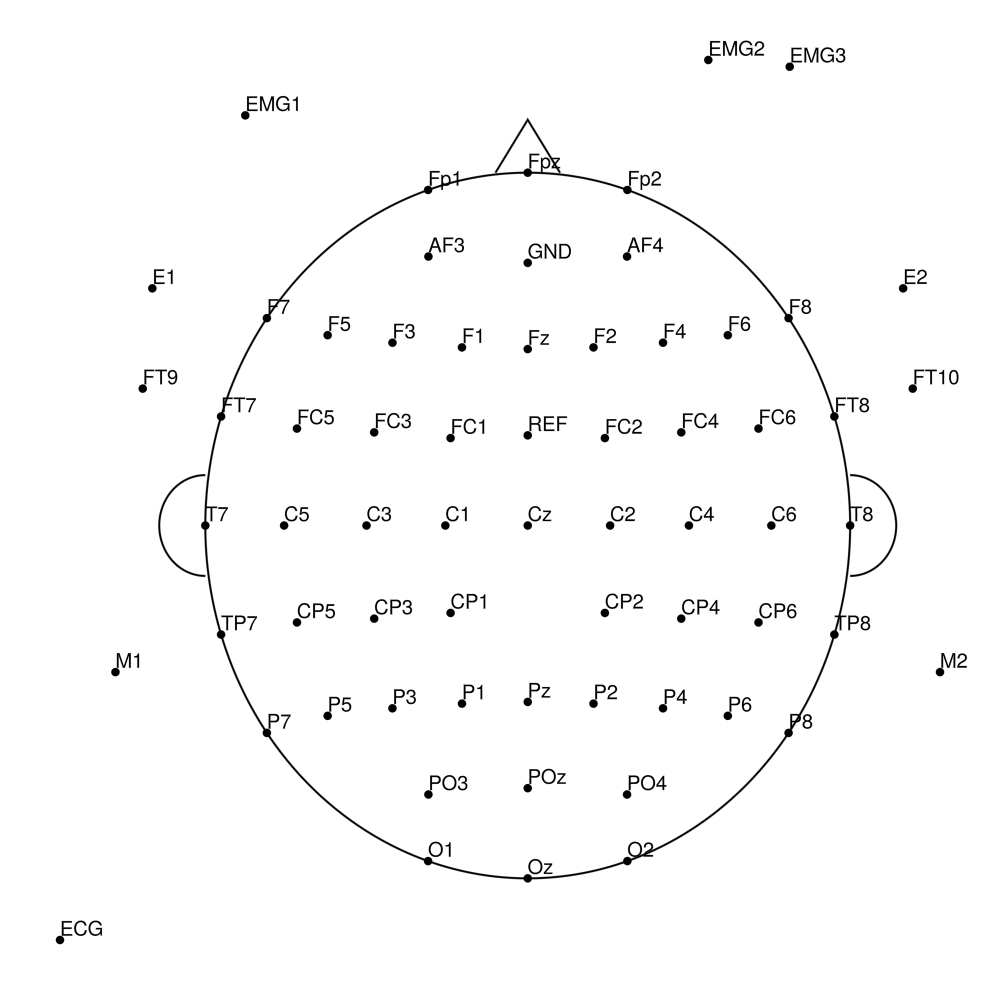
BC-SL-32 (32 channels) - BrainAmp sleep
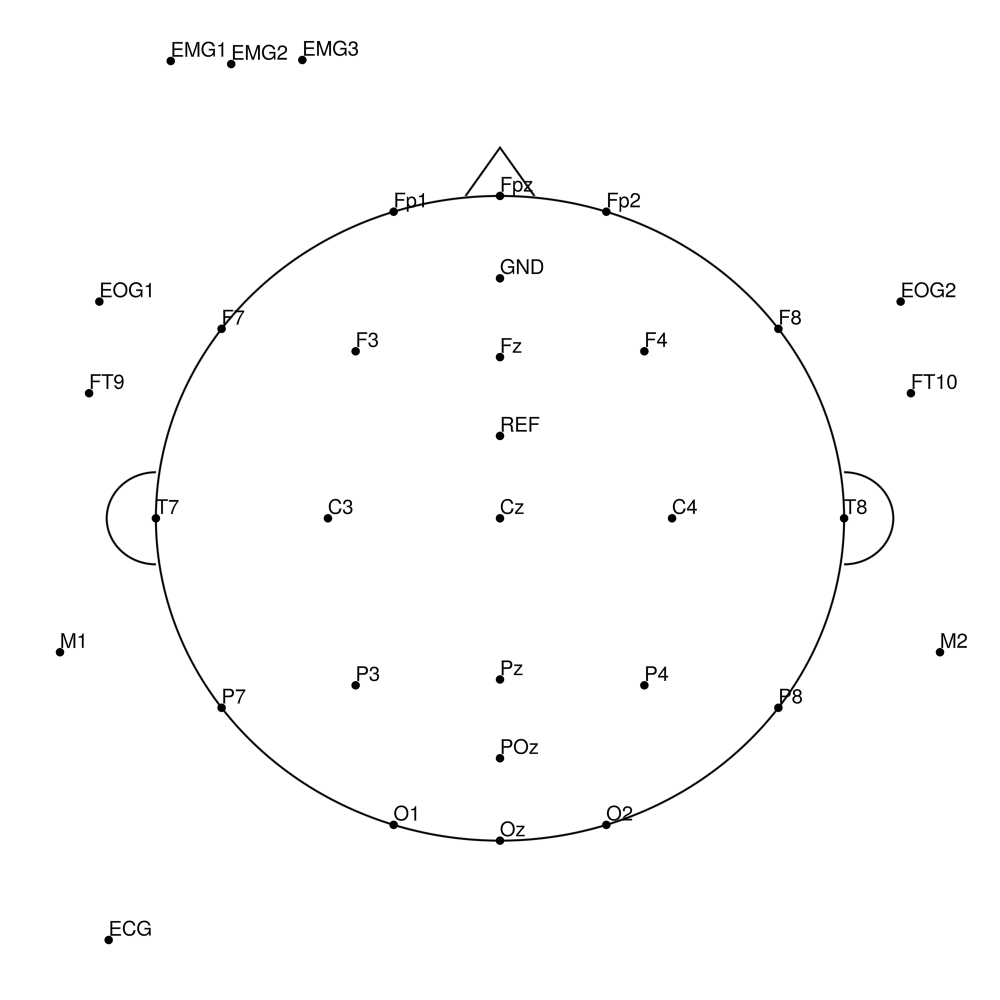
BC-SL-64 (64 channels) - BrainAmp sleep
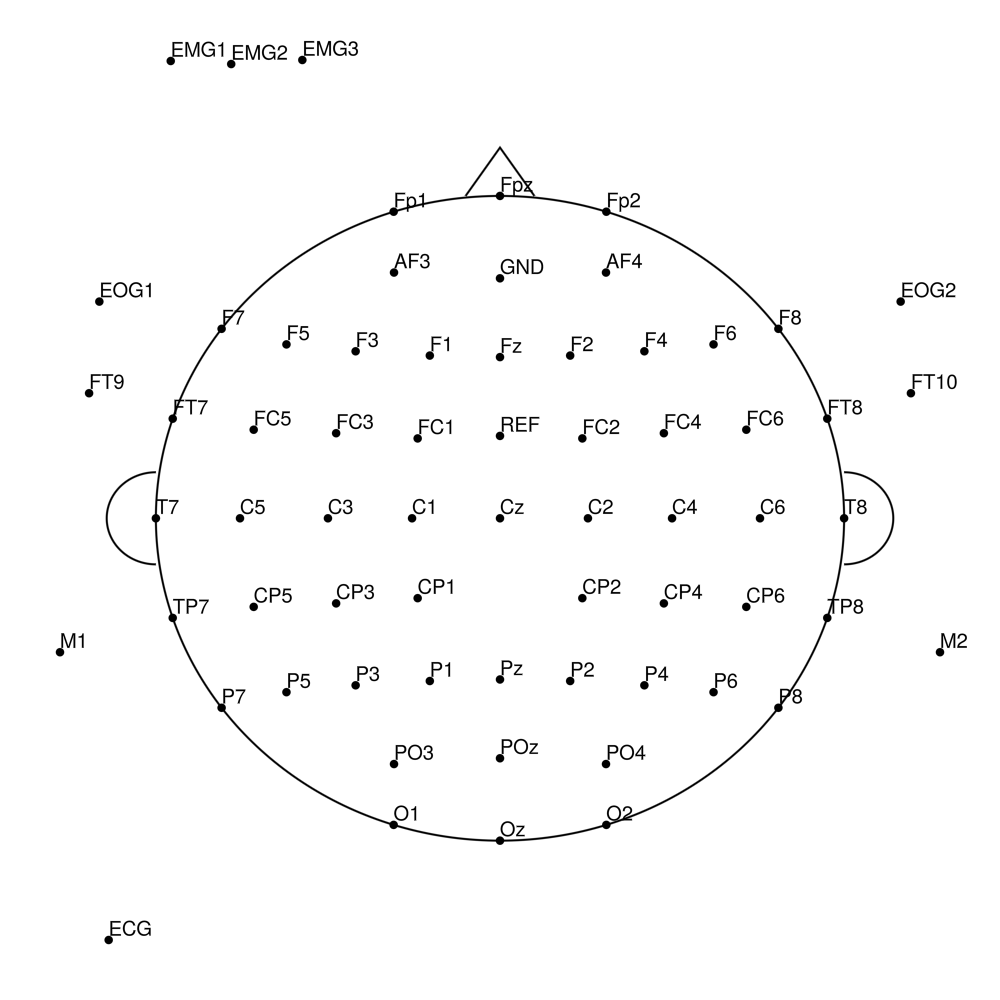
BC-SL-128 (128 channels) - BrainAmp sleep
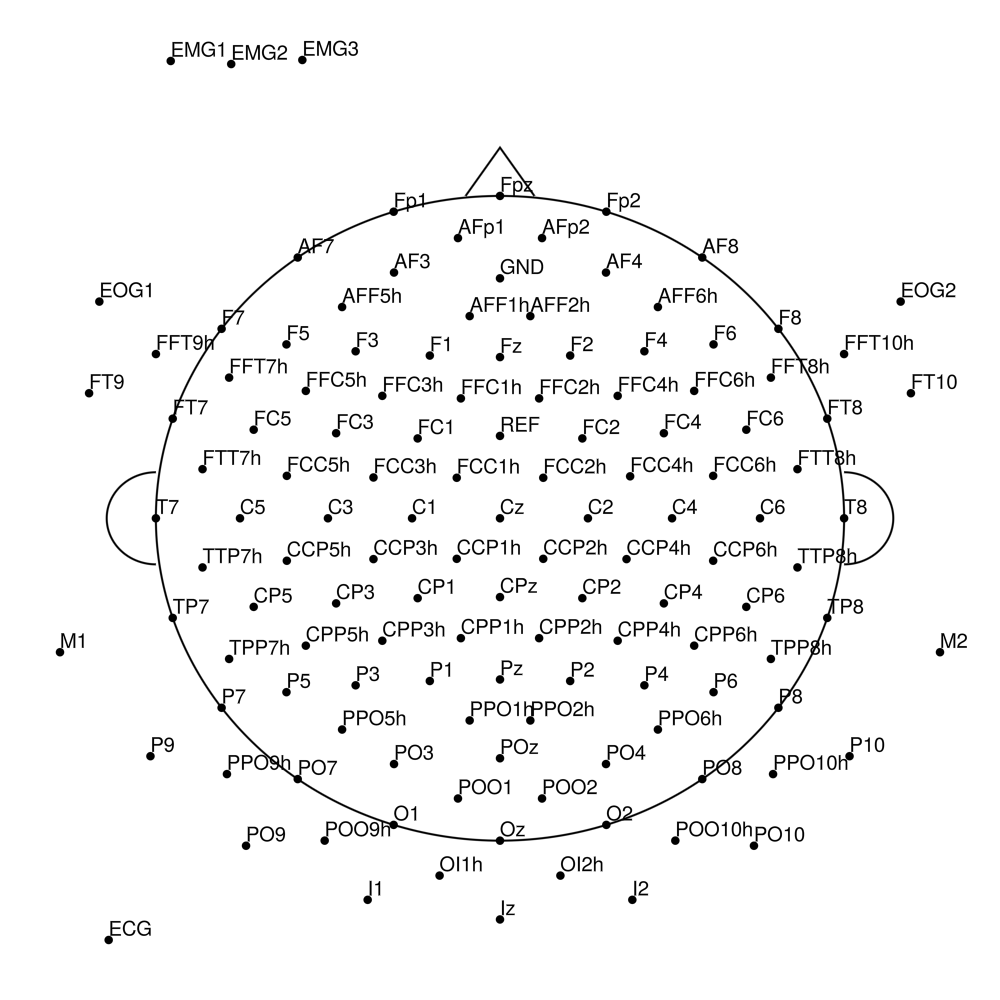
BCA-SL-32 (32 channels) - actiCHamp sleep
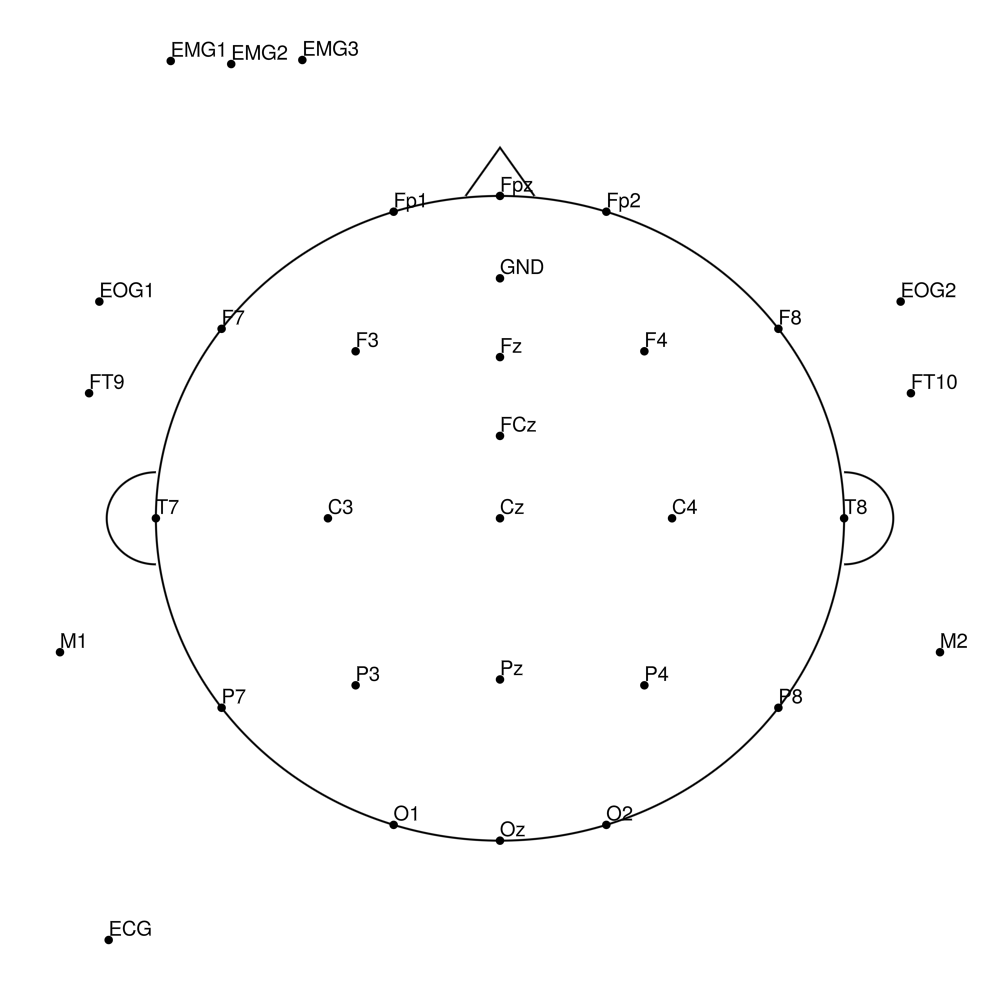
BCA-SL-64 (64 channels) - actiCHamp sleep
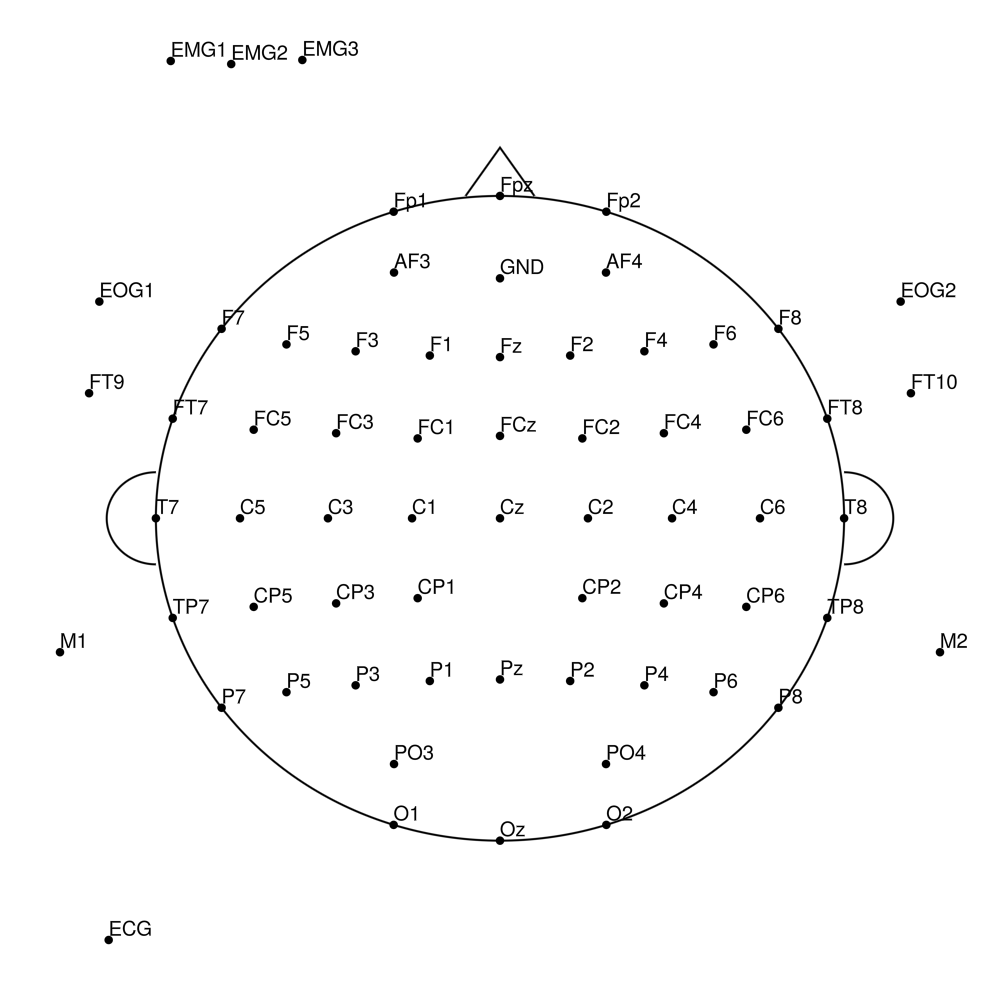
BCA-SL-128 (128 channels) - actiCHamp sleep
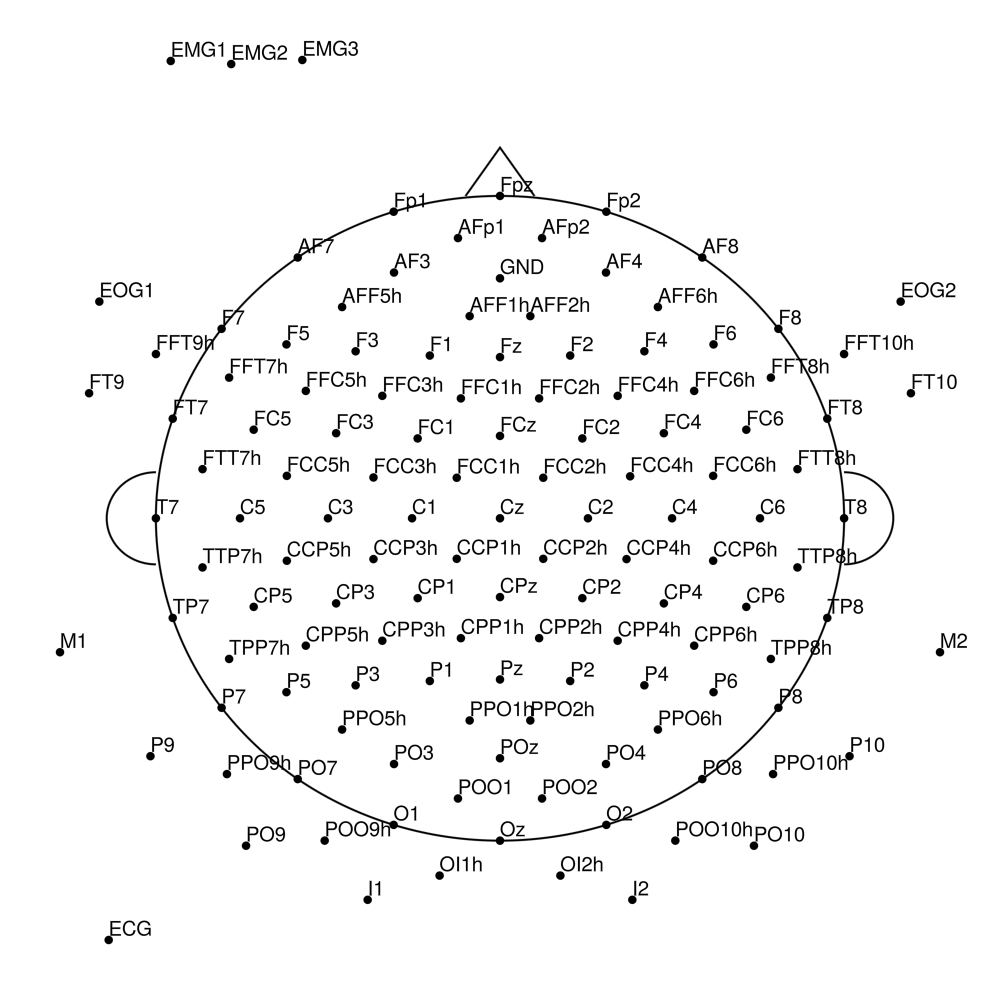
LiveCap
LC-32 (32 channels) - LiveAmp 32-channel
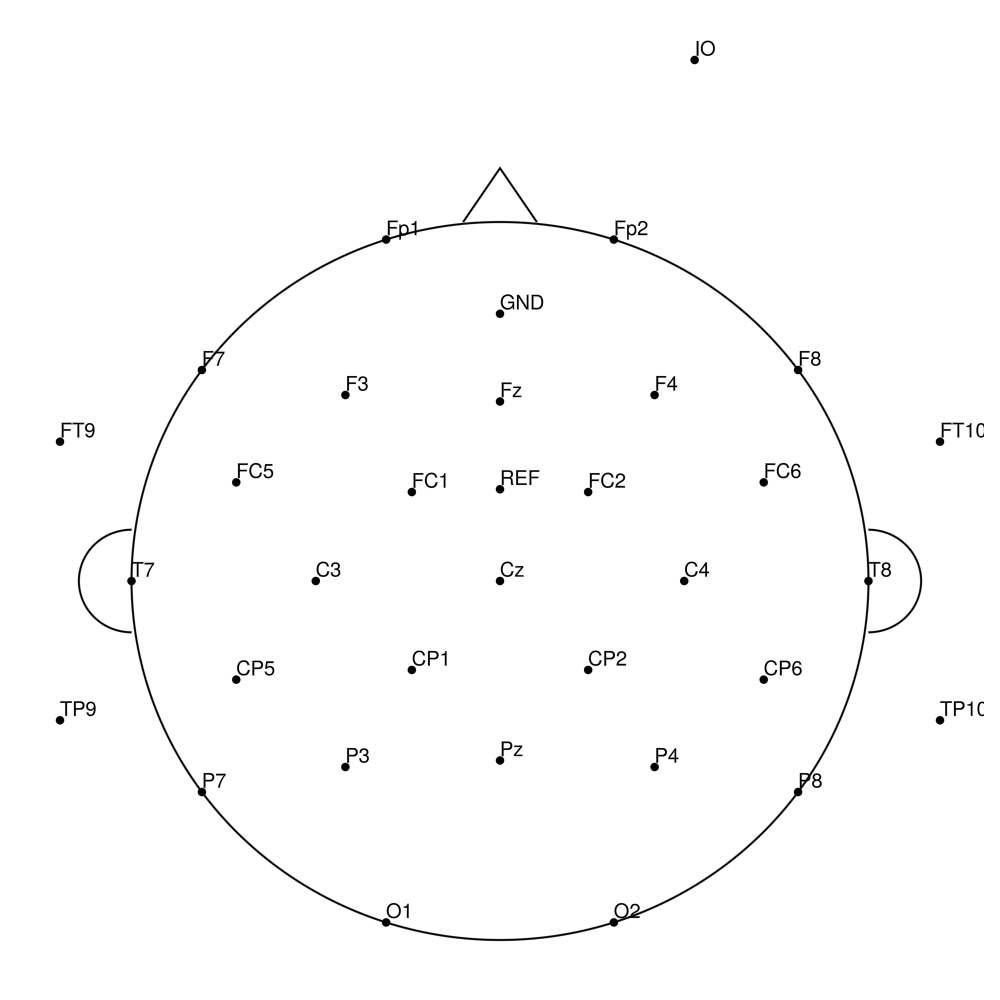
LC-64 (64 channels) - LiveAmp 64-channel
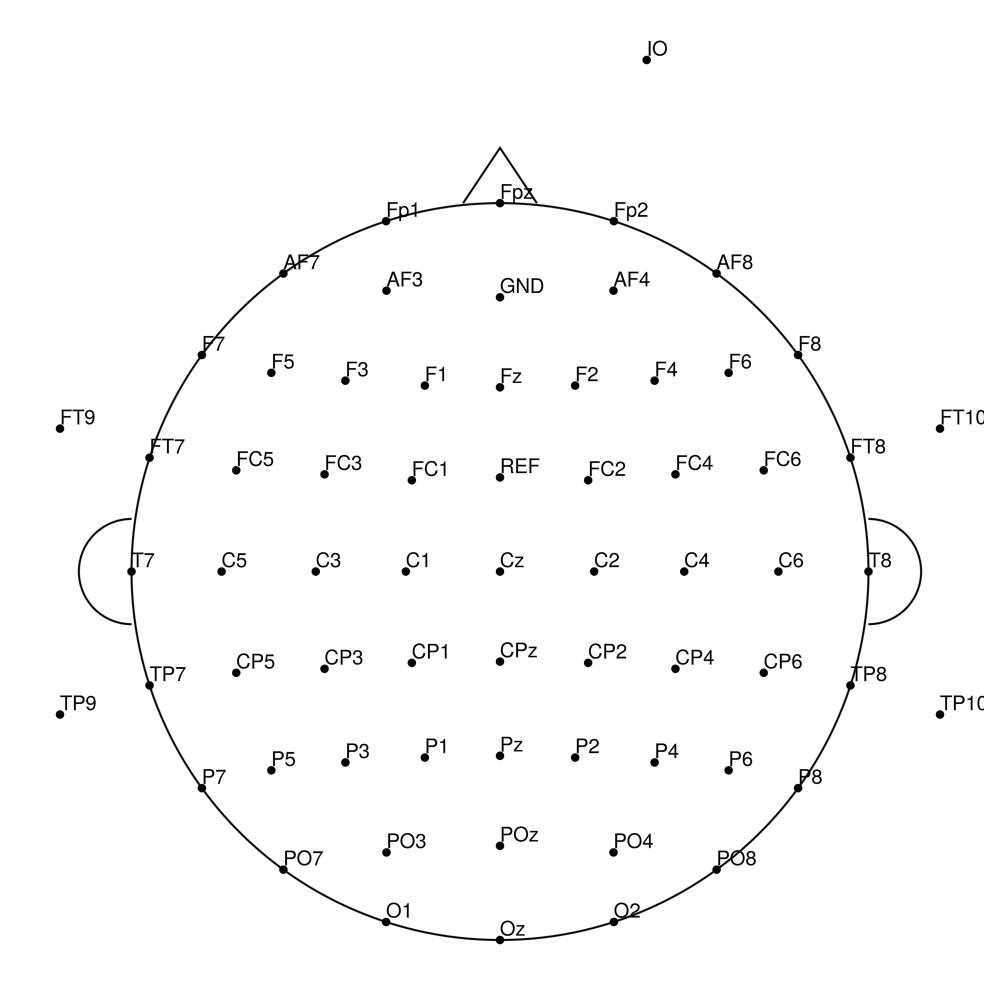
LC-SL-32_Ch1-24 (32 channels) - LiveCap sleep (24 + 8)
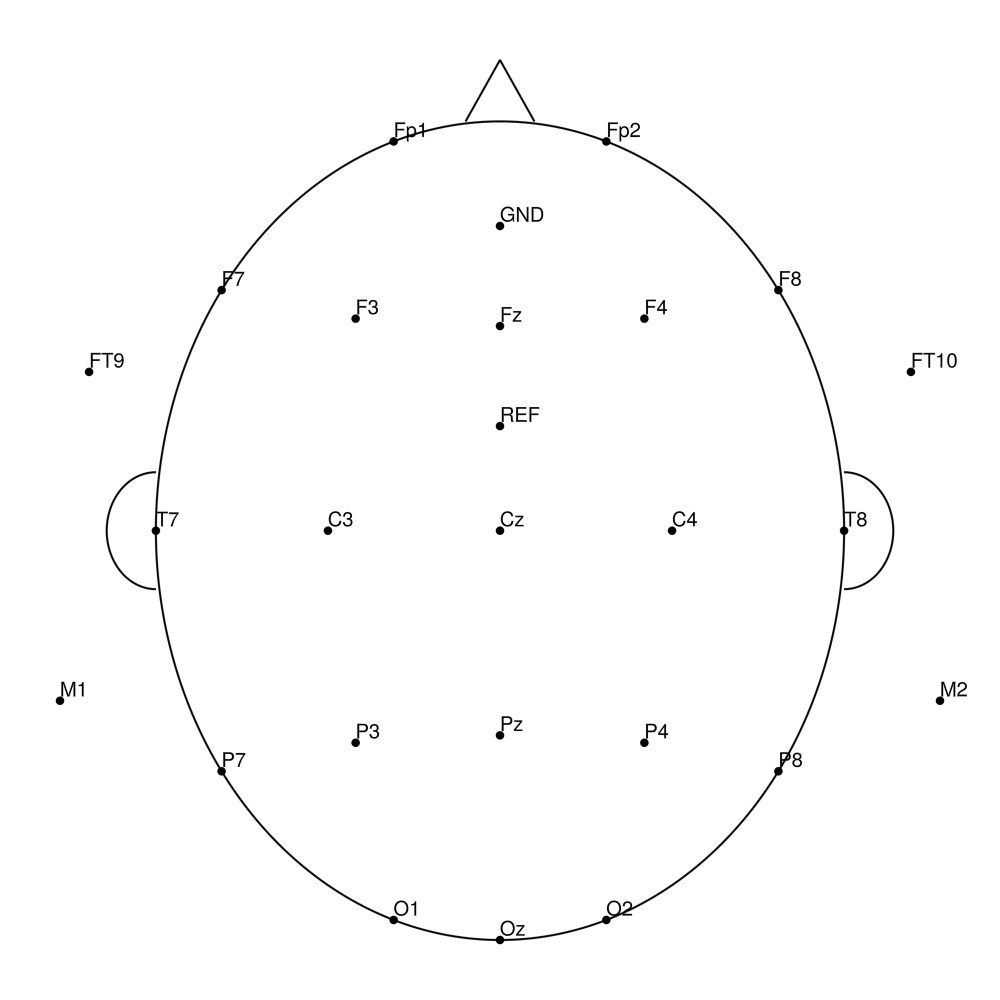
R-Net
RNP-BA-32 (32 channels) - R-Net for BrainAmp
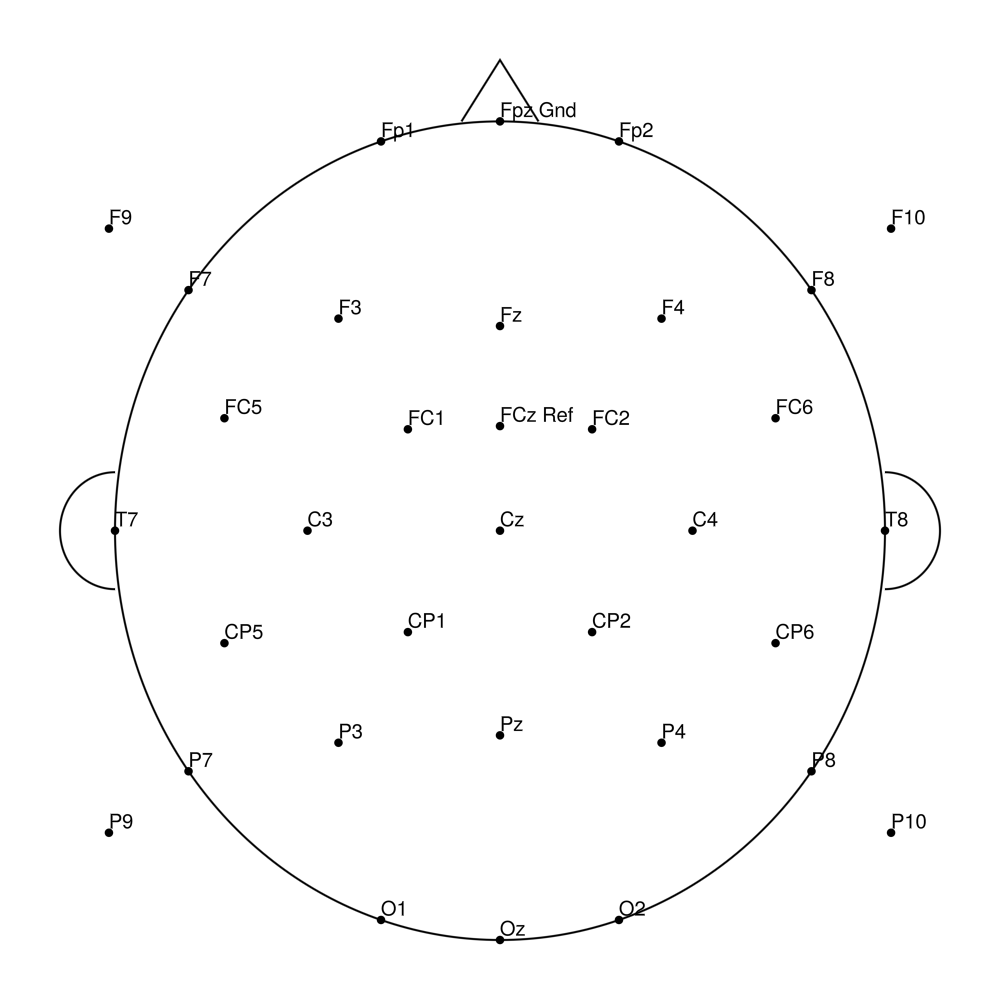
RNP-BA-64 (64 channels) - R-Net for BrainAmp
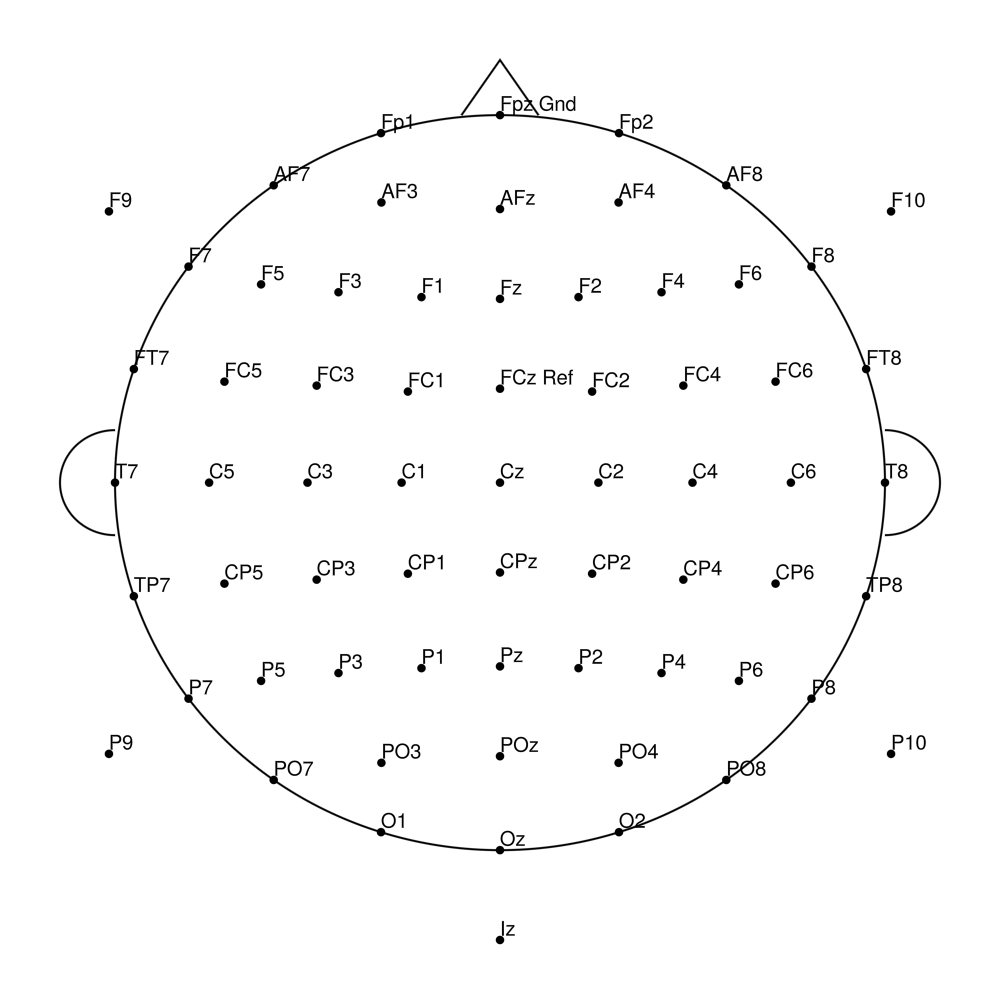
RNP-BA-96 (96 channels) - R-Net for BrainAmp
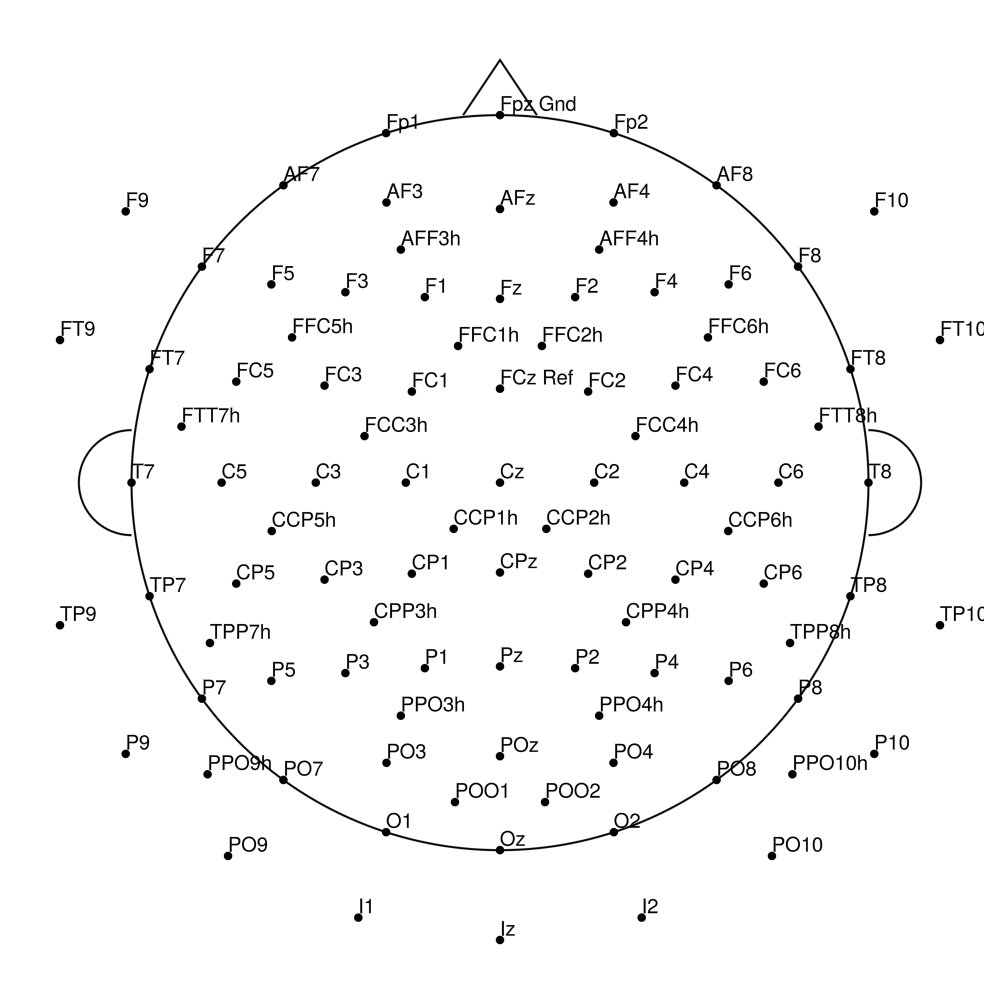
RNP-BA-128 (128 channels) - R-Net for BrainAmp
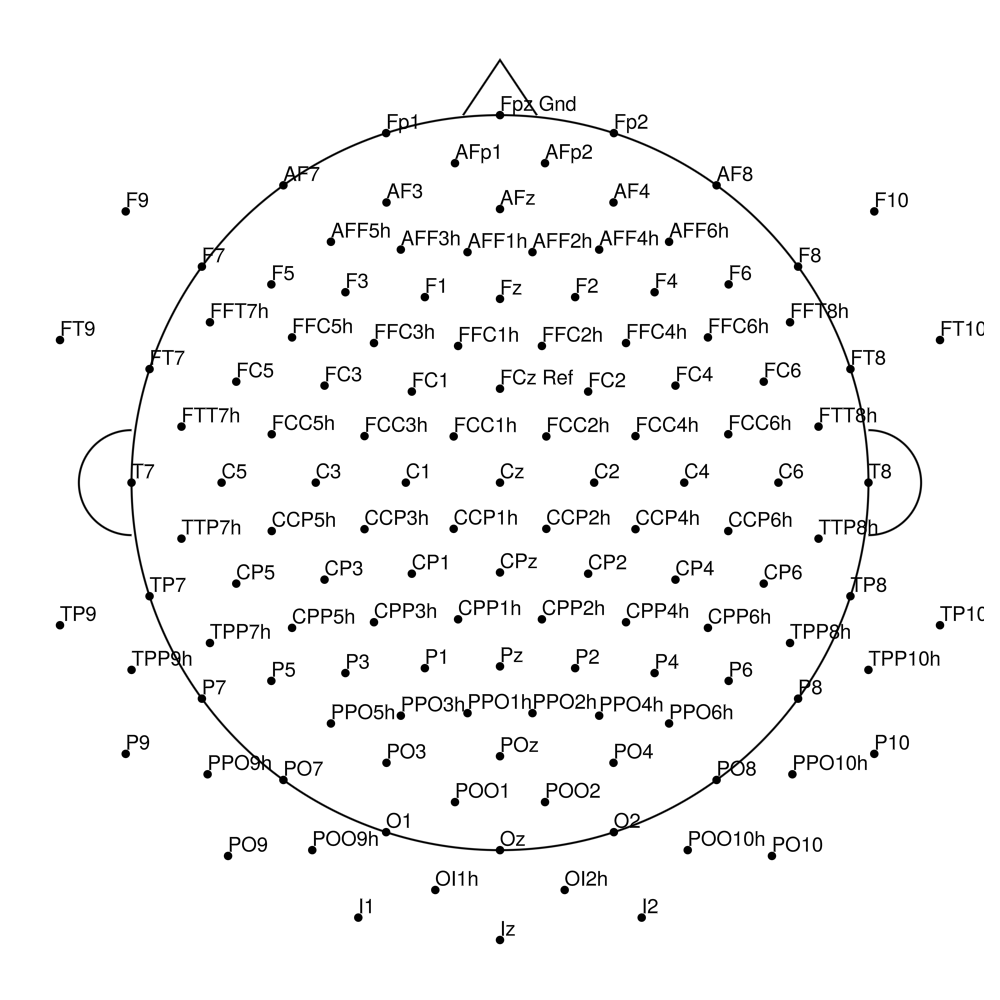
RNP-LA-32 (32 channels) - R-Net for LiveAmp
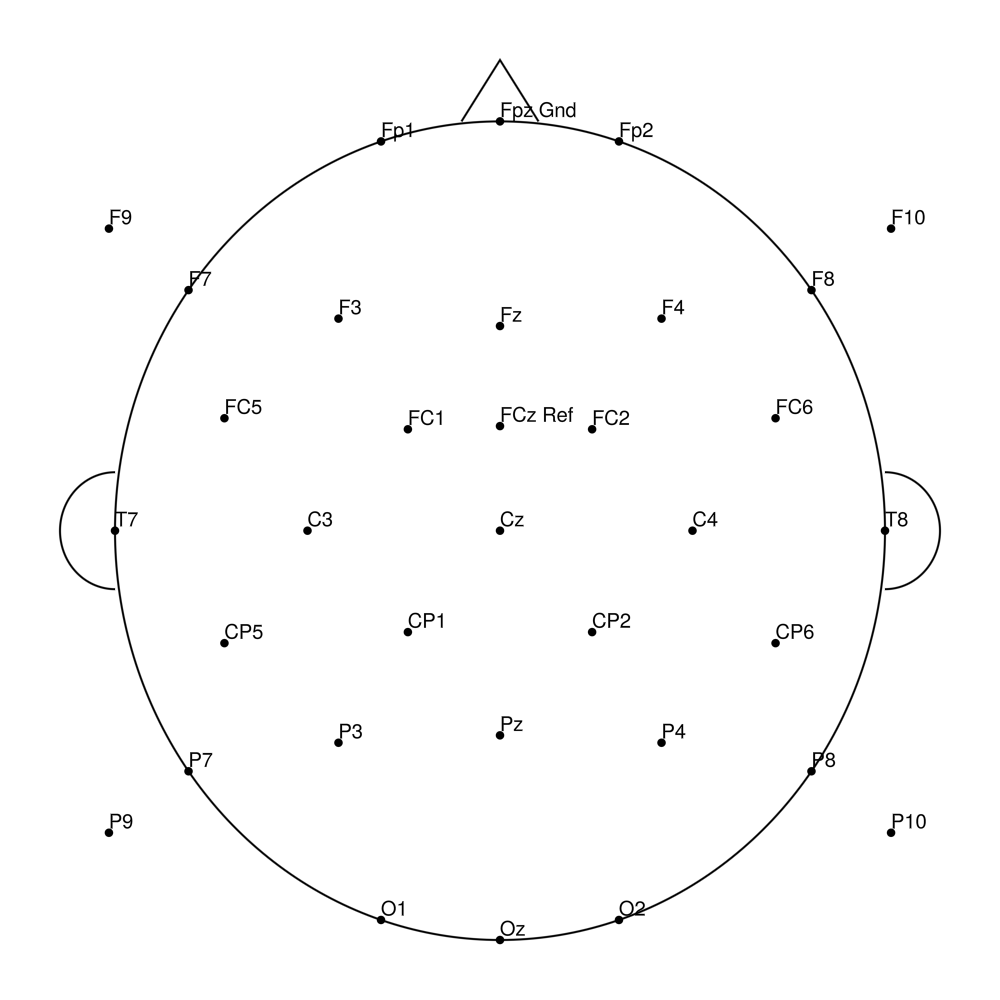
RNP-LA-64 (64 channels) - R-Net for LiveAmp
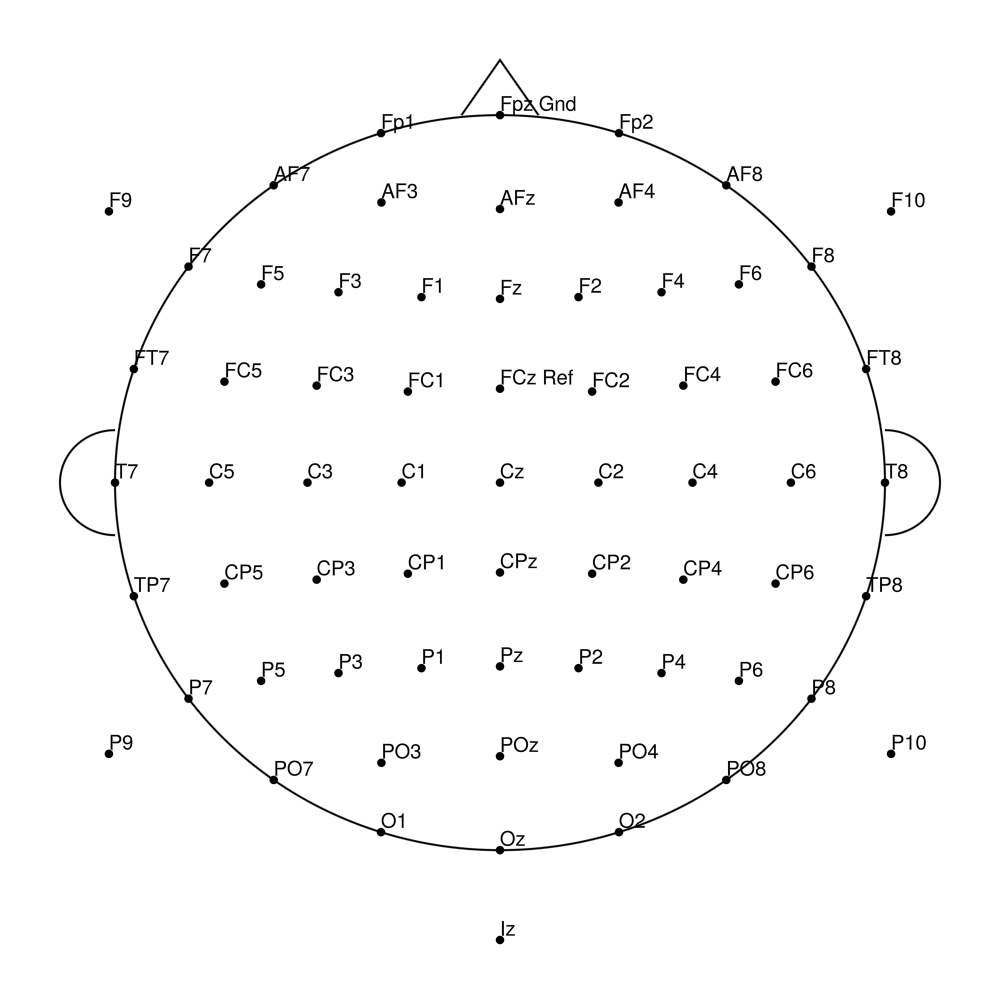
RNP-AC-32 (32 channels) - R-Net for actiCHamp Plus
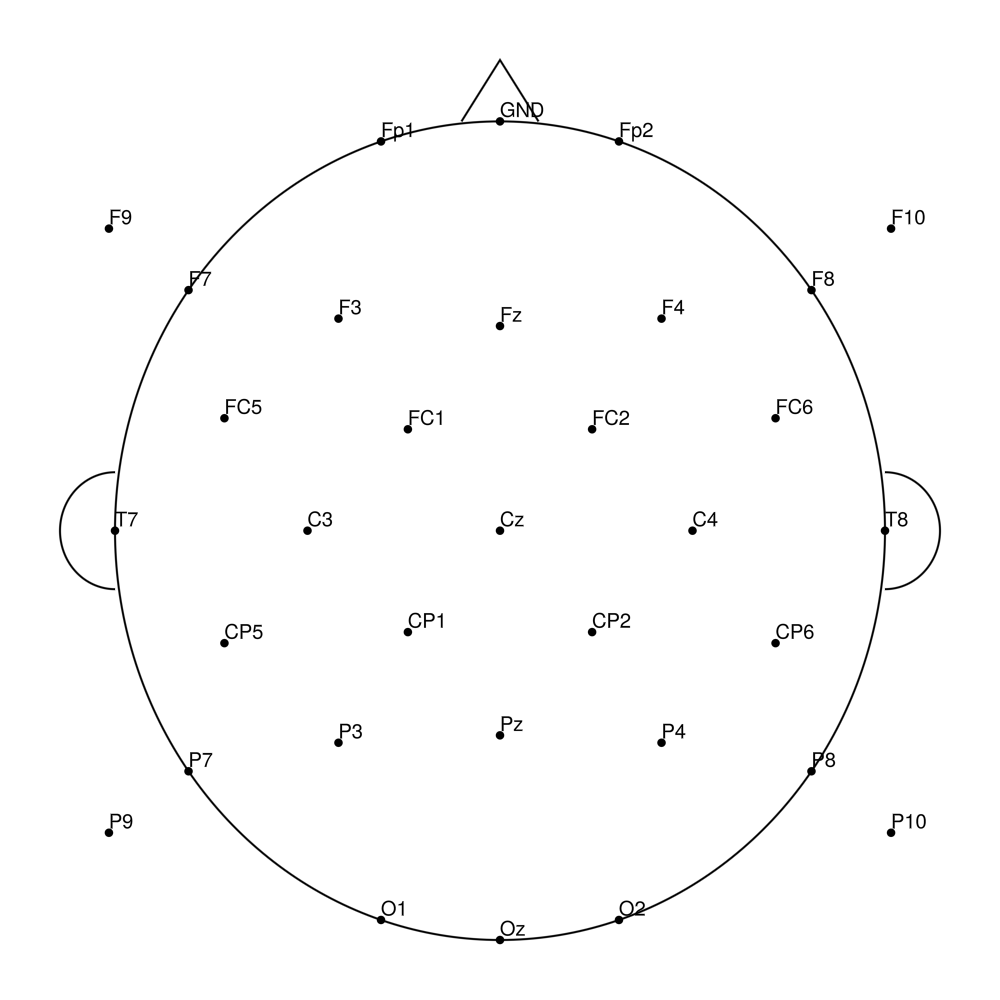
RNP-AC-64 (64 channels) - R-Net for actiCHamp Plus (with Iz)
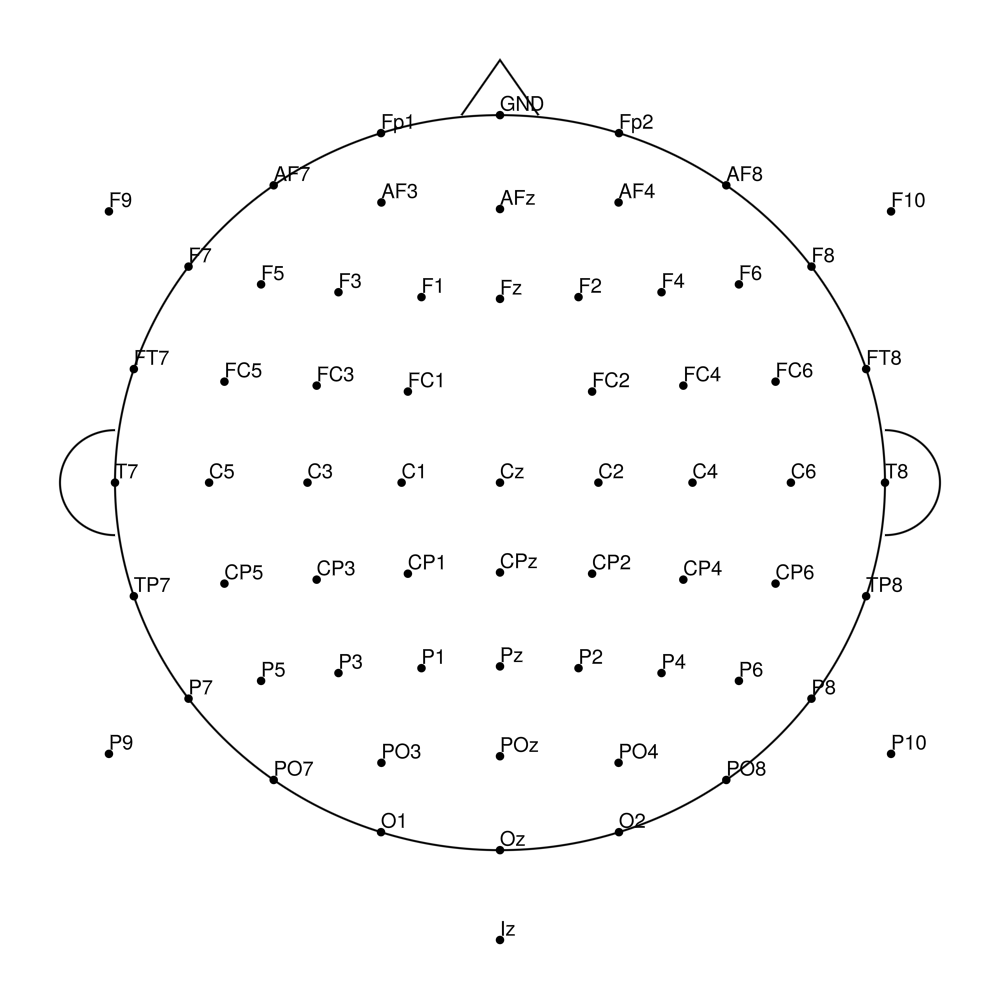
RNP-AC-96 (96 channels) - R-Net for actiCHamp Plus (with Iz)
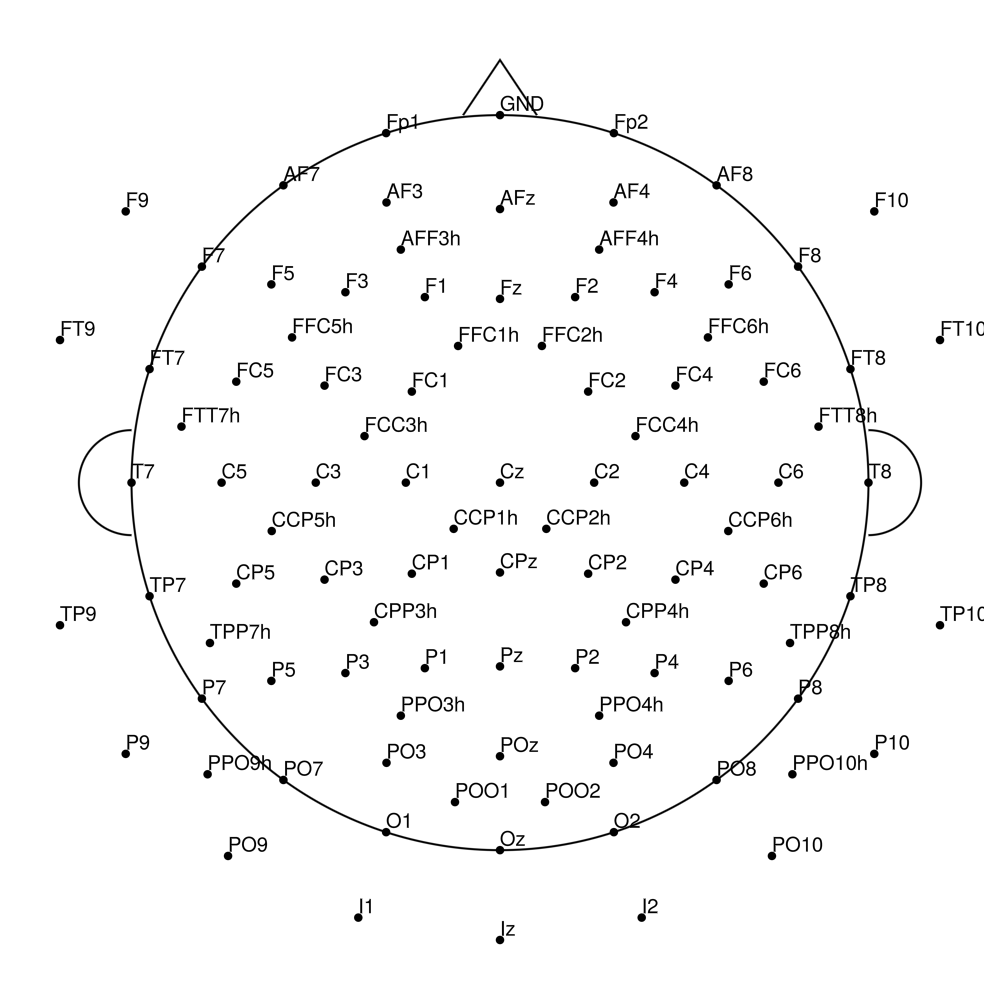
RNP-AC-128 (128 channels) - R-Net for actiCHamp Plus (with Iz)
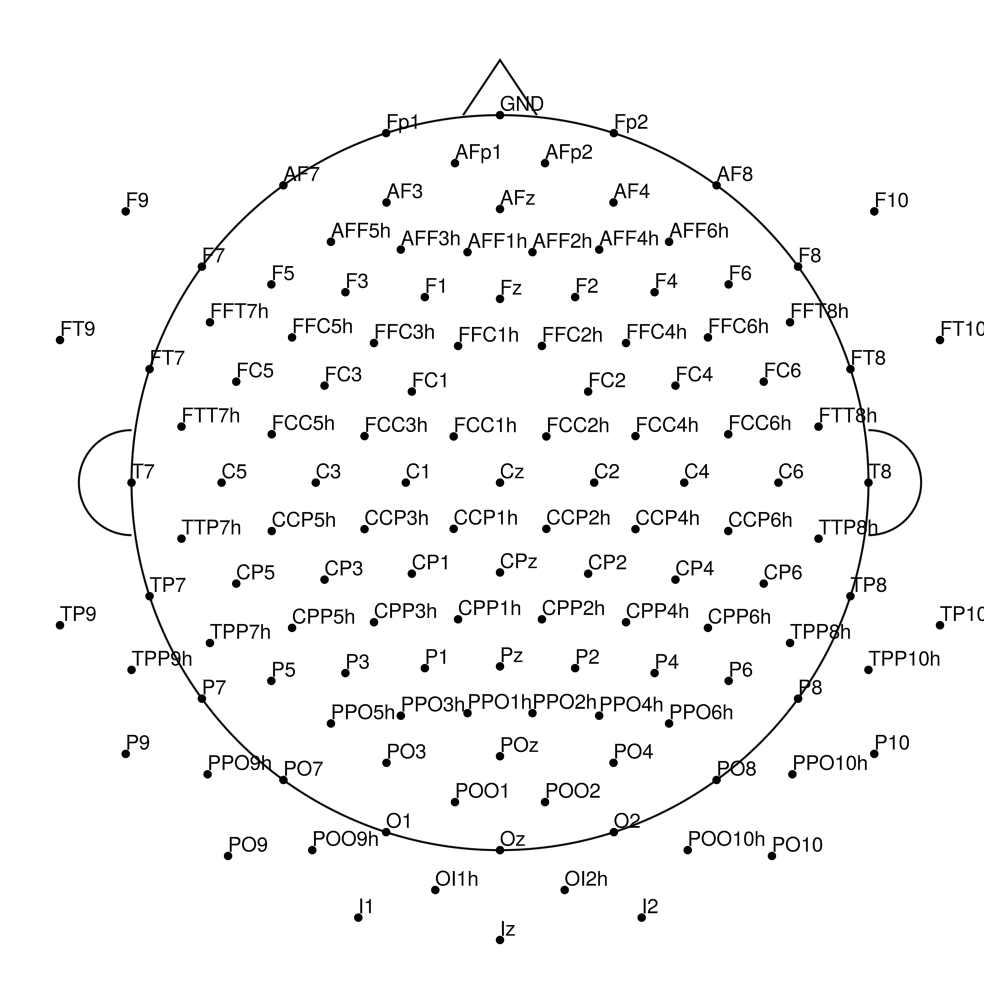
RNP-BA-MR3-32 (32 channels) - R-Net MR3
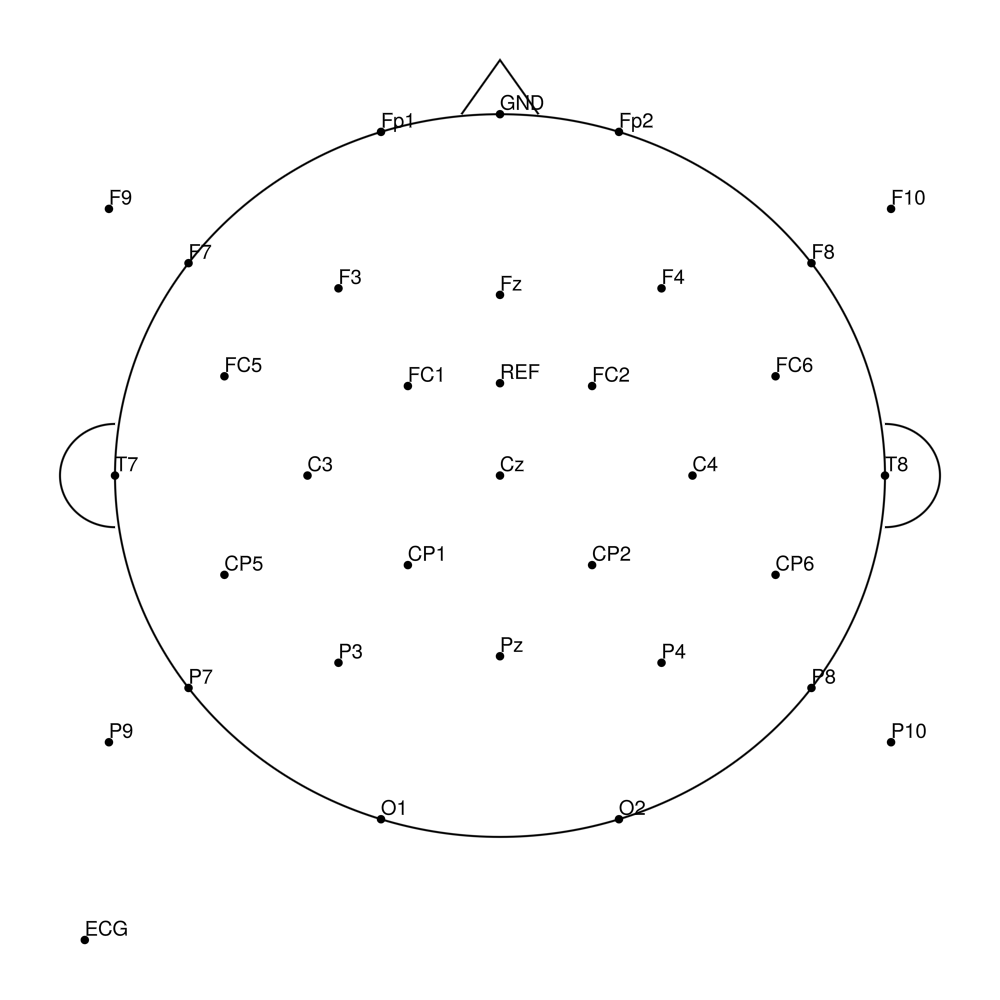
RNP-BA-MR3-64 (64 channels) - R-Net MR3
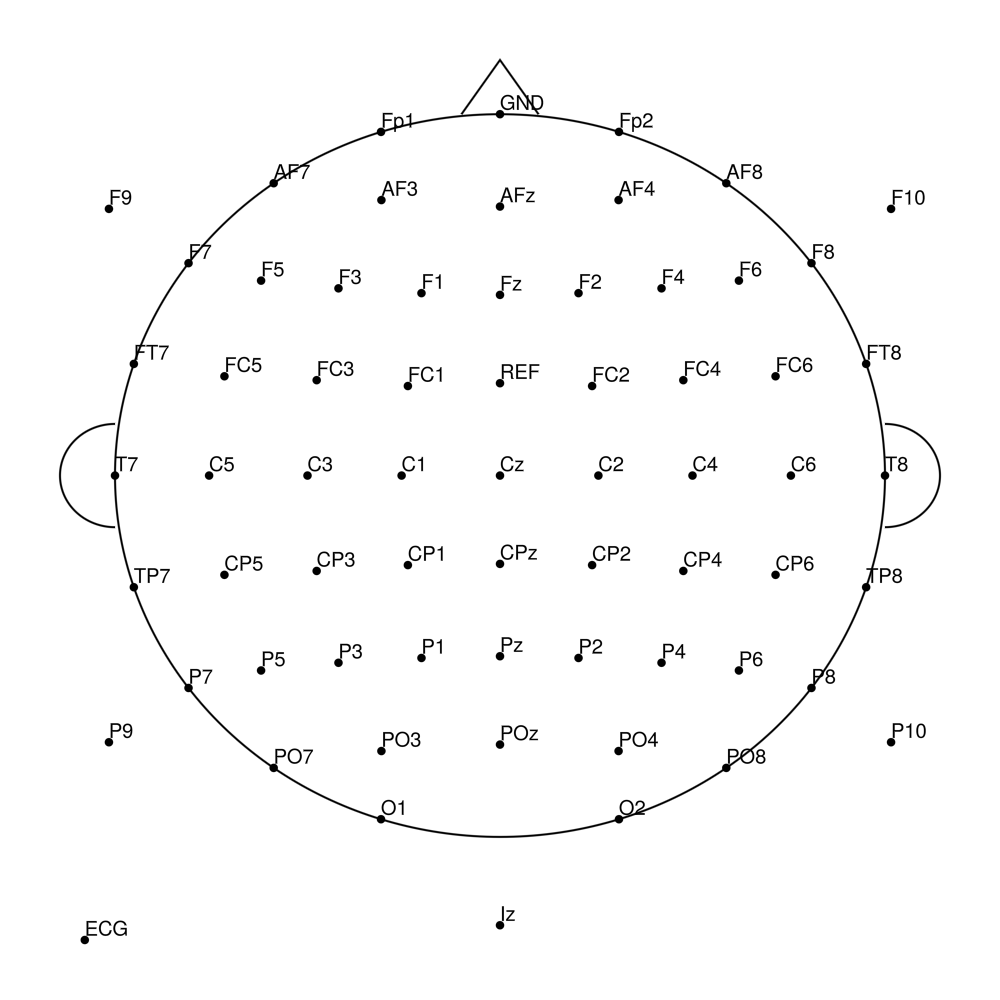
RNP-BA-MR3-96 (96 channels) - R-Net MR3
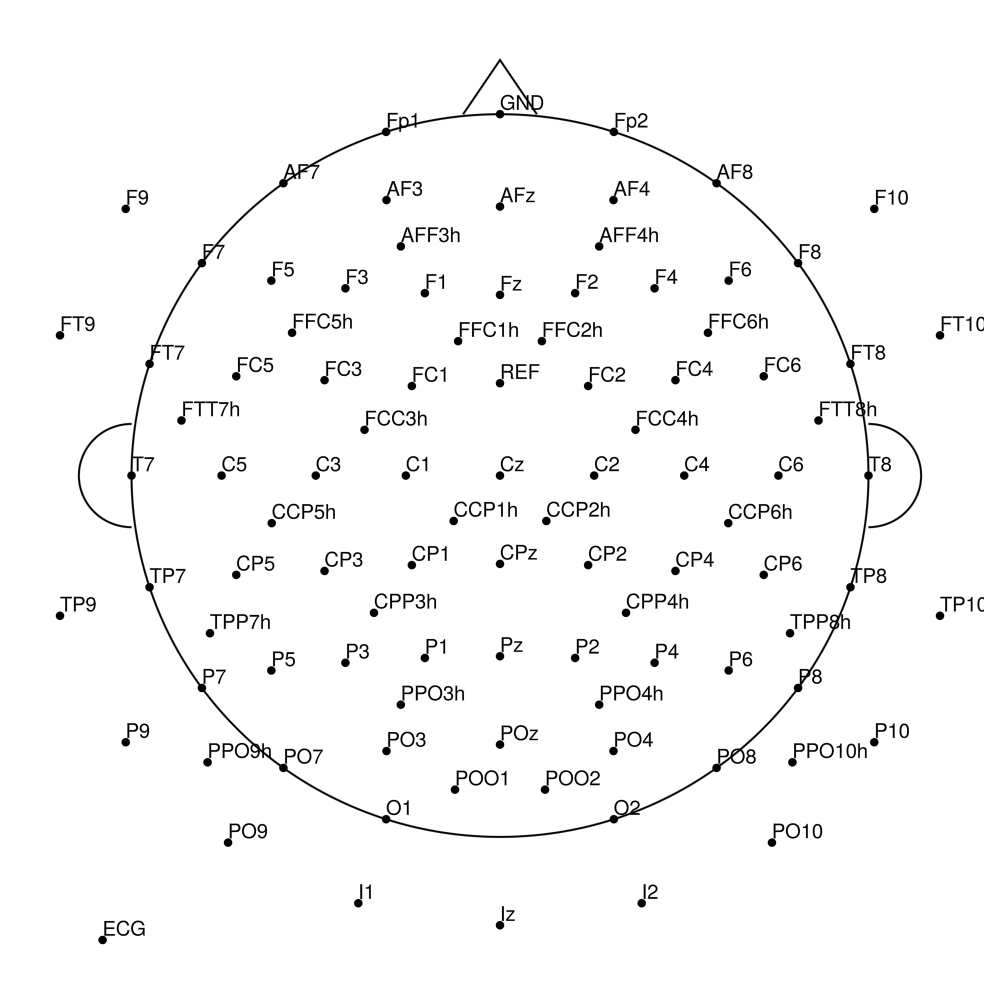
RNP-BA-MR3-128 (128 channels) - R-Net MR3
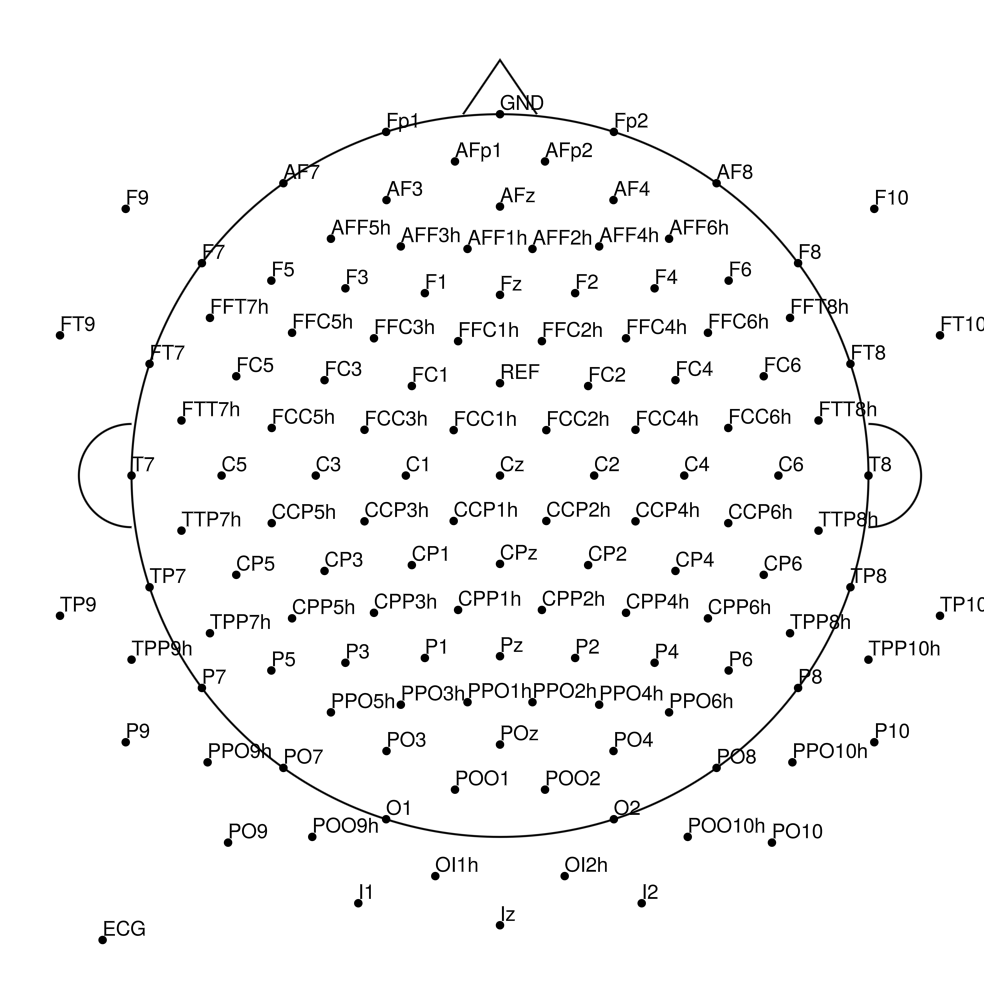
RNP-AP-32 (32 channels) - R-Net for actiCHamp Plus
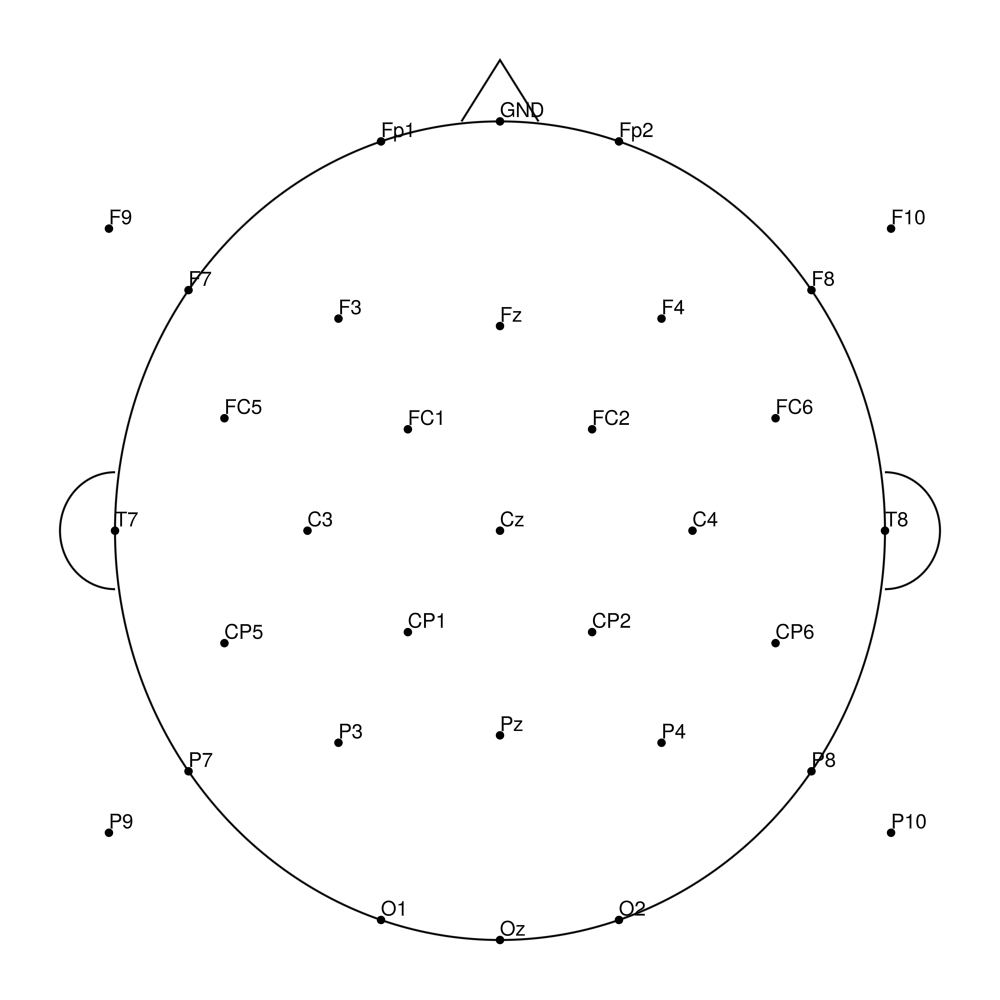
RNP-AP-64 (64 channels) - R-Net for actiCHamp Plus (with FCz)
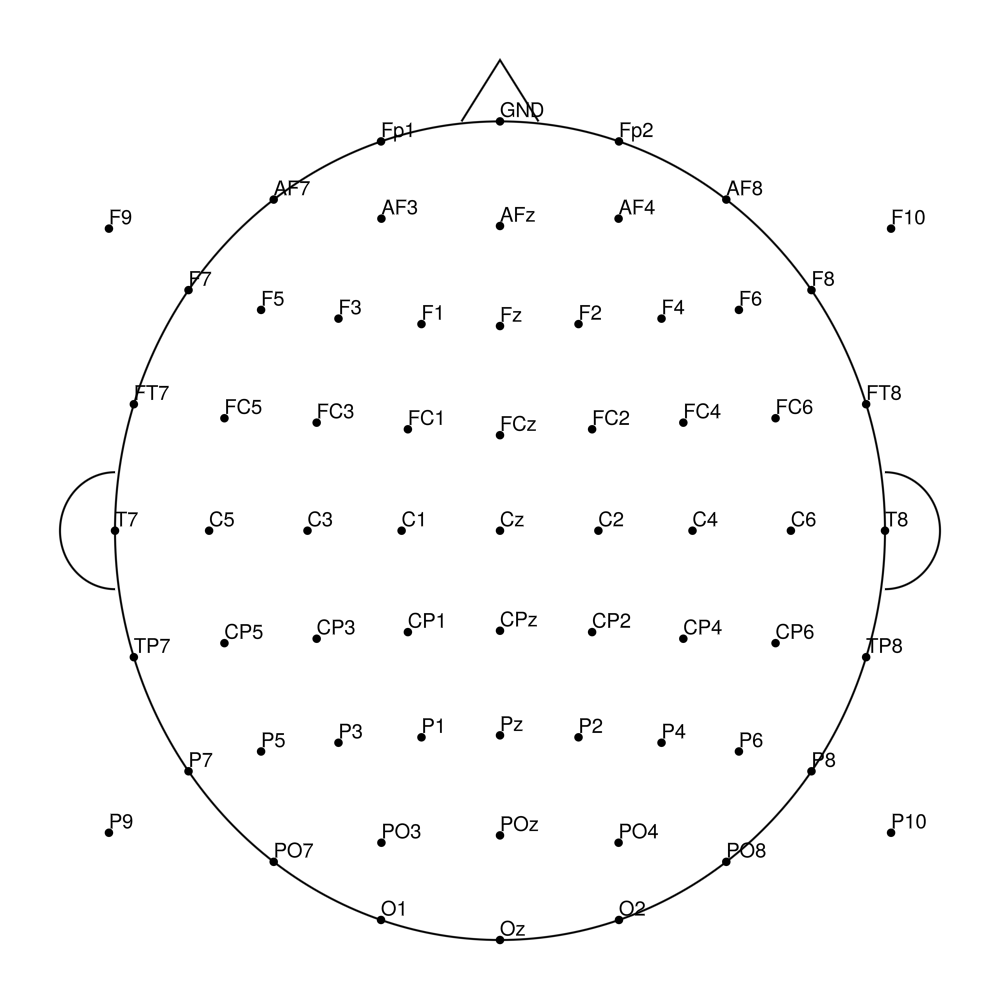
RNP-AP-96 (96 channels) - R-Net for actiCHamp Plus (with FCz)
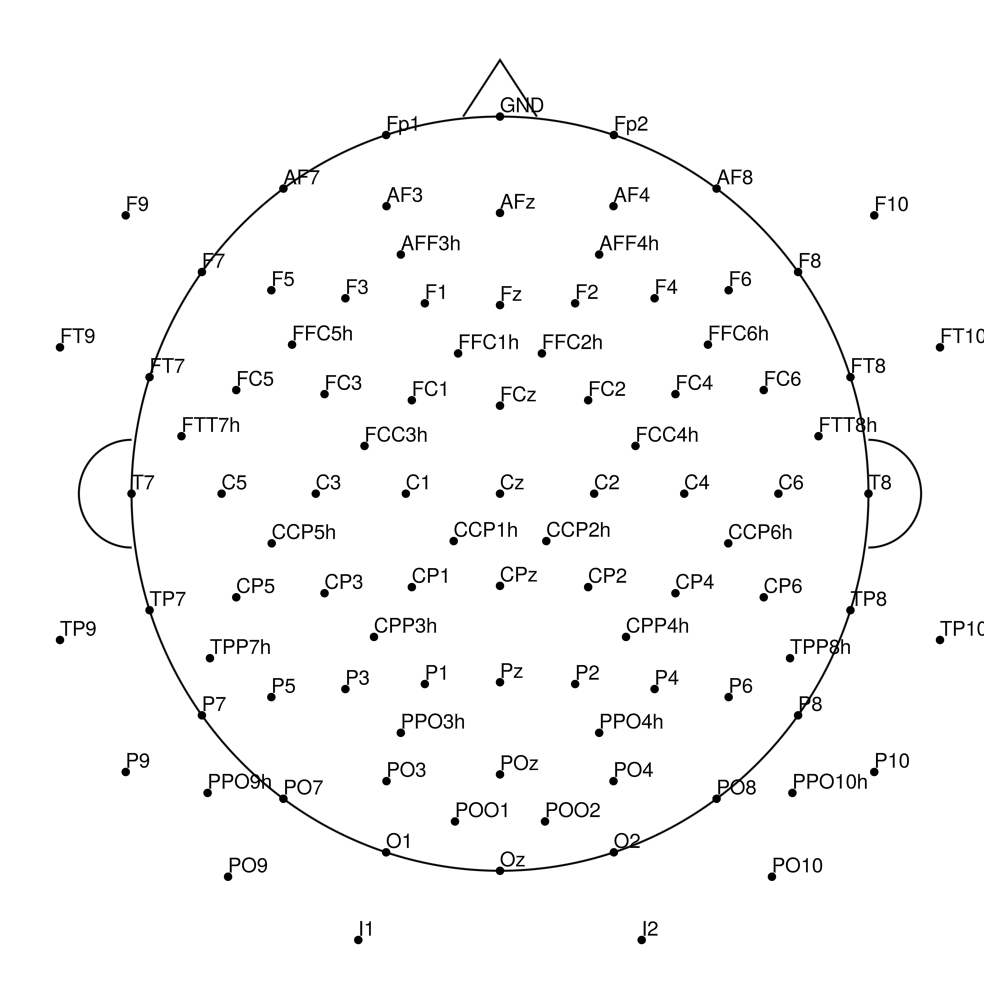
RNP-AP-128 (128 channels) - R-Net for actiCHamp Plus (with FCz)
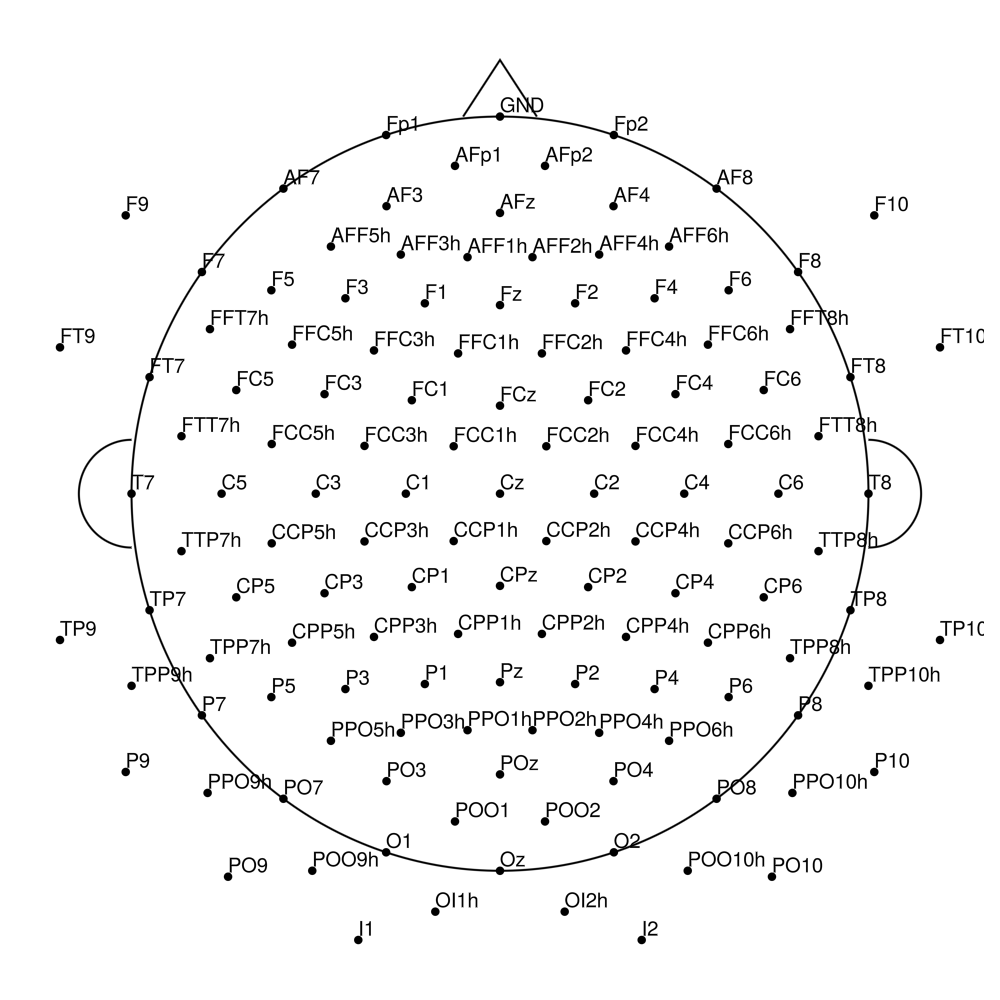
RNP-AP-160 (160 channels) - R-Net for actiCHamp Plus (with FCz, Iz)
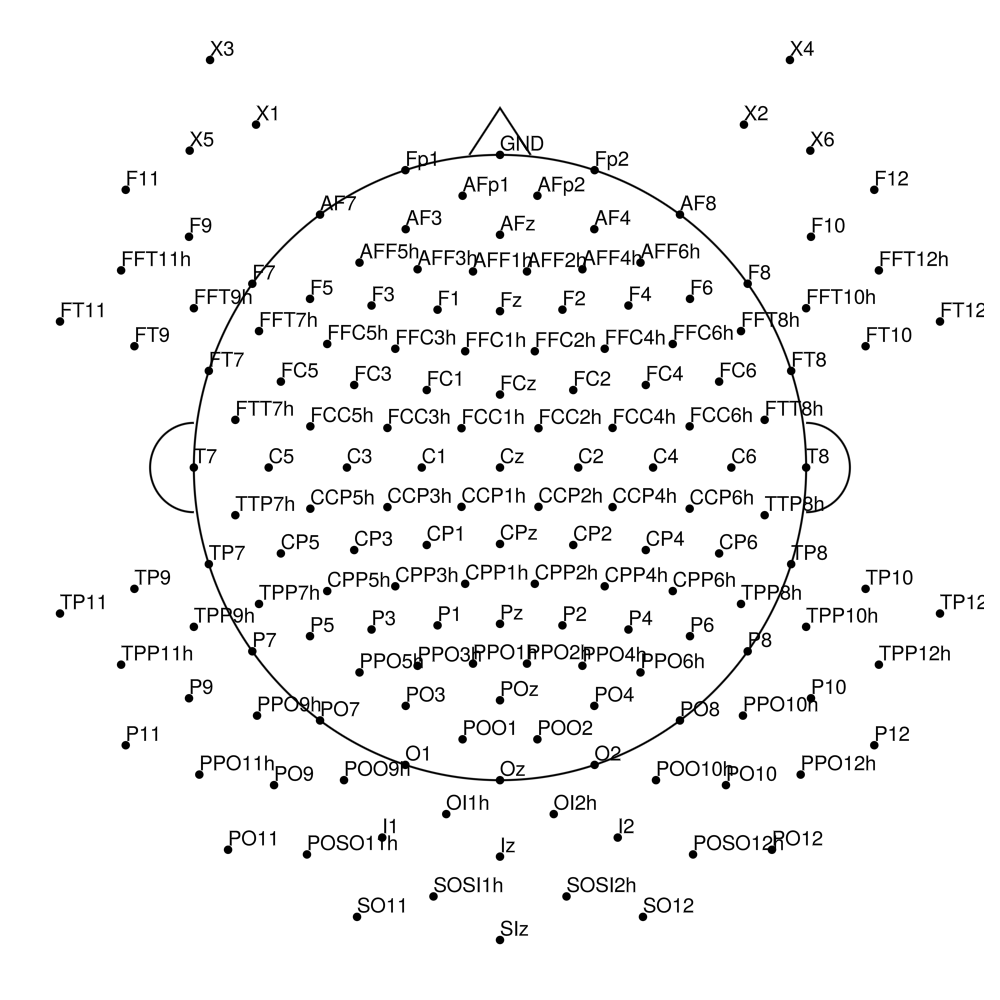
actiCAP (Standard)
AC-21 (21 channels)
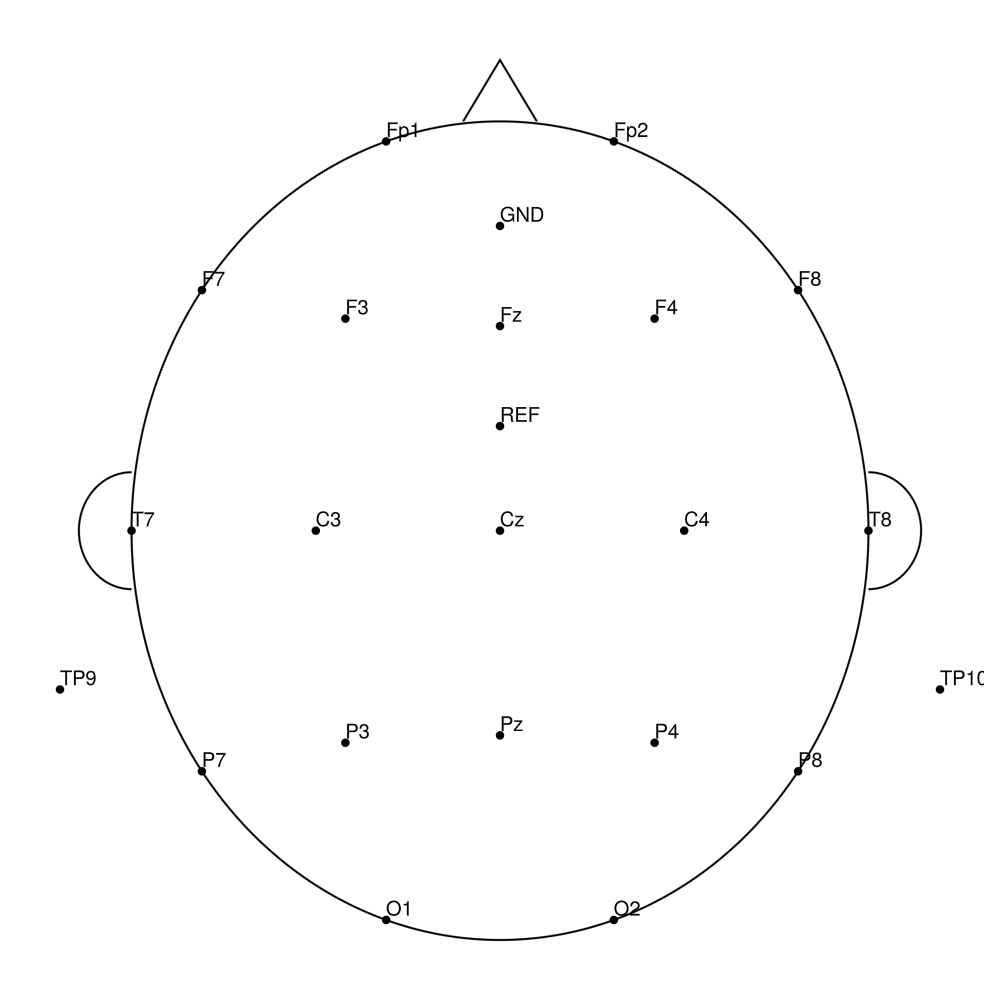
AC-32 (32 channels)
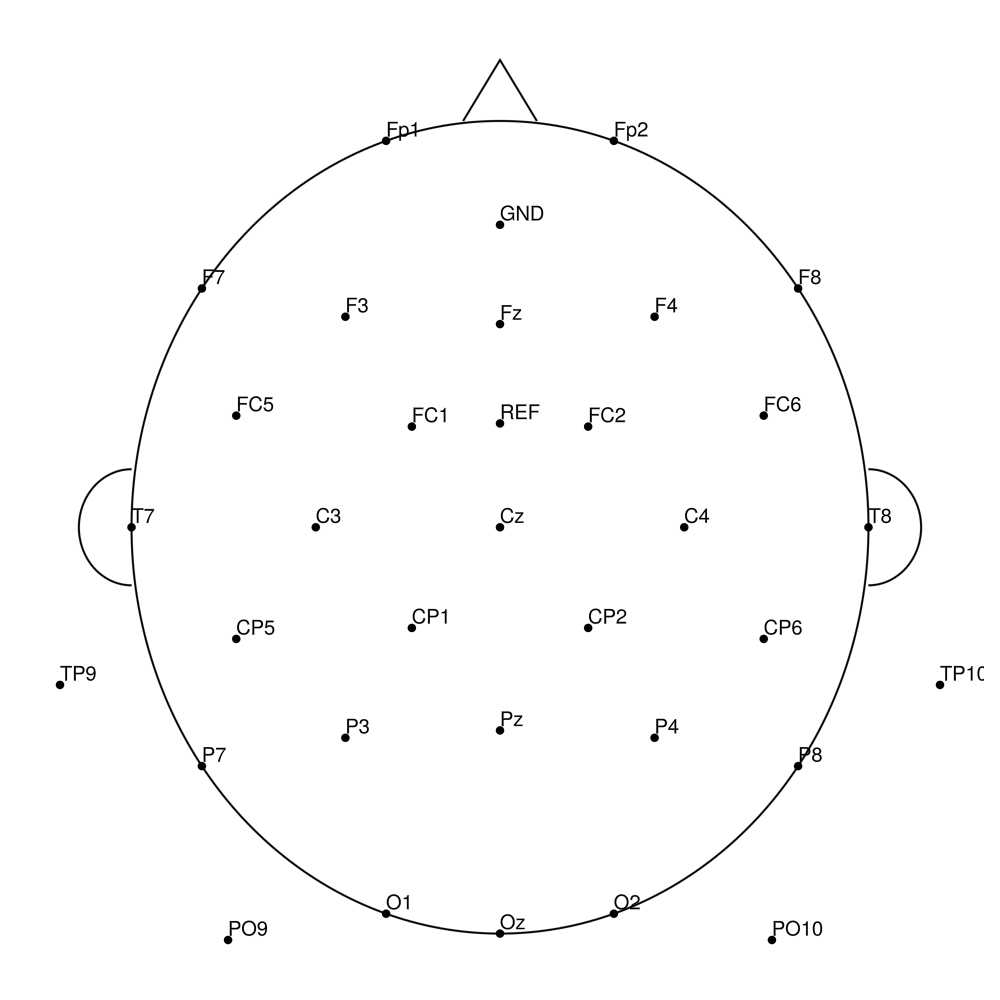
AC-64 (64 channels)
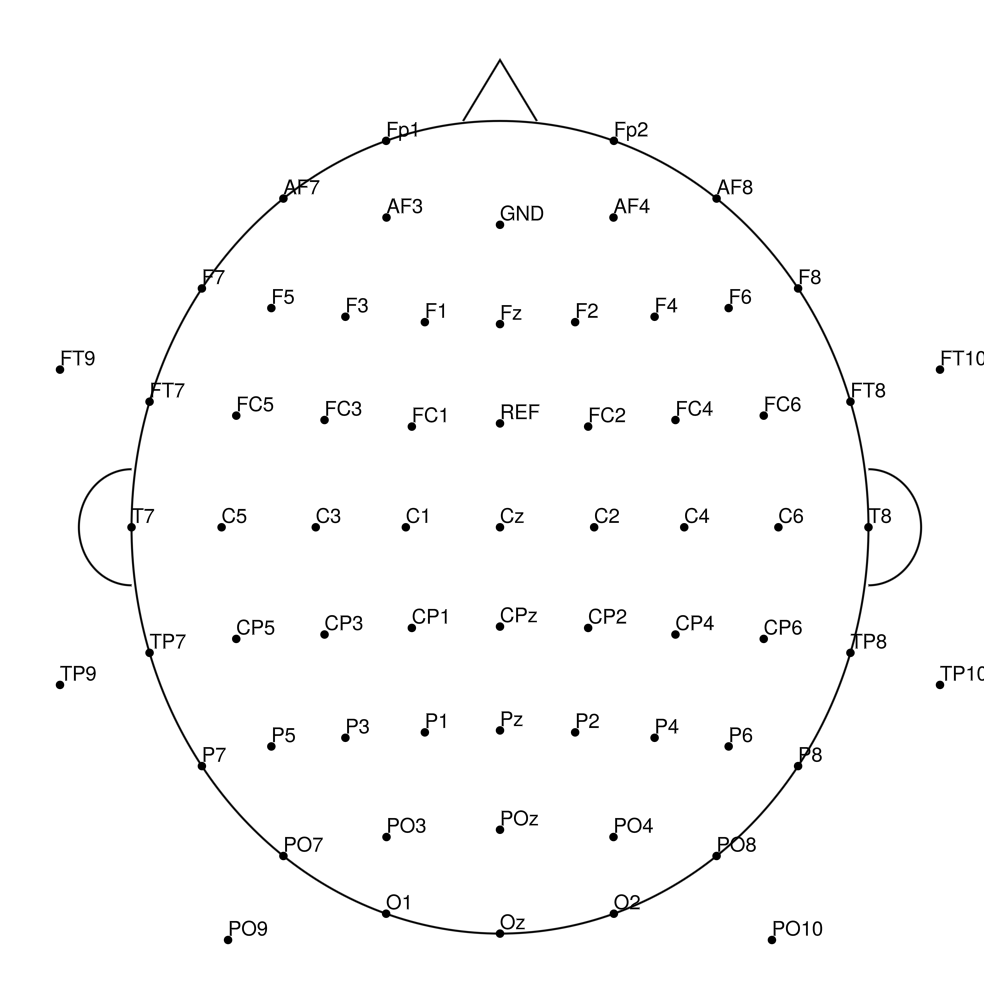
AC-96 (96 channels)
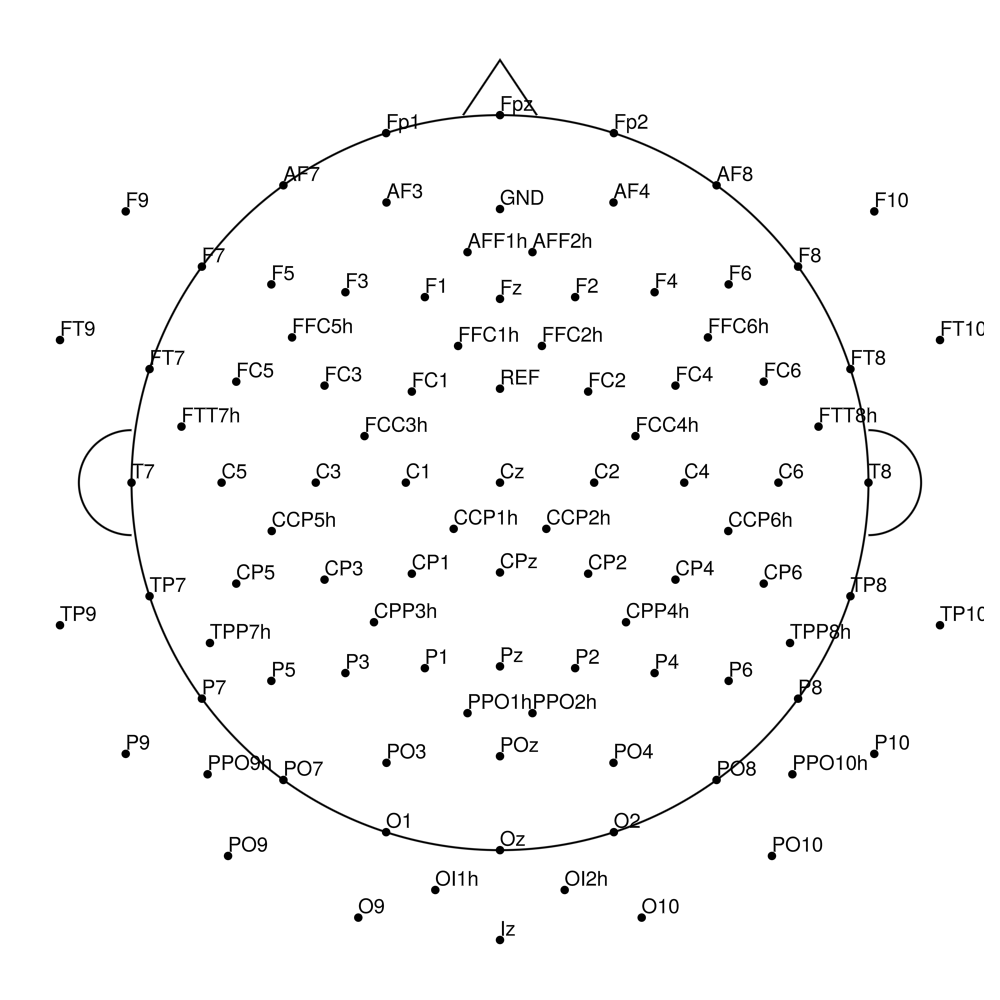
AC-128 (128 channels)
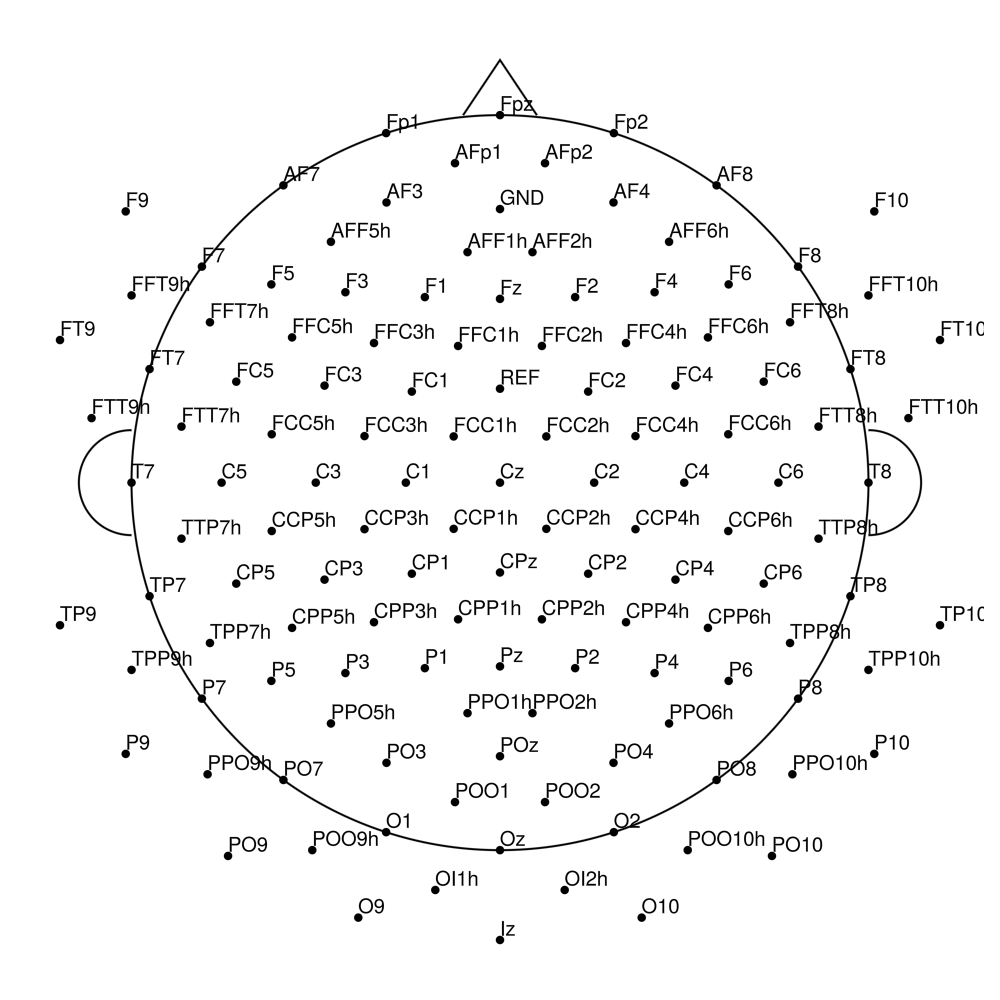
actiCAP Xpress Twist
Twist-32 (32 channels) - for BrainAmp (with REF)
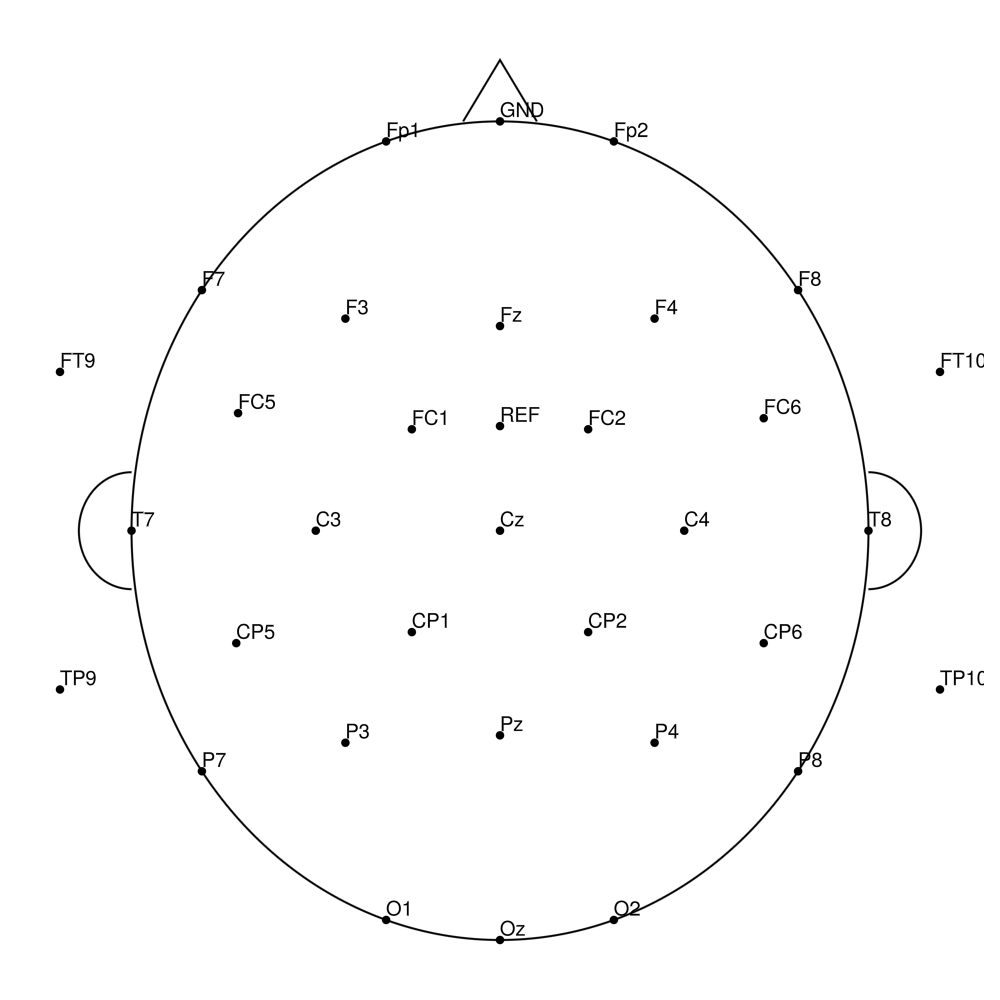
Twist-32 (32 channels) - for LiveAmp (with REF)
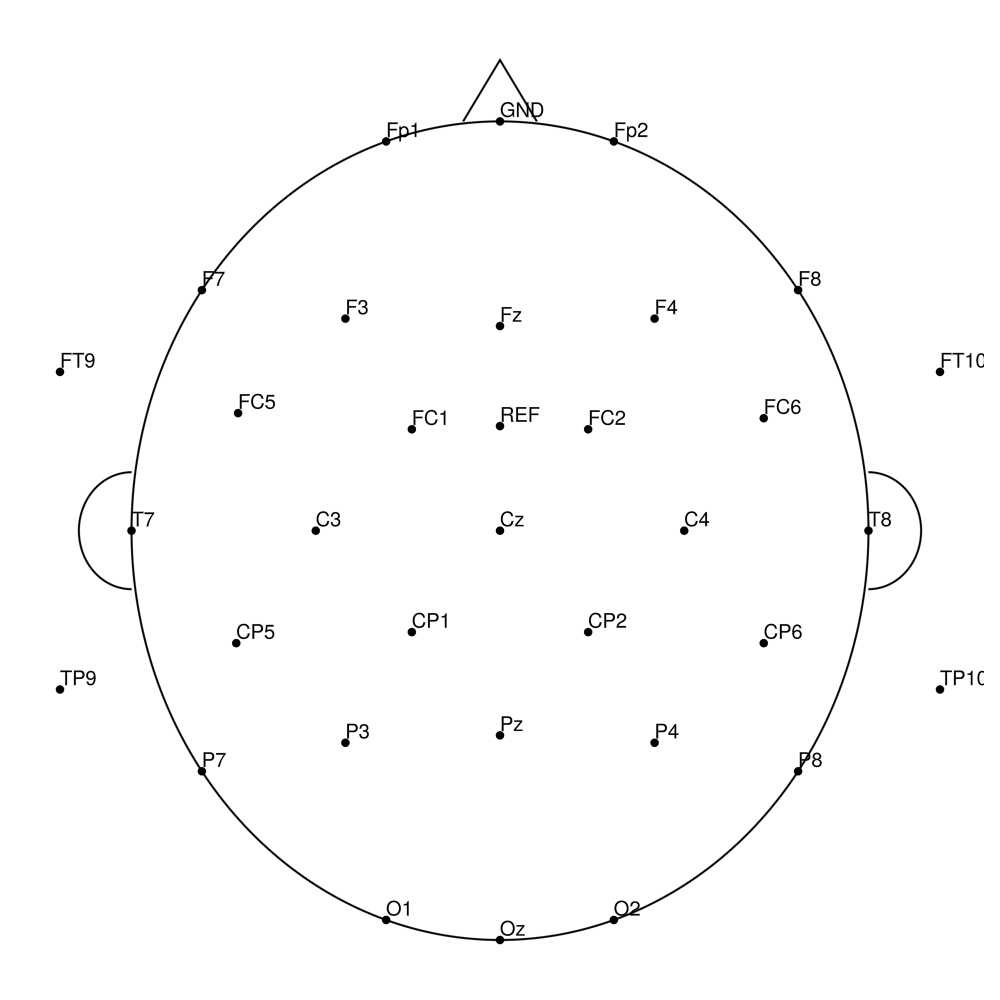
Twist-32 (32 channels) - for actiCHamp (no REF)
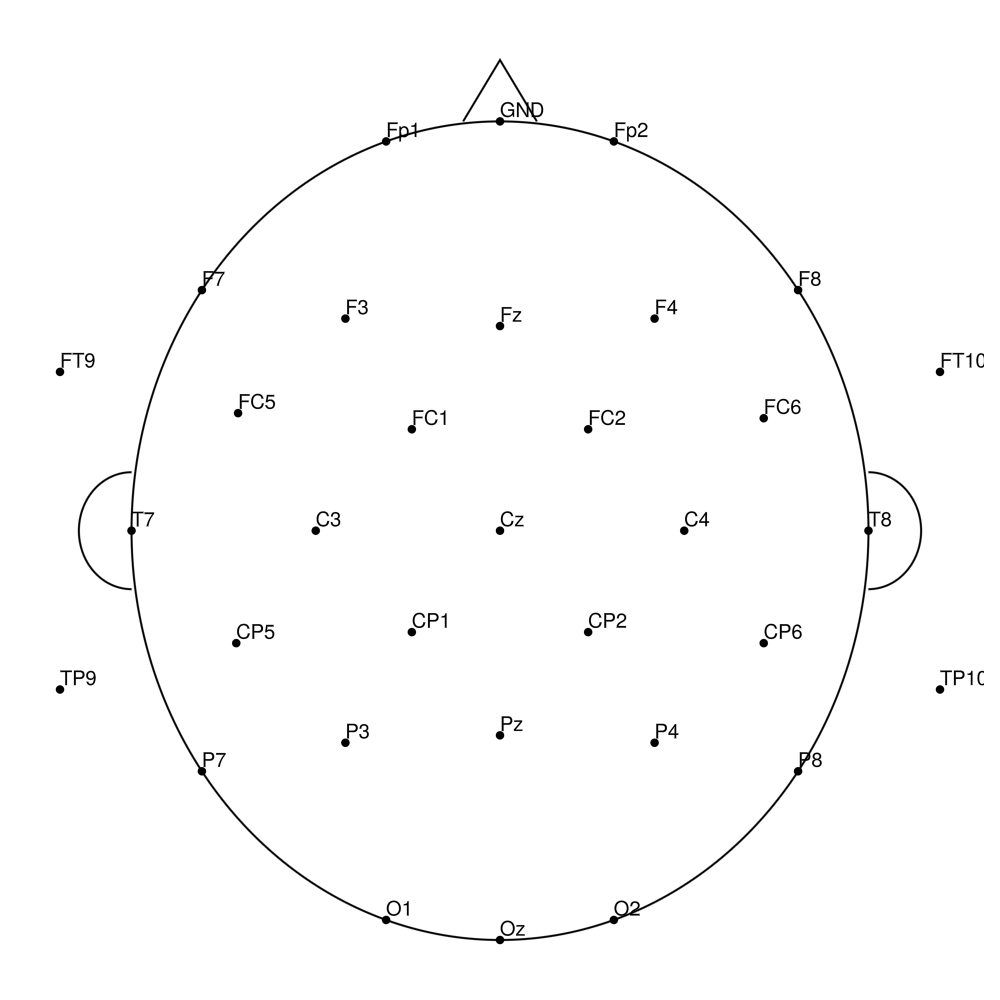
actiCHamp Electrode Cap (CAP)
AP-32 (32 channels) - actiCHamp electrode cap
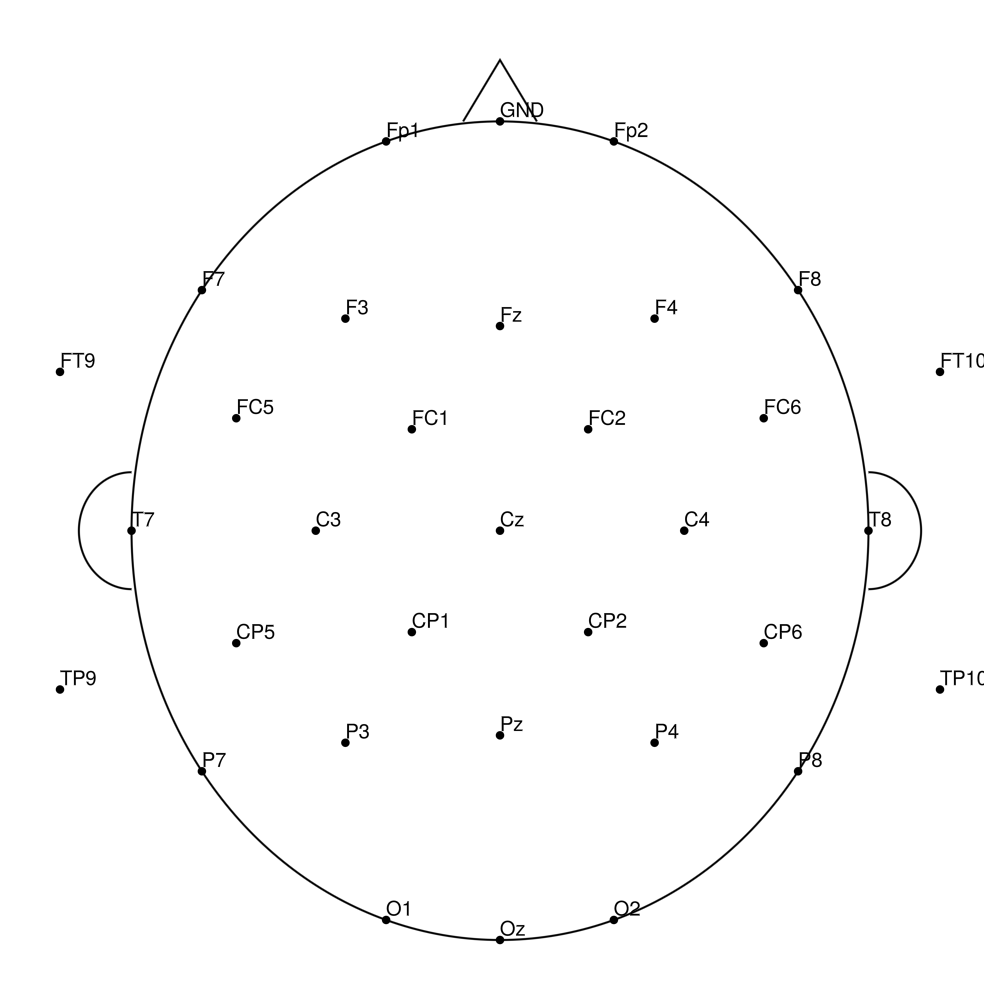
AP-64 (64 channels) - actiCHamp electrode cap
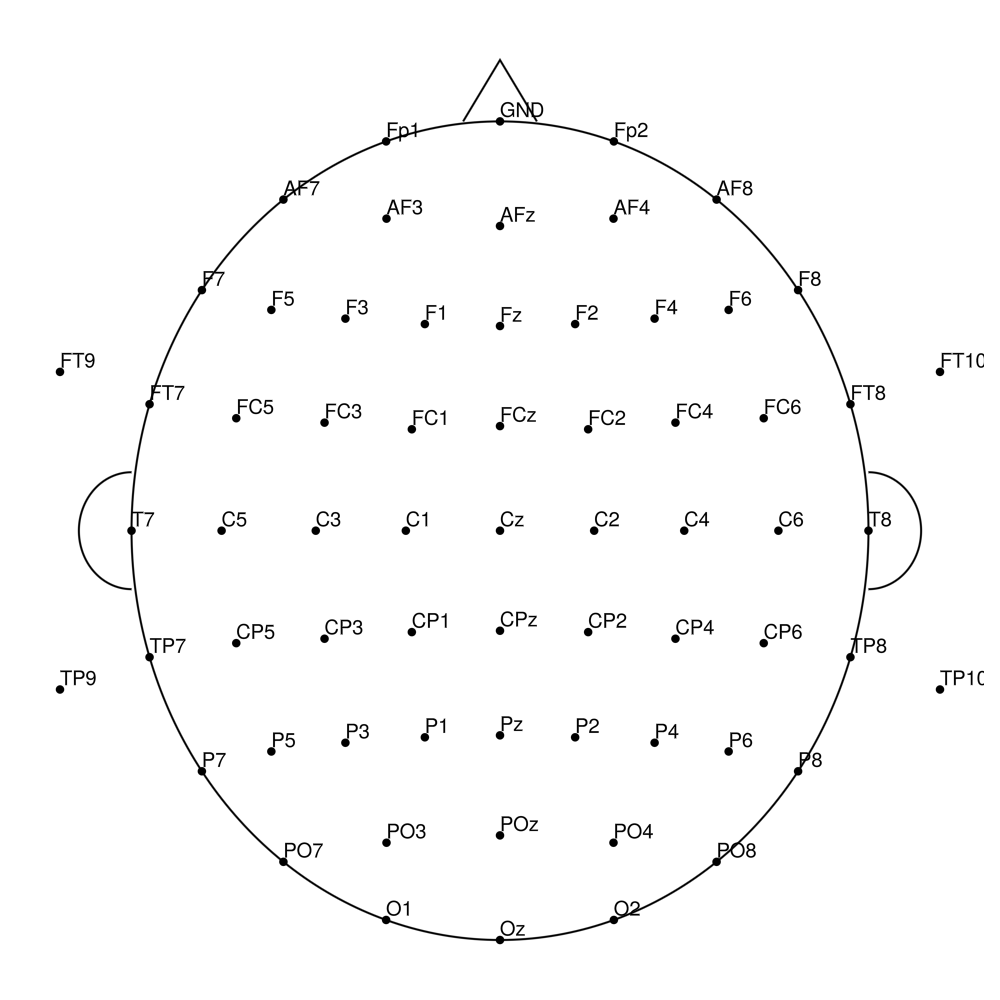
AP-96 (96 channels) - actiCHamp electrode cap
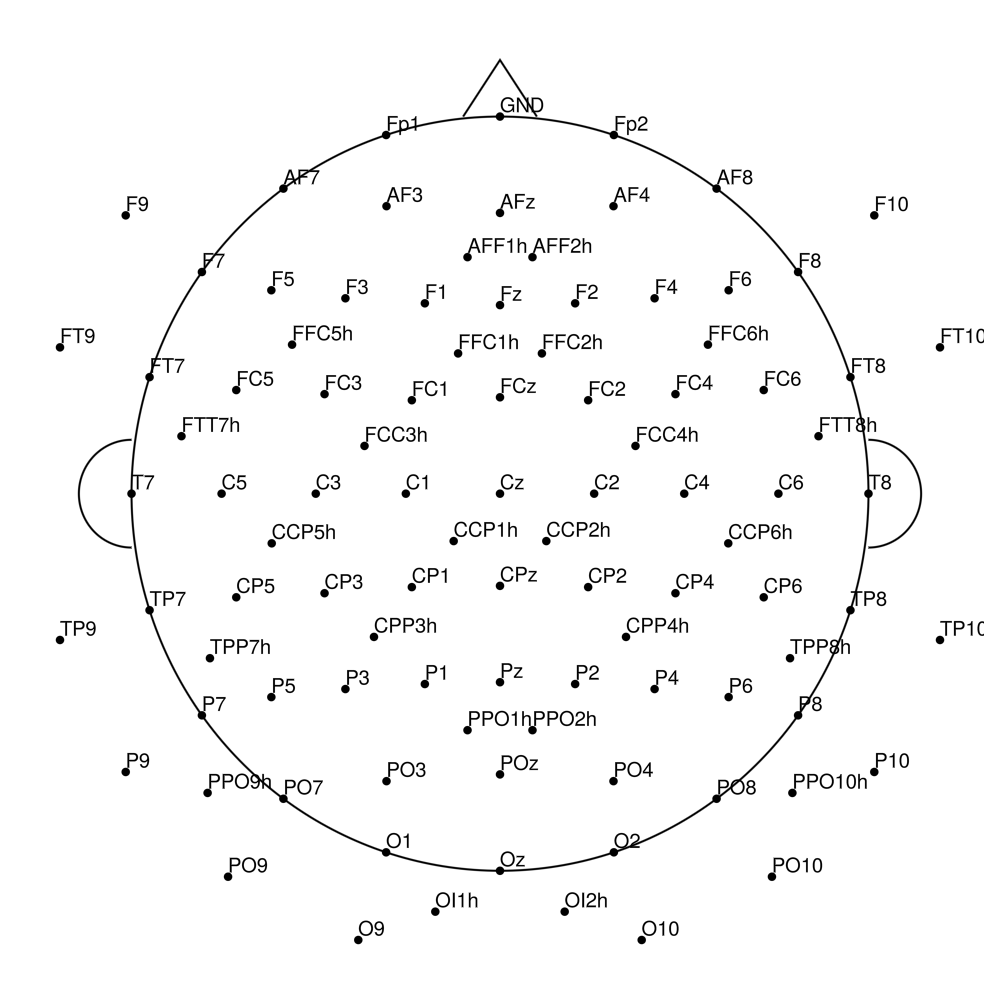
AP-128 (128 channels) - actiCHamp electrode cap
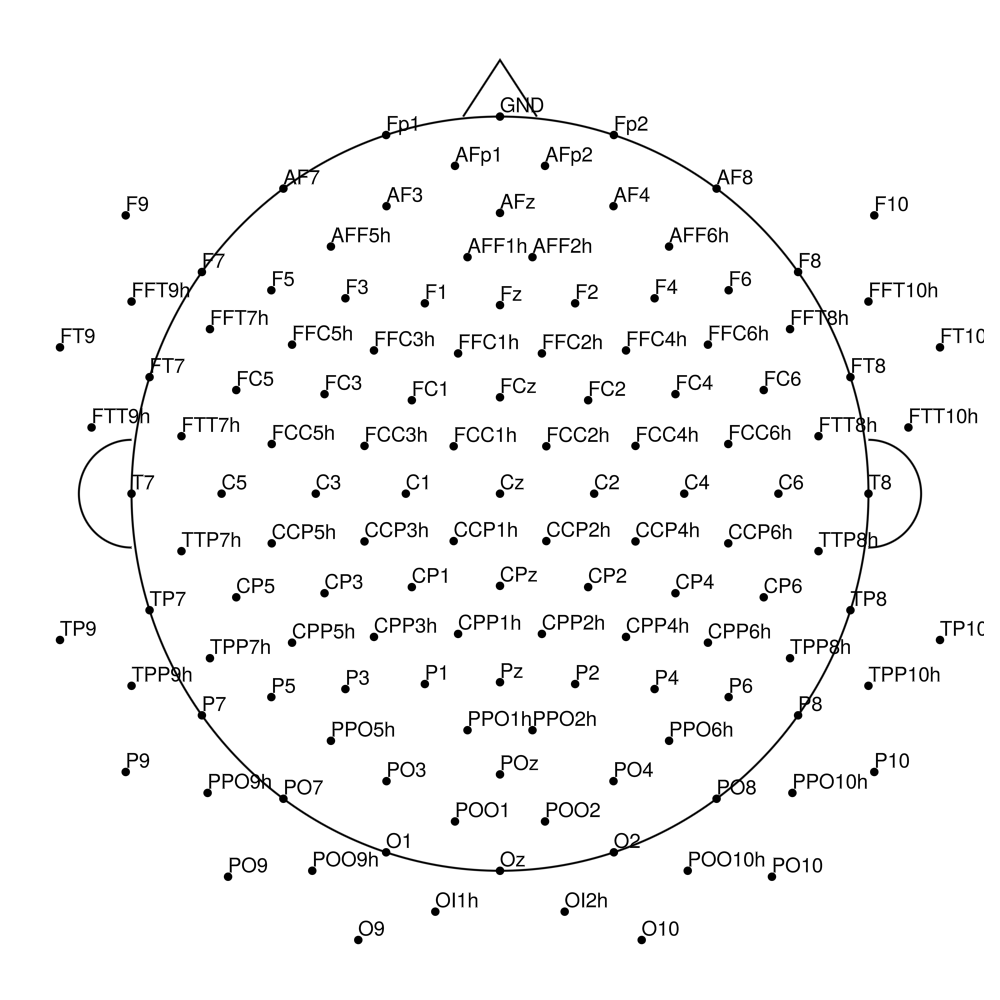
AP-160 (160 channels) - actiCHamp electrode cap
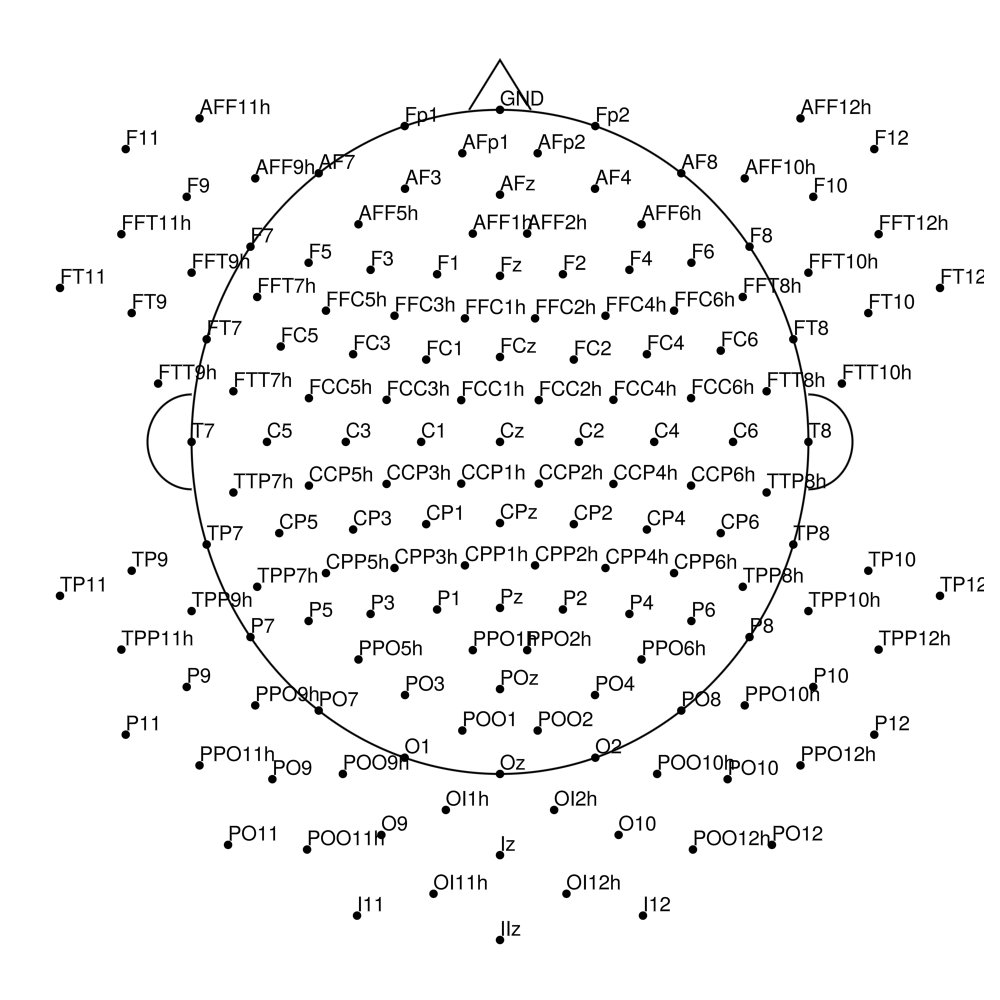
actiCAP UOL (CMA)
CMA-21 (21 channels) - BrainAmp (with REF)
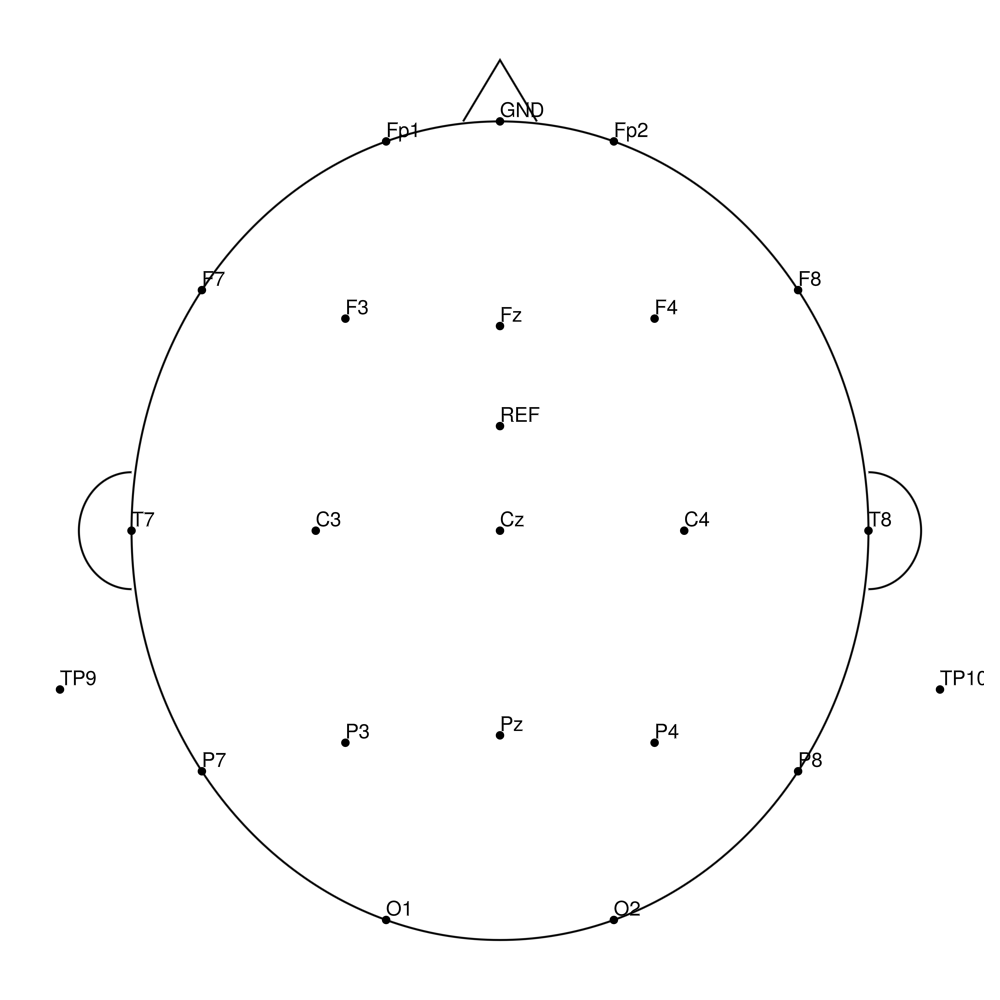
CMA-21 (21 channels) - actiCHamp (no REF)
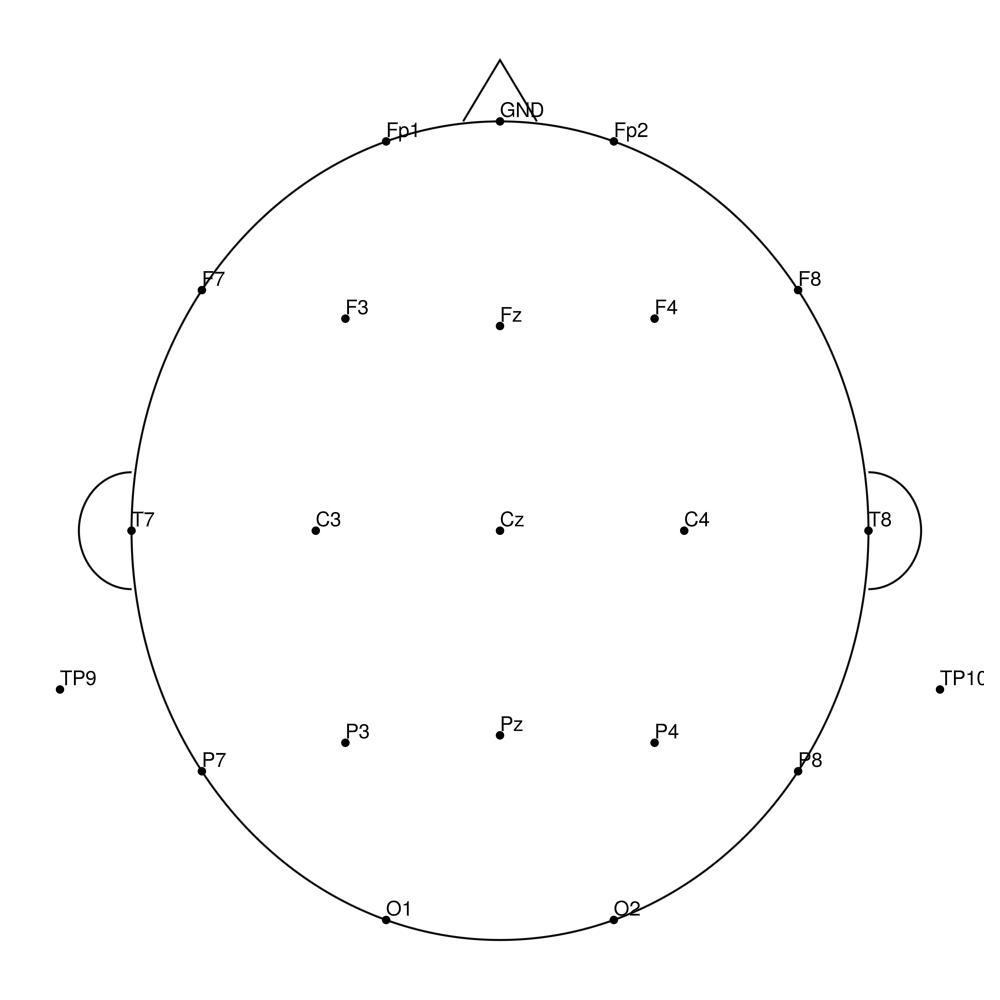
CMA-21 (21 channels) - actiCHamp Plus (no REF)
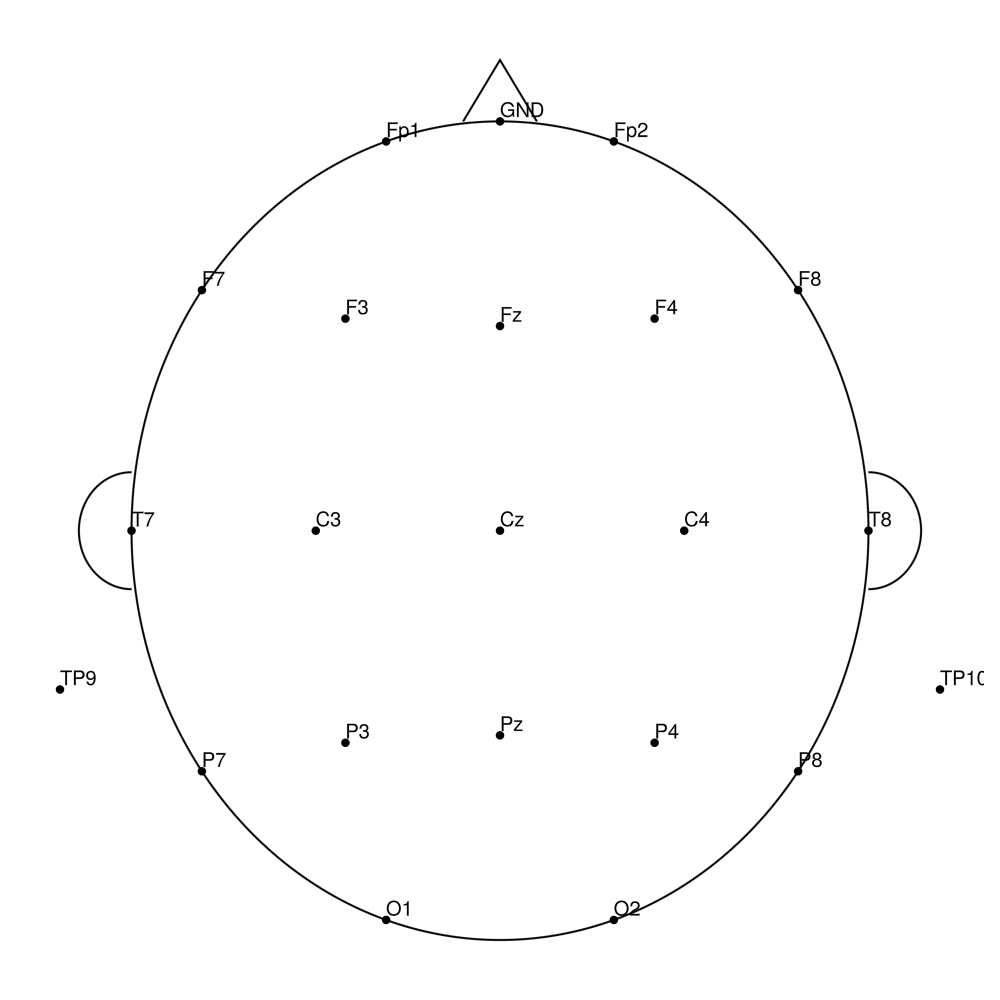
CMA-32 (32 channels) - BrainAmp (with REF)
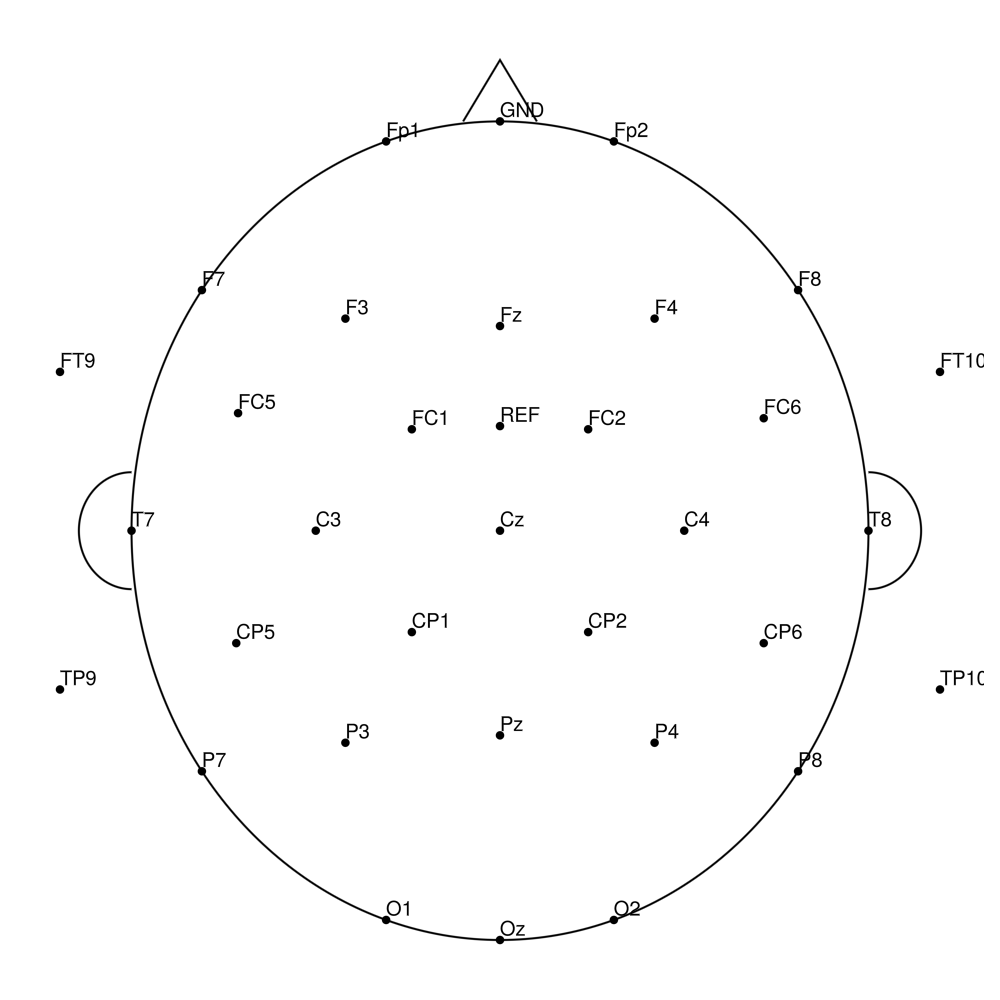
CMA-32 (32 channels) - actiCHamp (no REF)
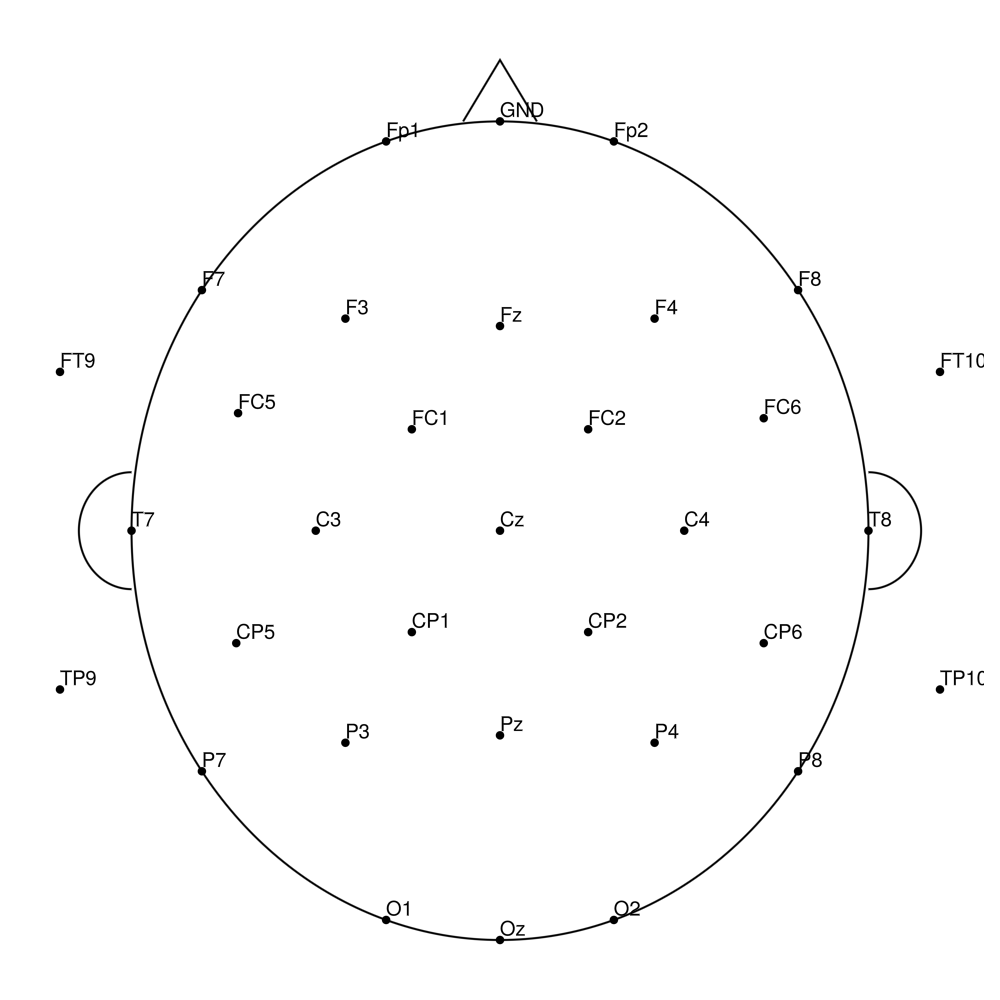
CMA-32 (32 channels) - actiCHamp Plus (no REF)
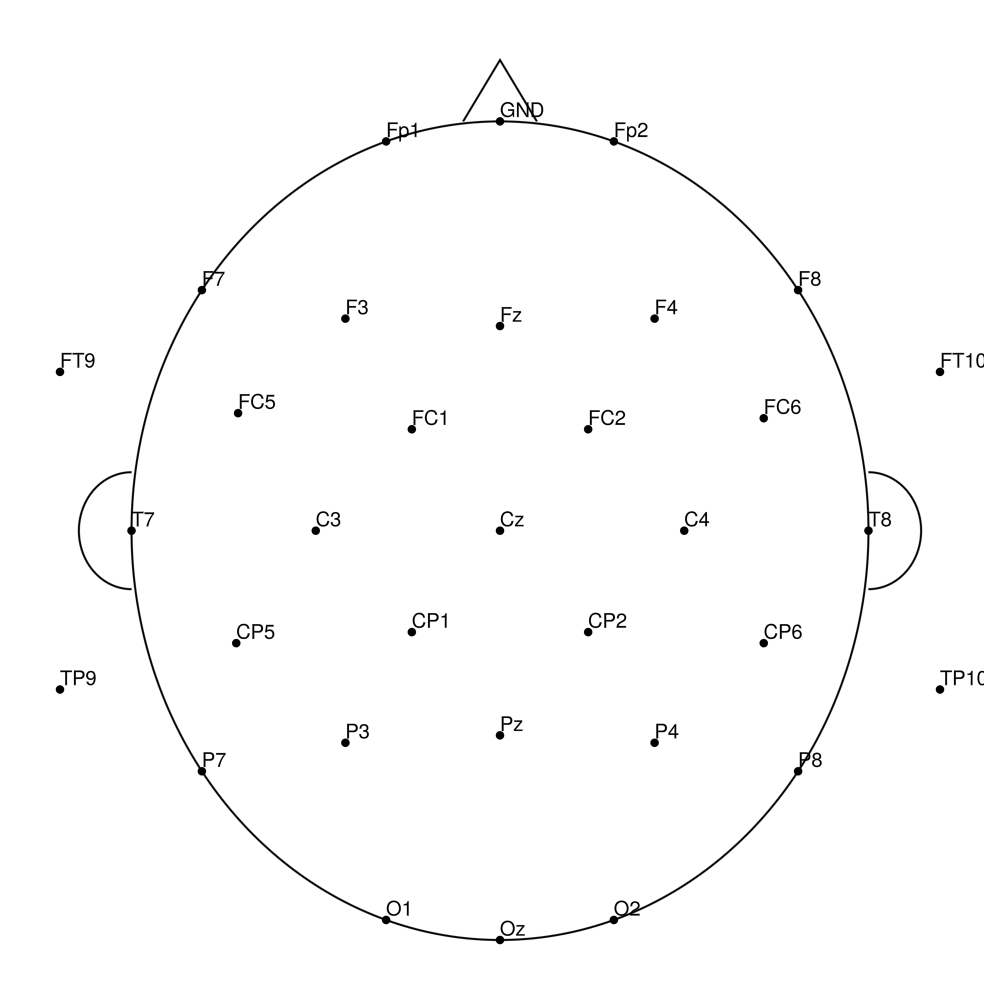
CMA-64 (64 channels) - BrainAmp (with REF)
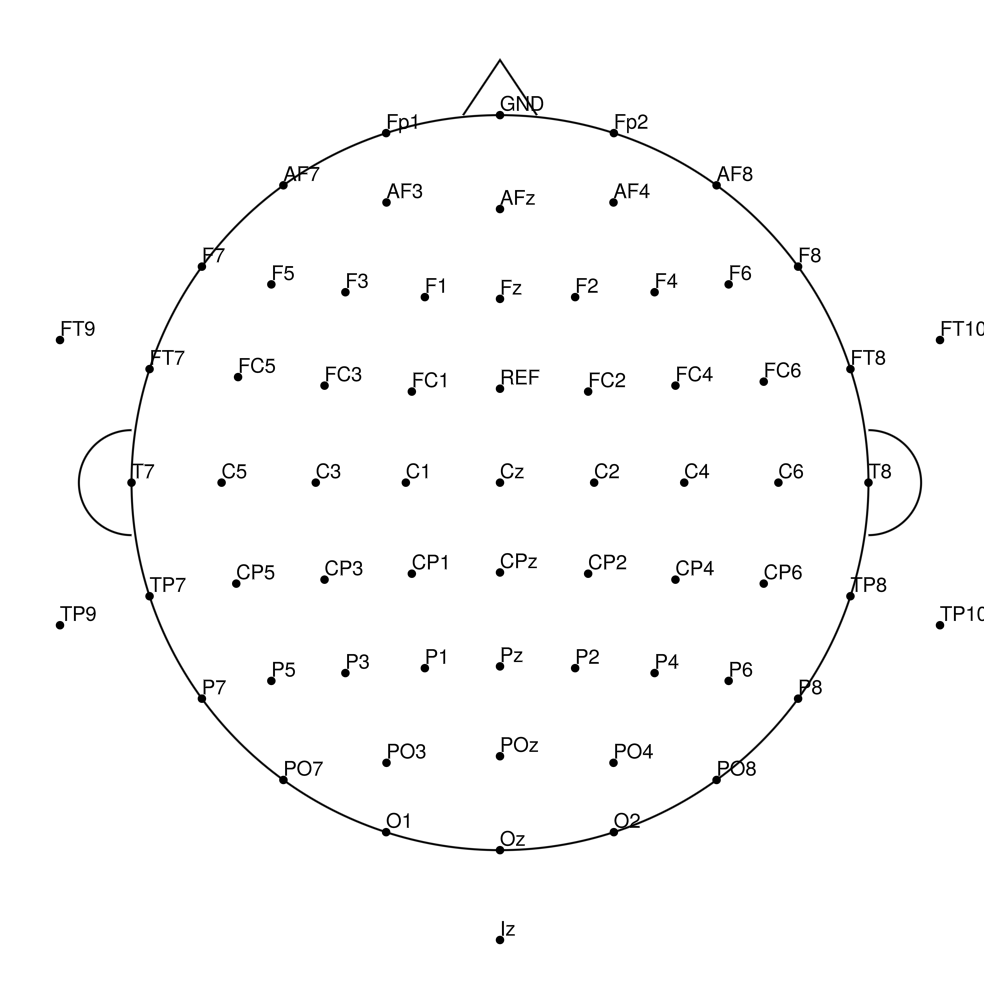
CMA-64 (64 channels) - actiCHamp (no REF)
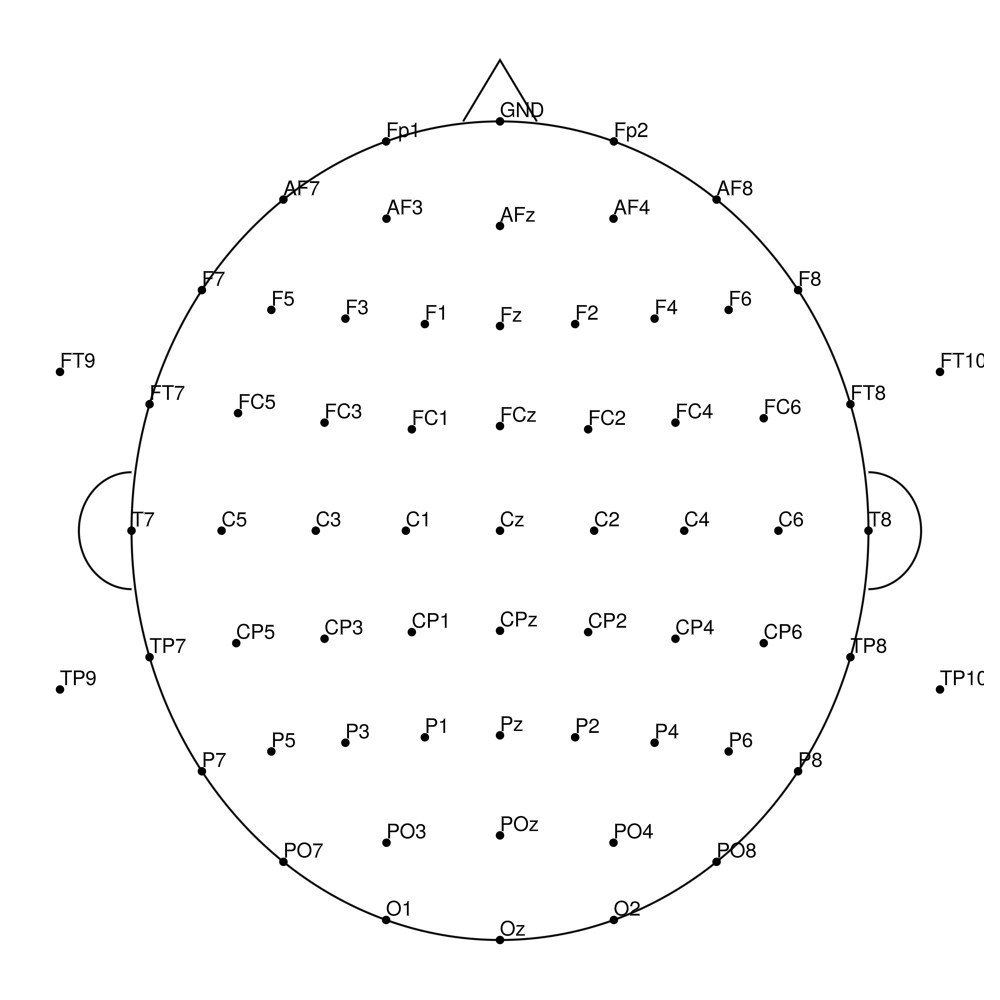
CMA-64 (64 channels) - actiCHamp Plus (no REF)
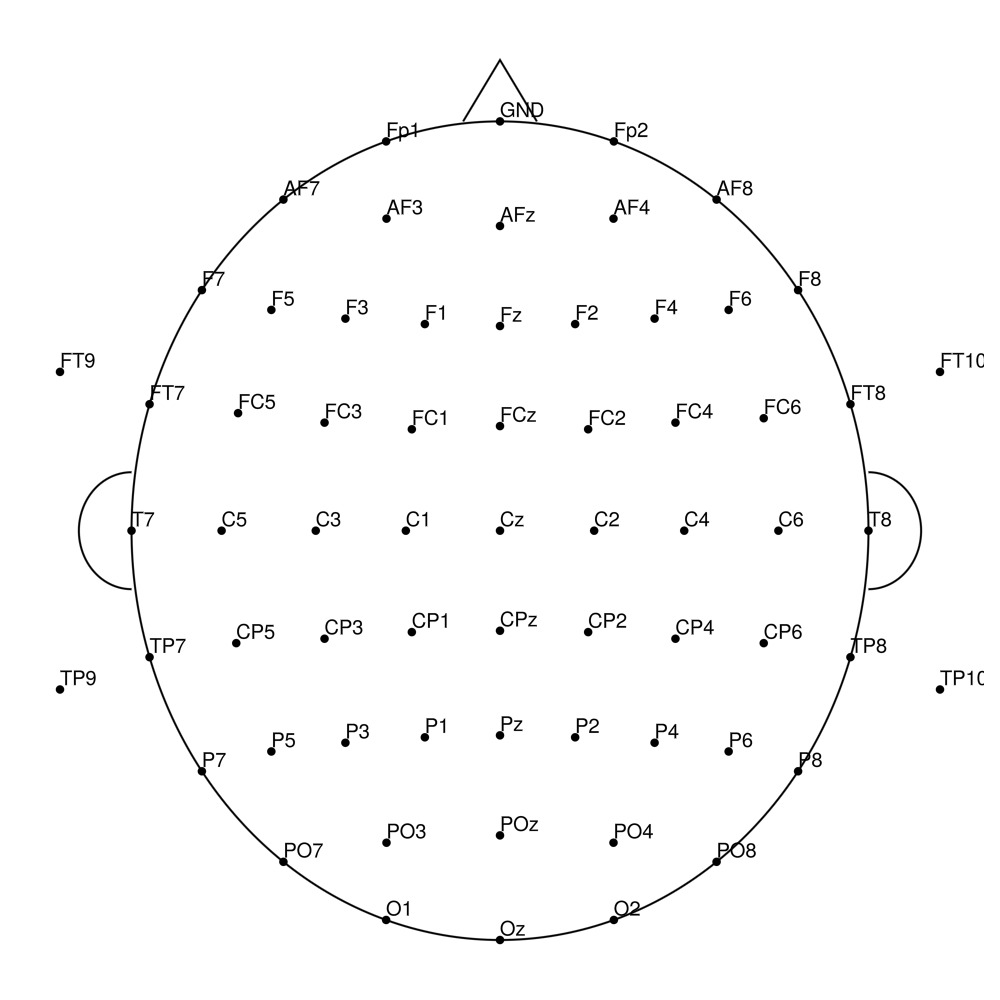
CMA-96 (96 channels) - BrainAmp (with REF)
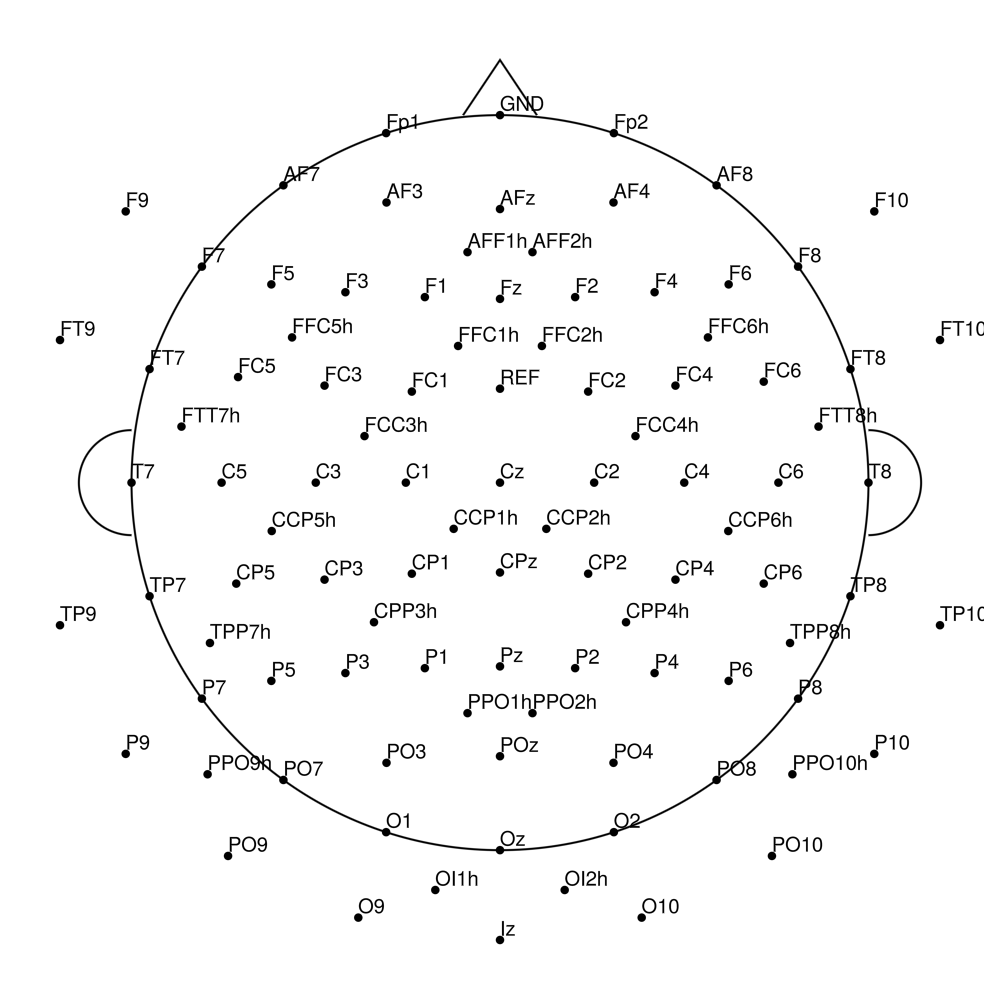
CMA-96 (96 channels) - actiCHamp (no REF)
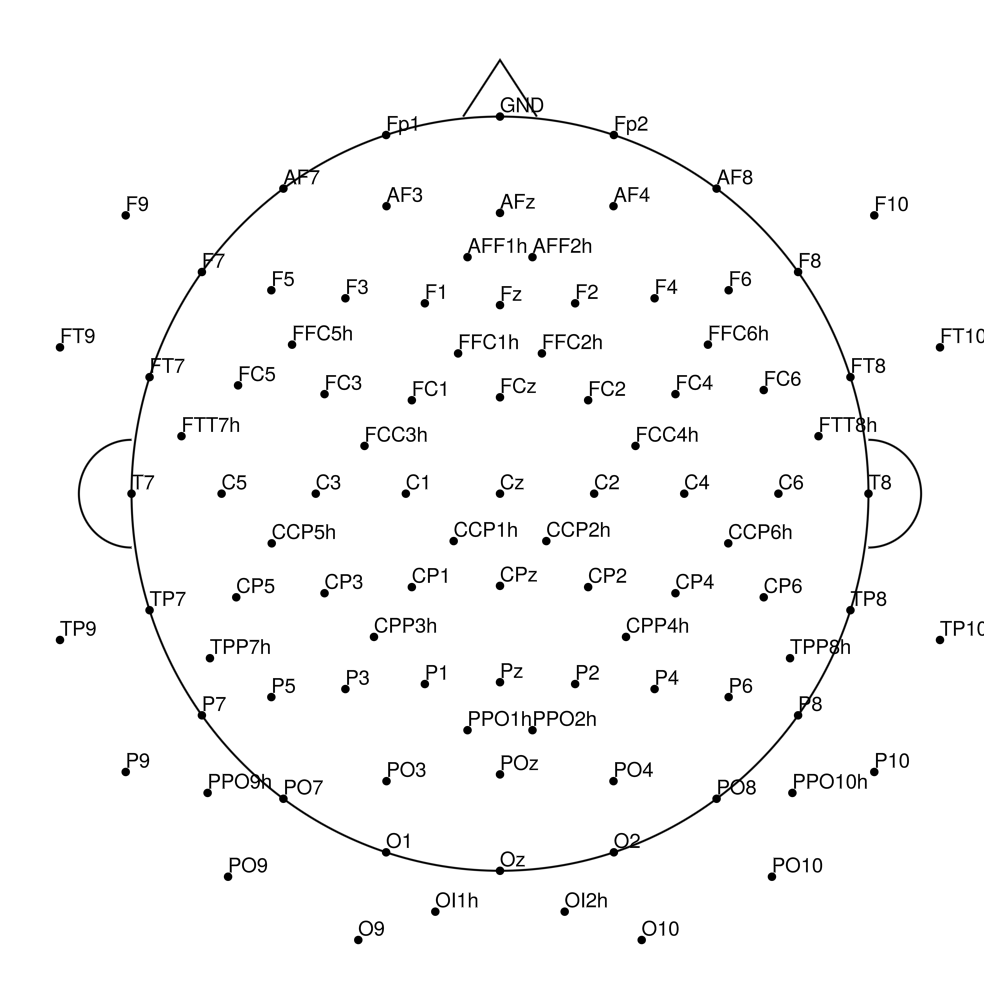
CMA-96 (96 channels) - actiCHamp Plus (no REF)
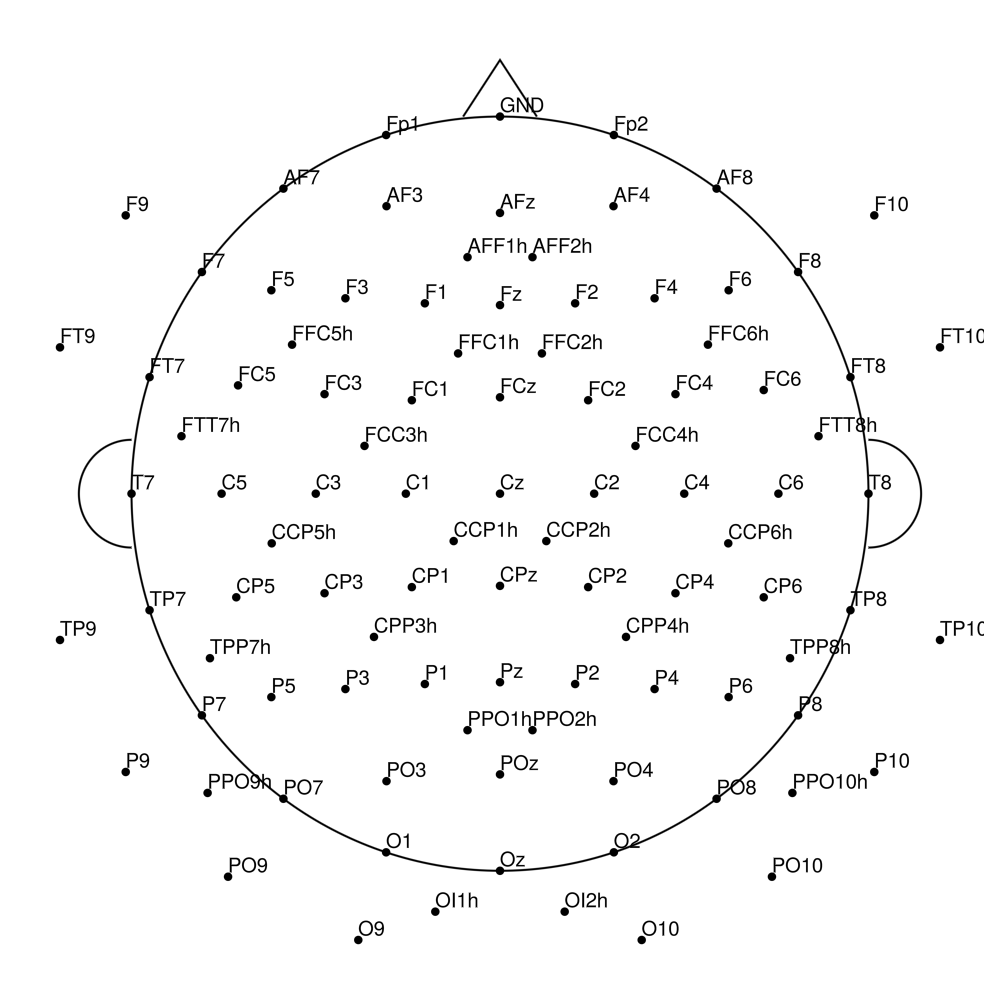
CMA-128 (128 channels) - BrainAmp (with REF)
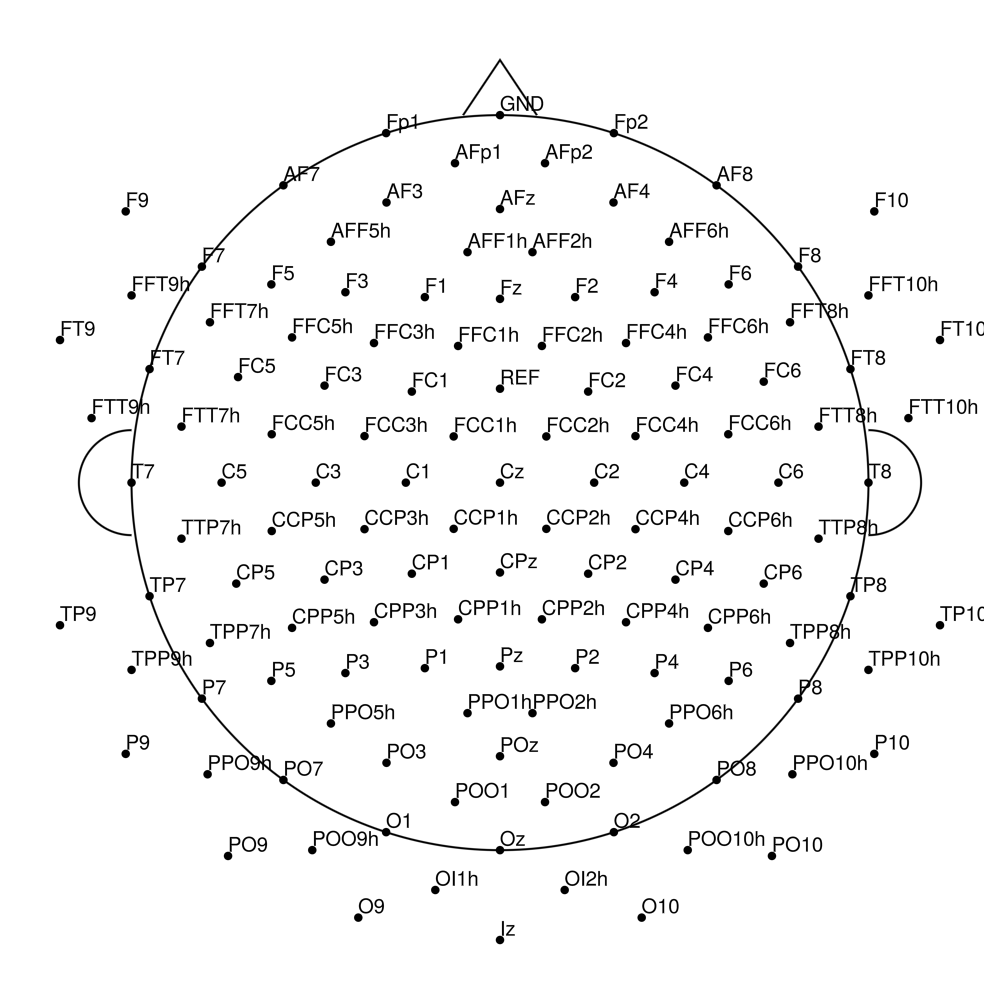
CMA-128 (128 channels) - actiCHamp (no REF)
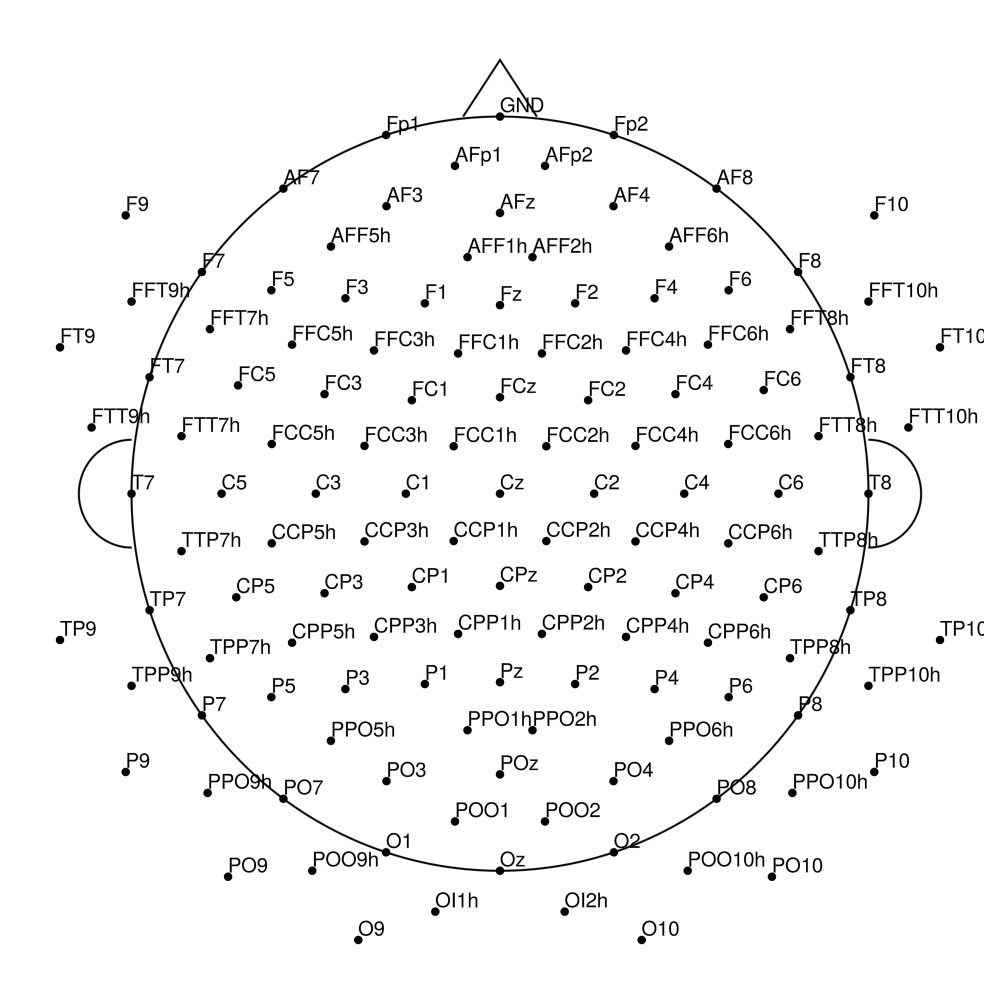
CMA-128 (128 channels) - actiCHamp Plus (no REF)
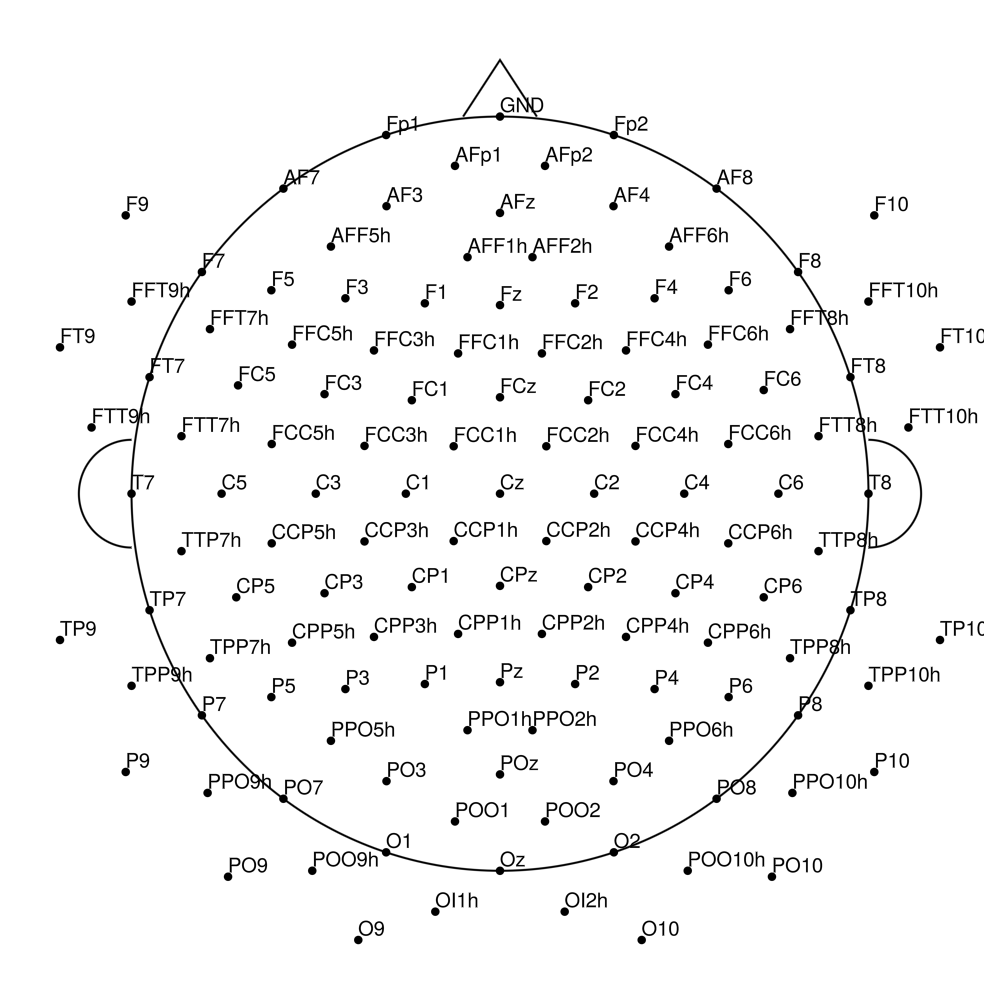
CMA-160 (160 channels) - BrainAmp (with REF)
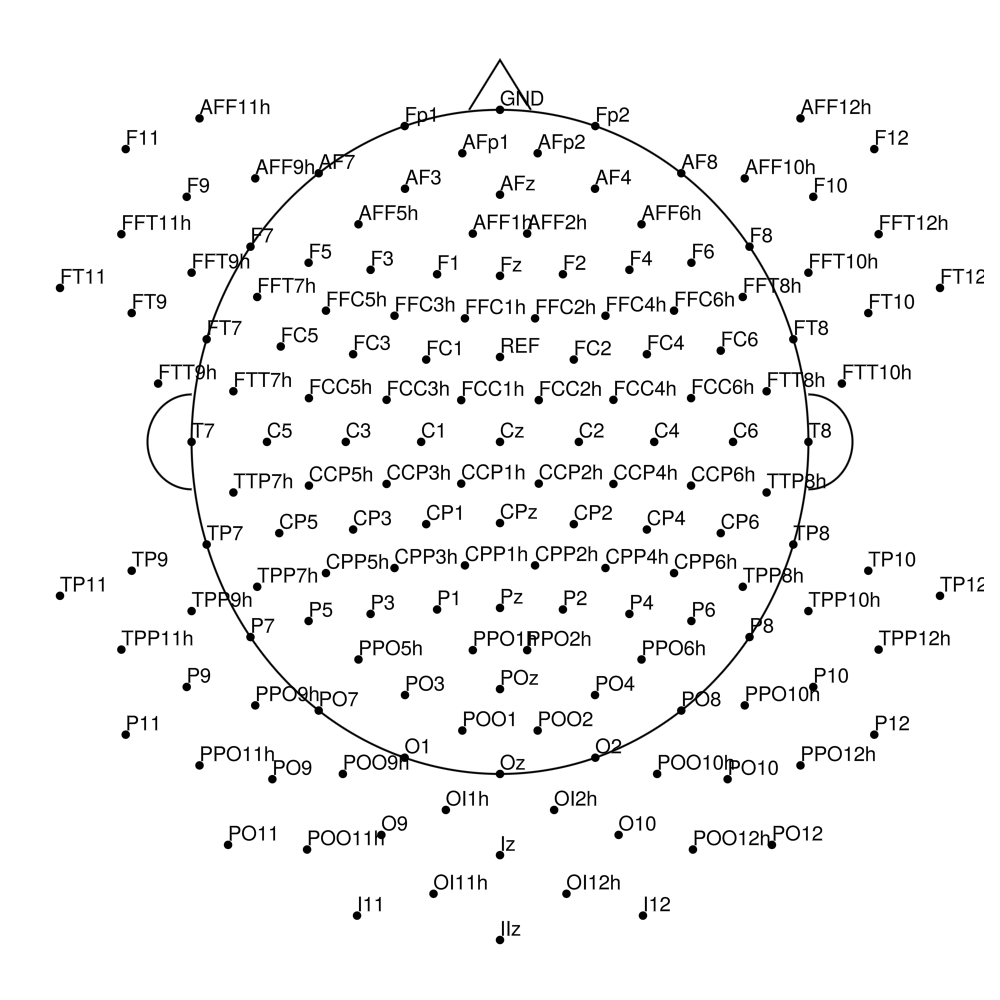
CMA-160 (160 channels) - actiCHamp (no REF)
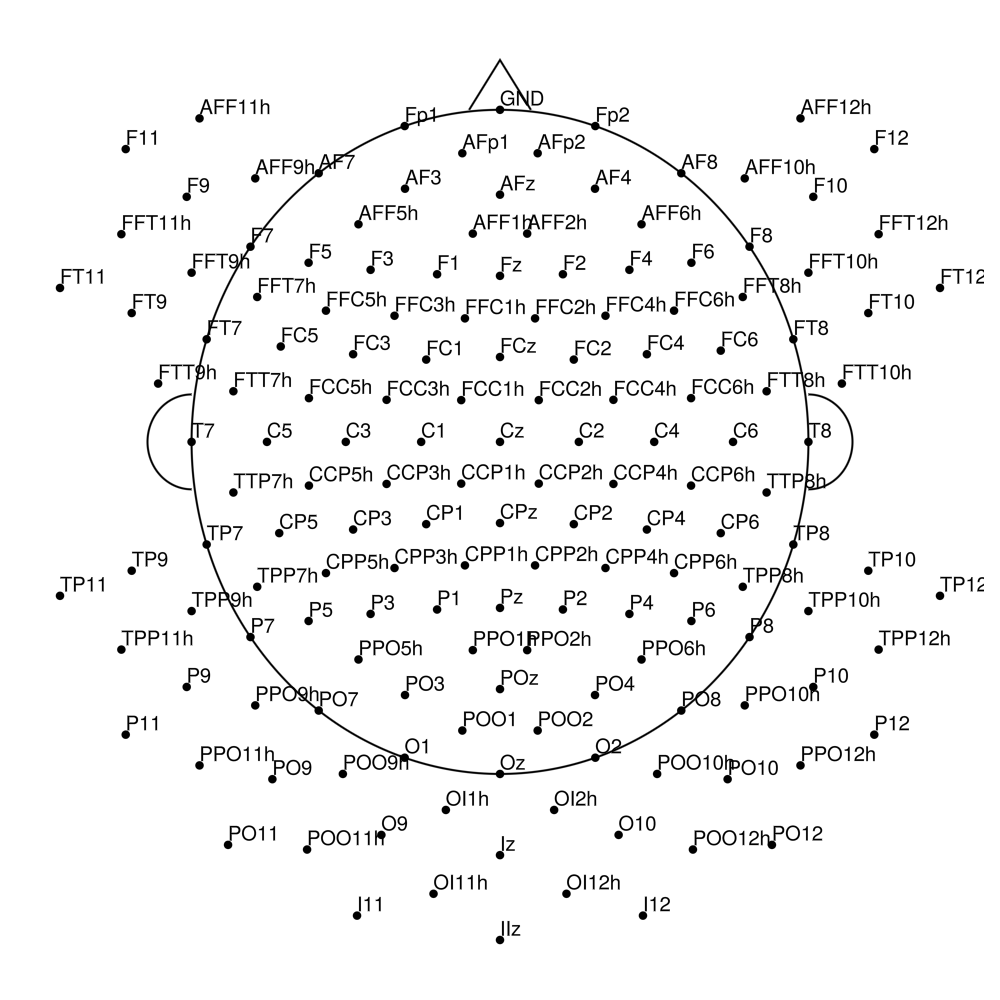
CMA-160 (160 channels) - actiCHamp Plus (no REF)
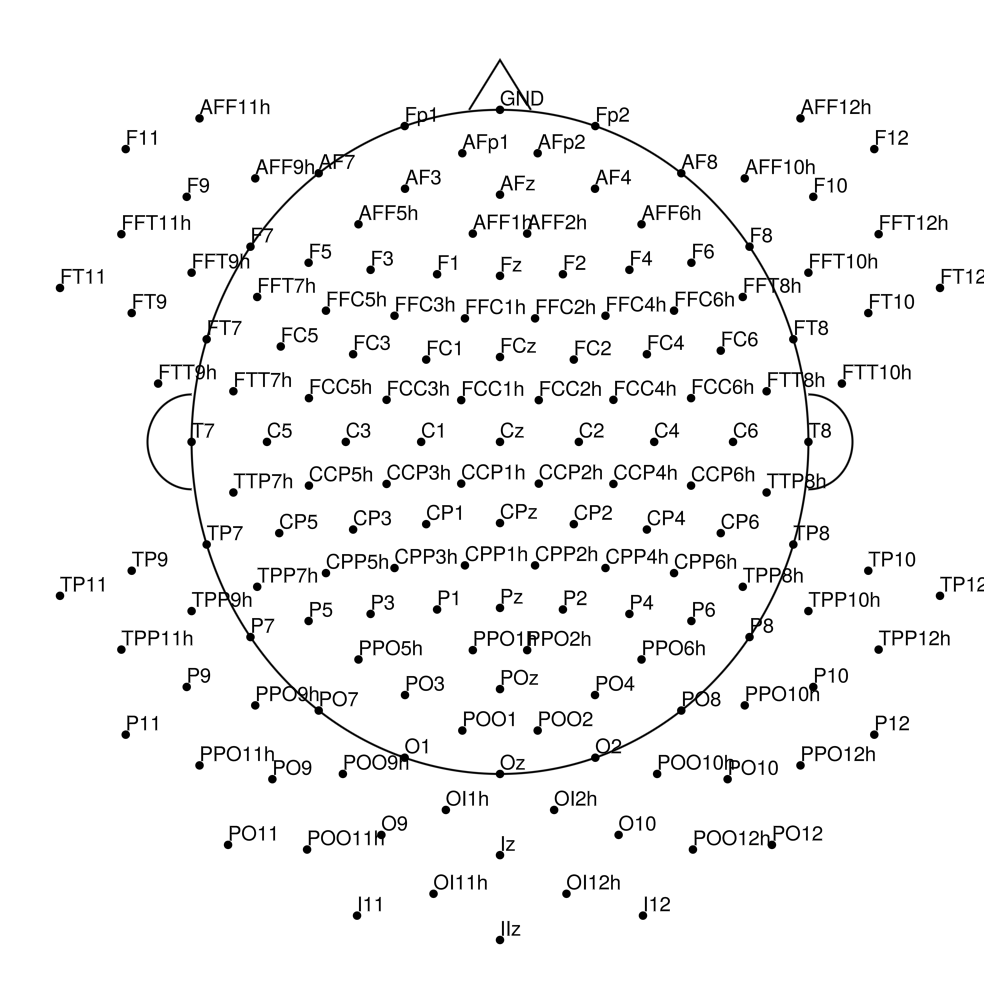
actiCAP Slim (AS)
AS-32 (32 channels) - with REF
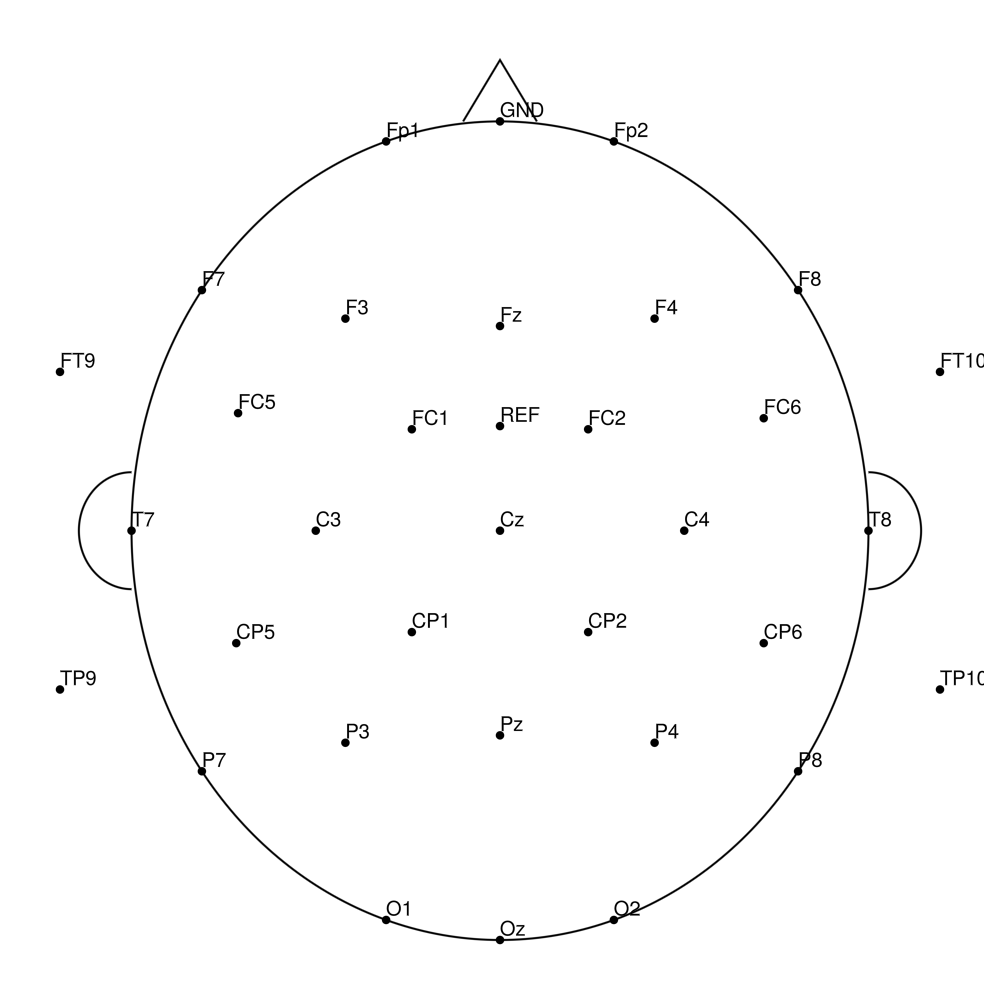
AS-32 (32 channels) - no REF
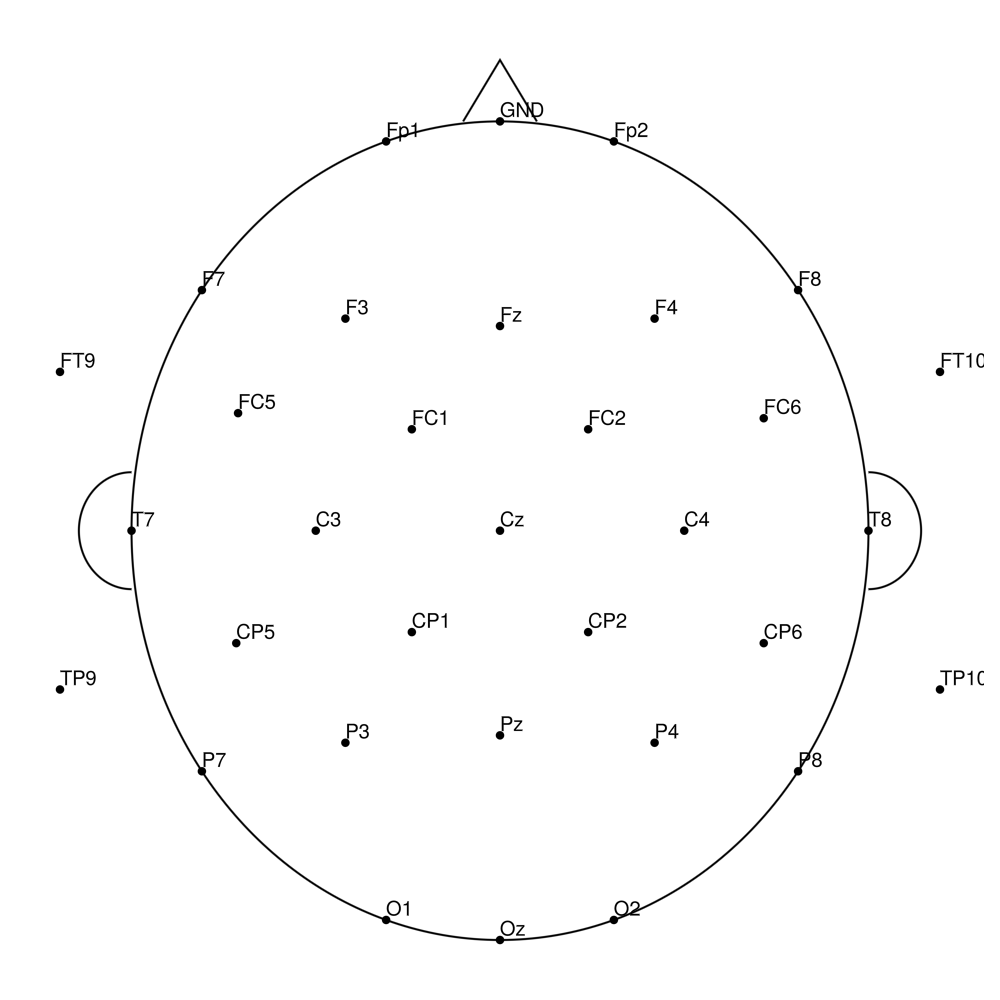
AS-64 (64 channels) - with REF
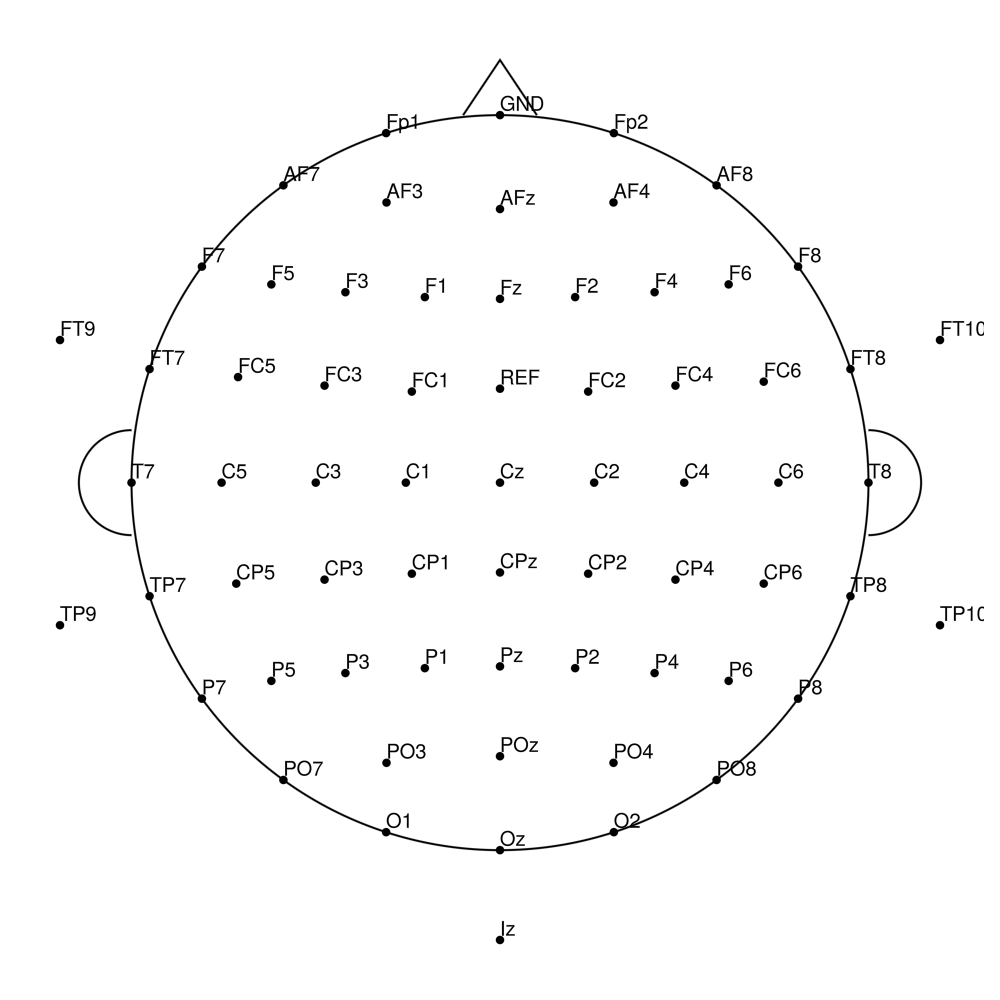
AS-64 (64 channels) - no REF
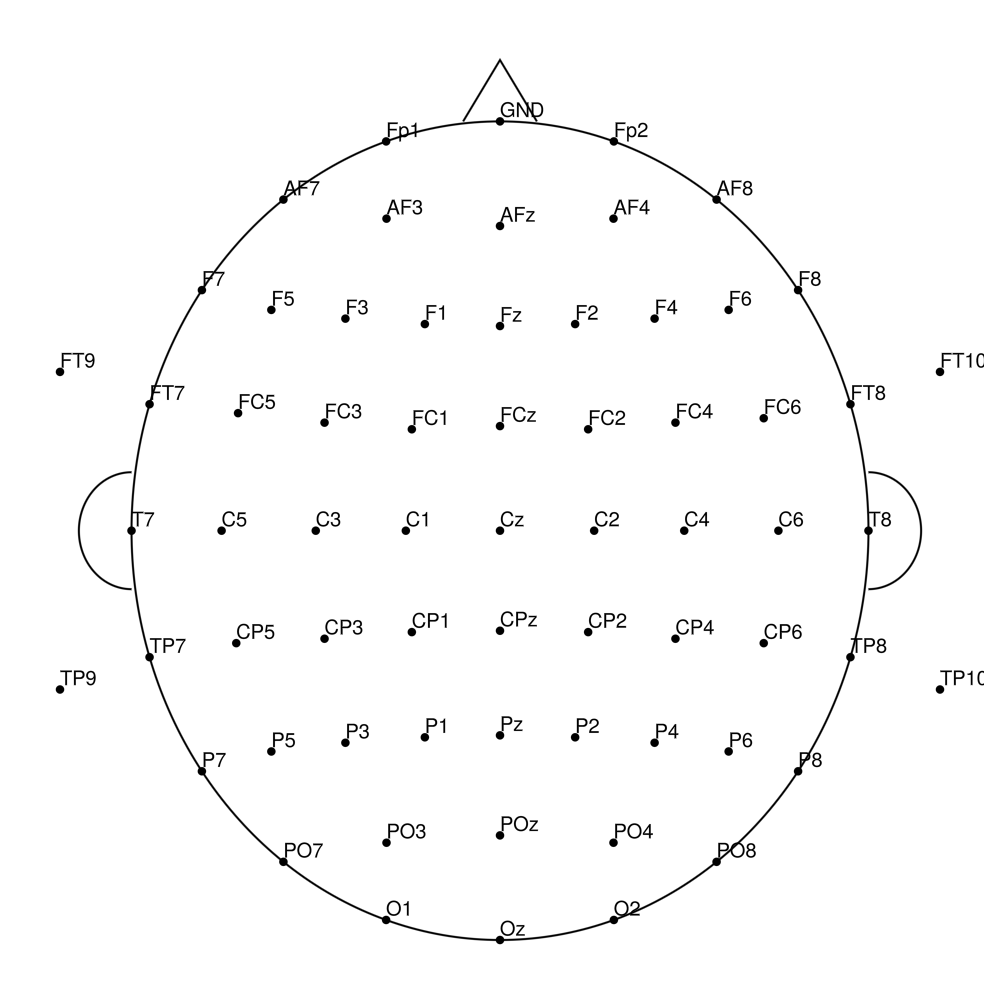
AS-96 (96 channels) - with REF
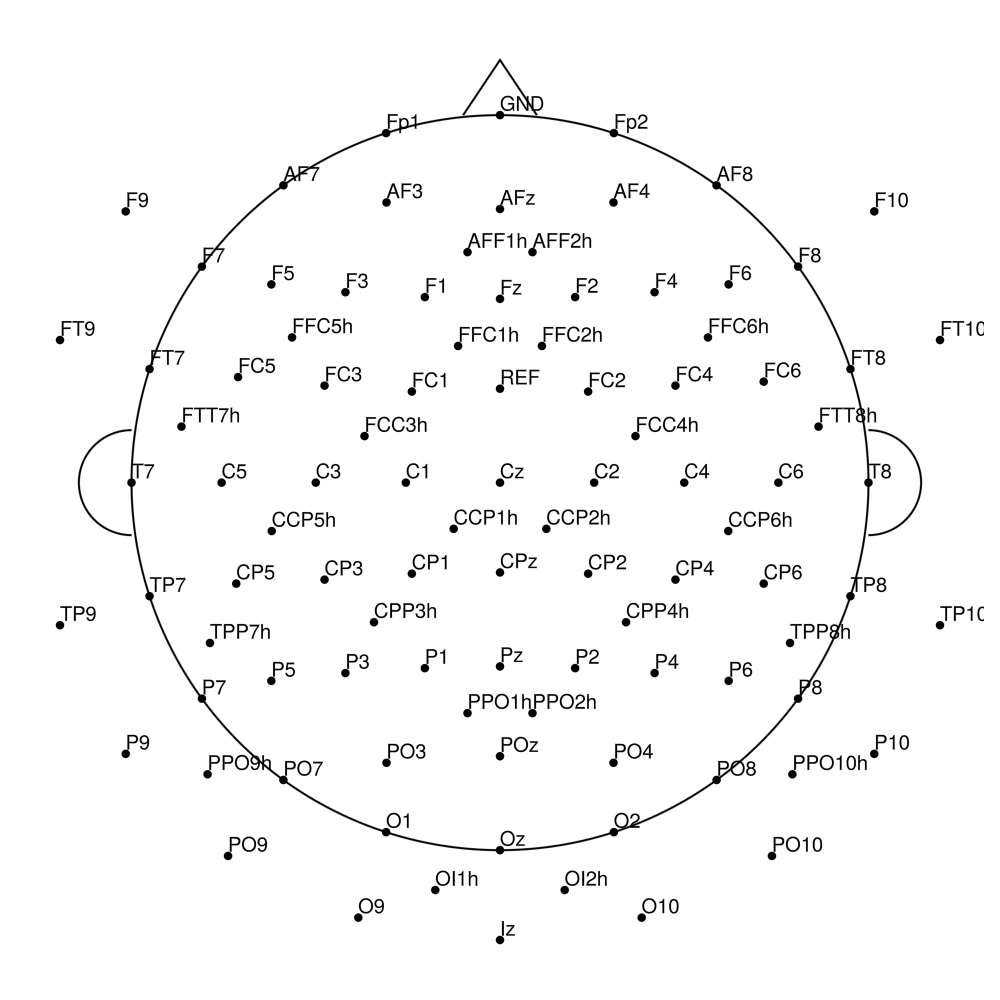
AS-96 (96 channels) - no REF
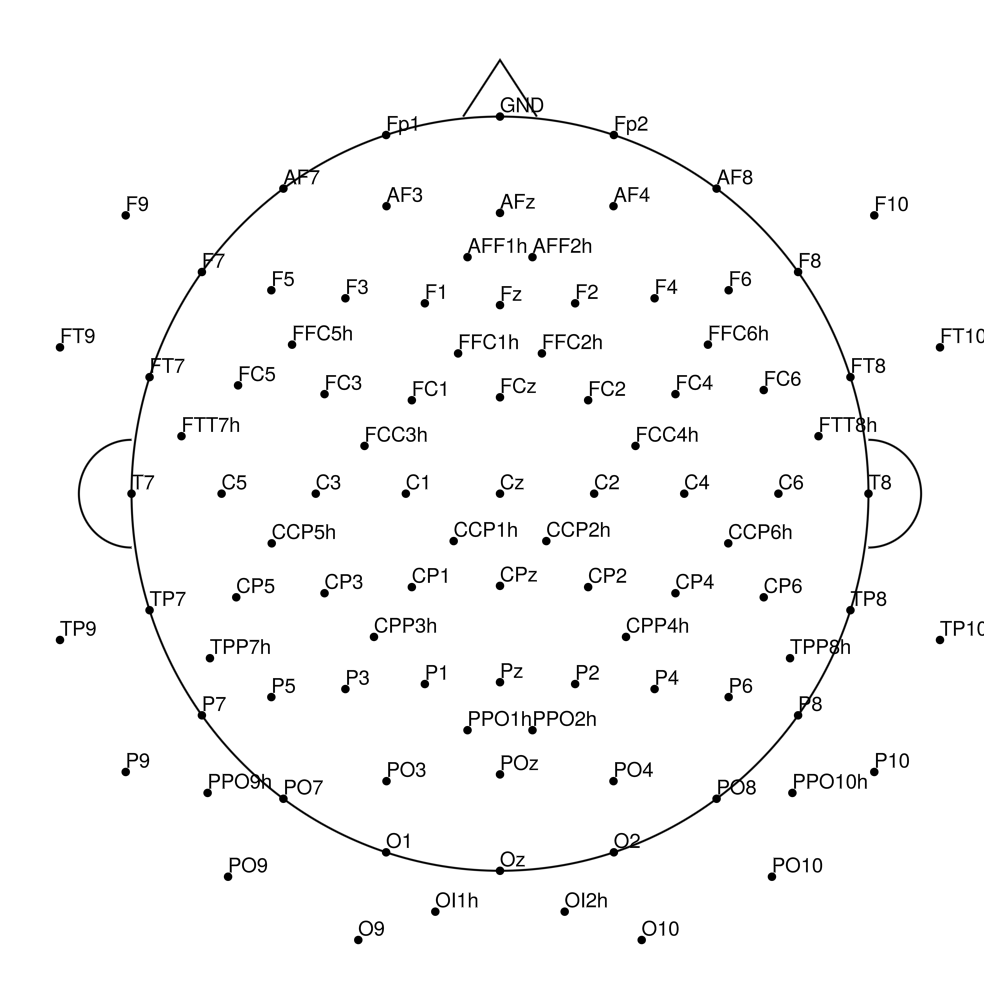
AS-128 (128 channels) - with REF
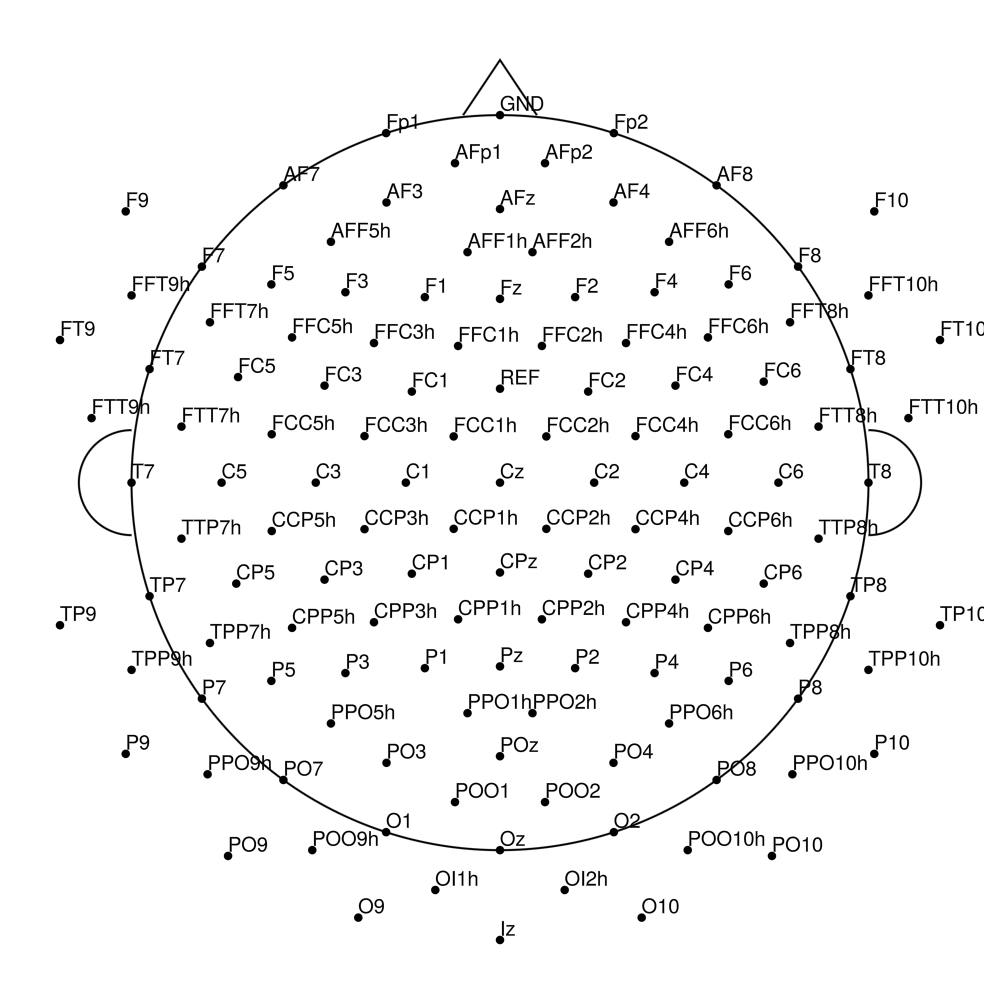
AS-128 (128 channels) - no REF
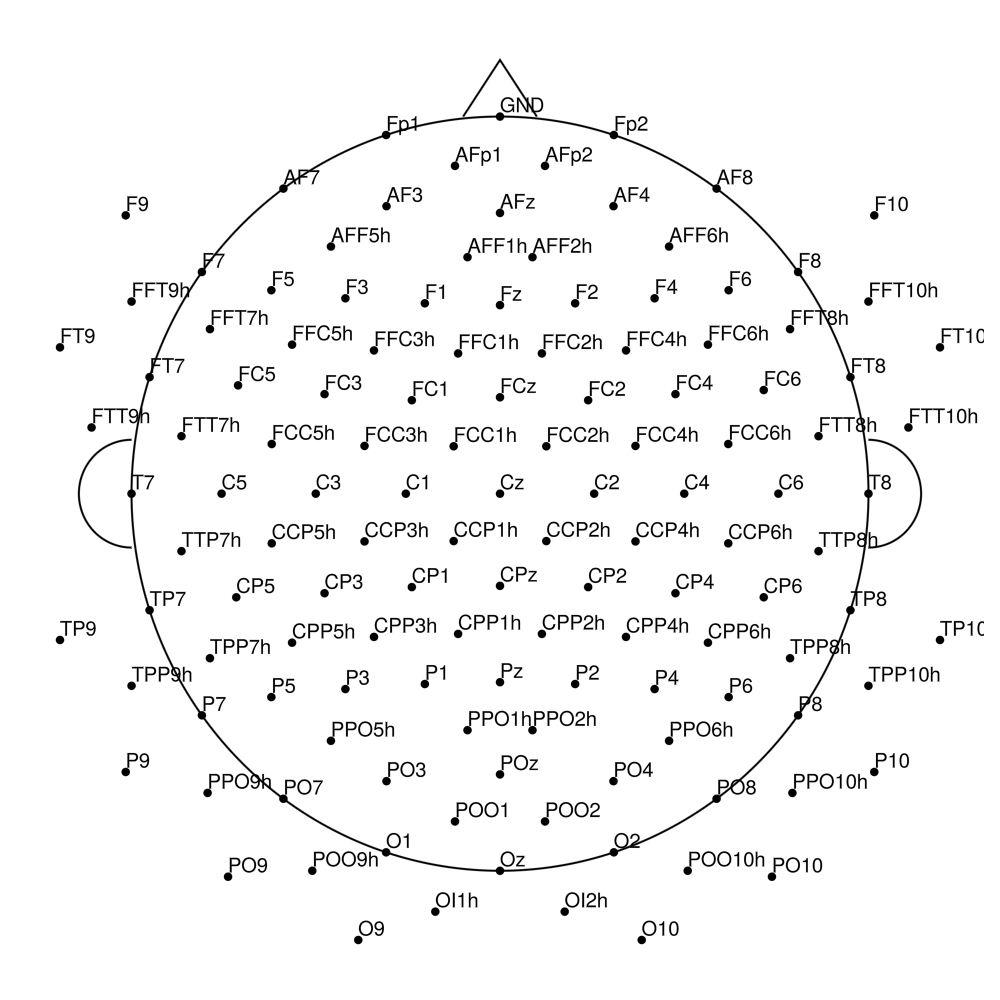
AS-160 (160 channels) - with REF
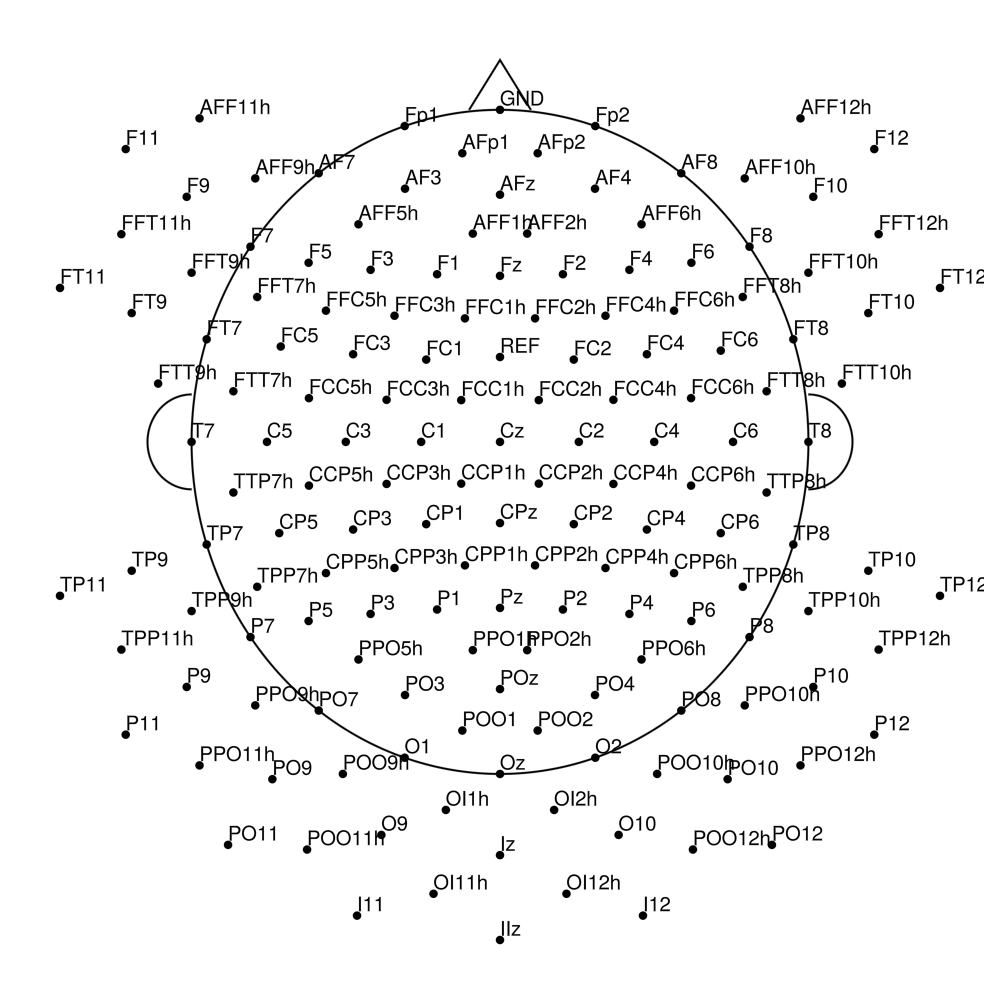
AS-160 (160 channels) - no REF
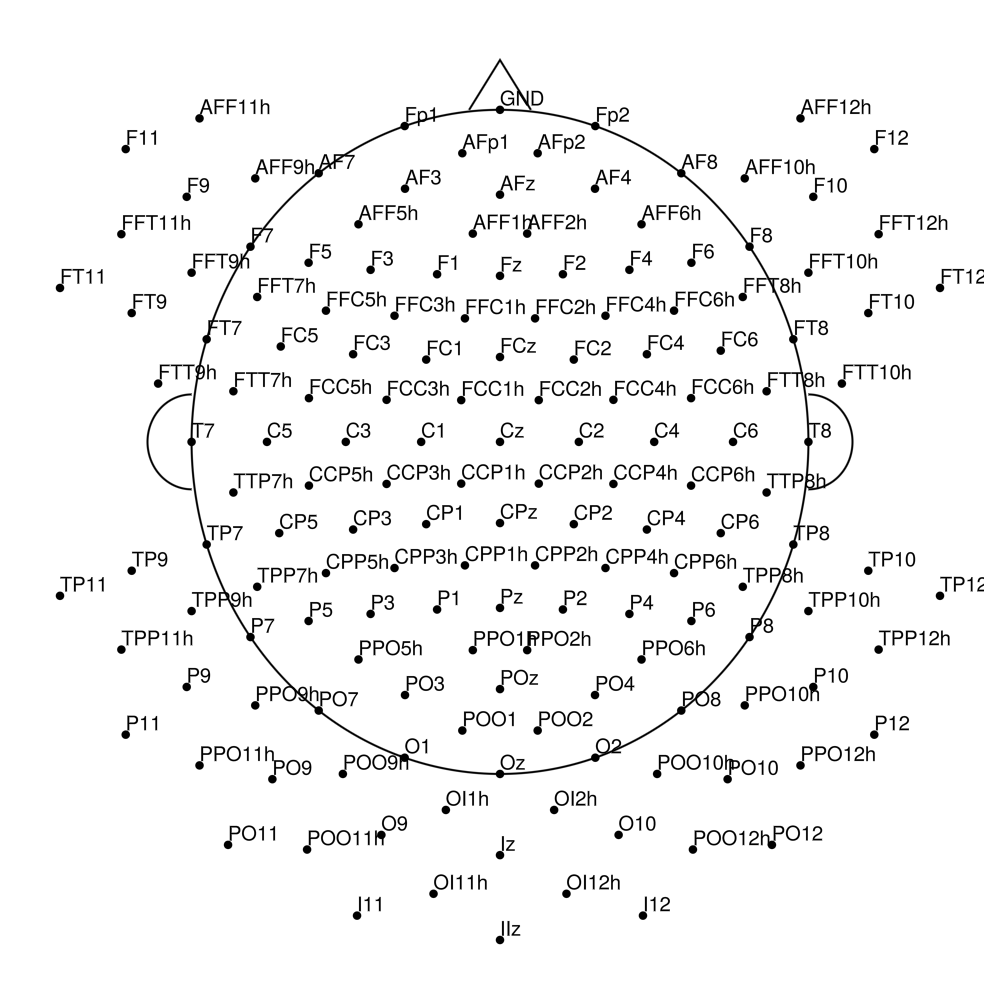
actiCap for LiveAmp
CLA-32 (32 channels) - actiCap for LiveAmp
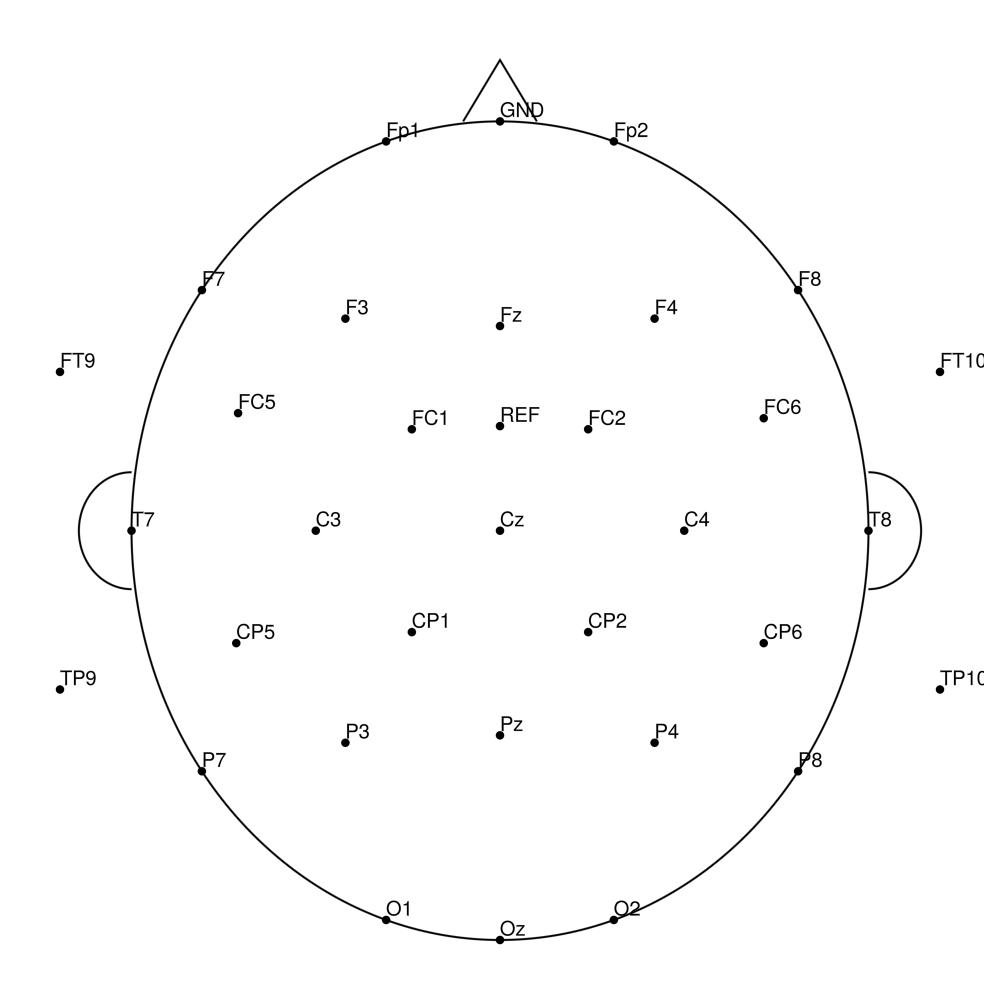
actiCap Slim (ASP) — for BrainAmpDC
CACS-32 (32 channels) - for BrainAmpDC (with REF)
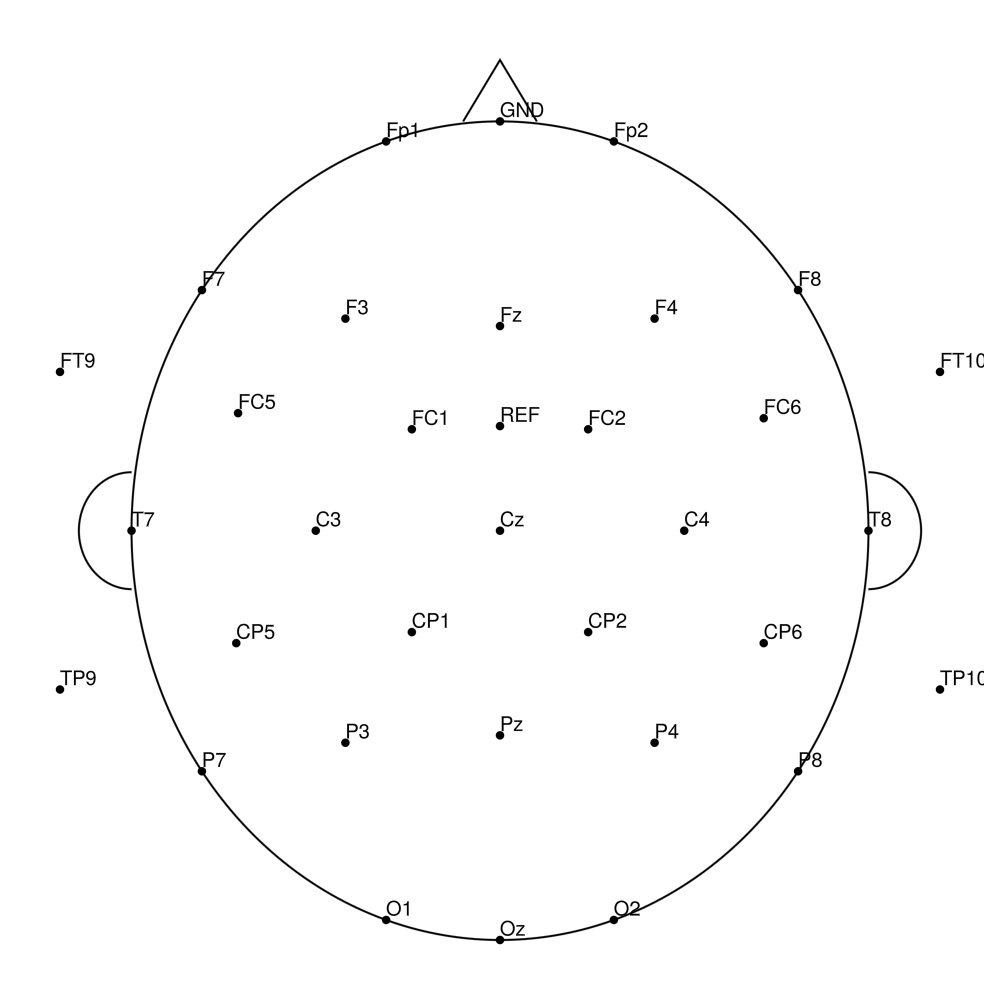
CACS-64 (64 channels) - for BrainAmpDC (with REF)
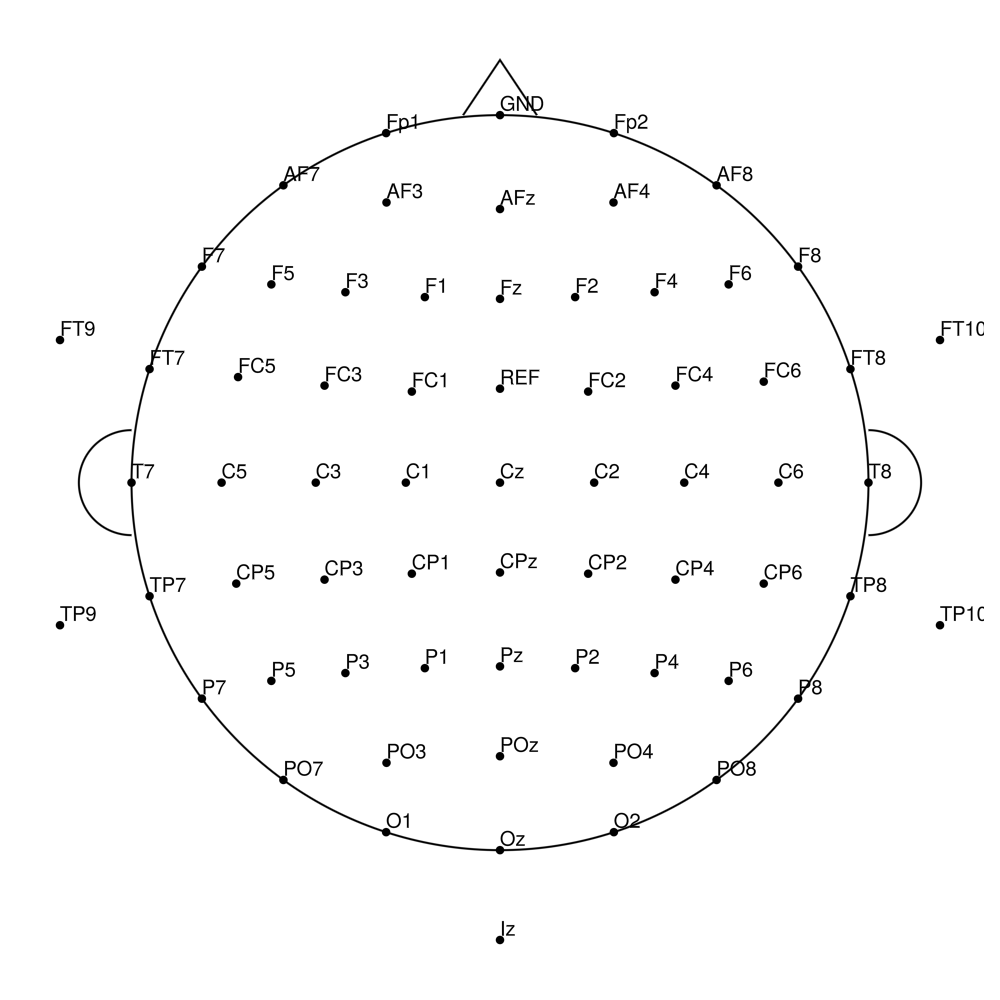
CACS-96 (96 channels) - for BrainAmpDC (with REF)
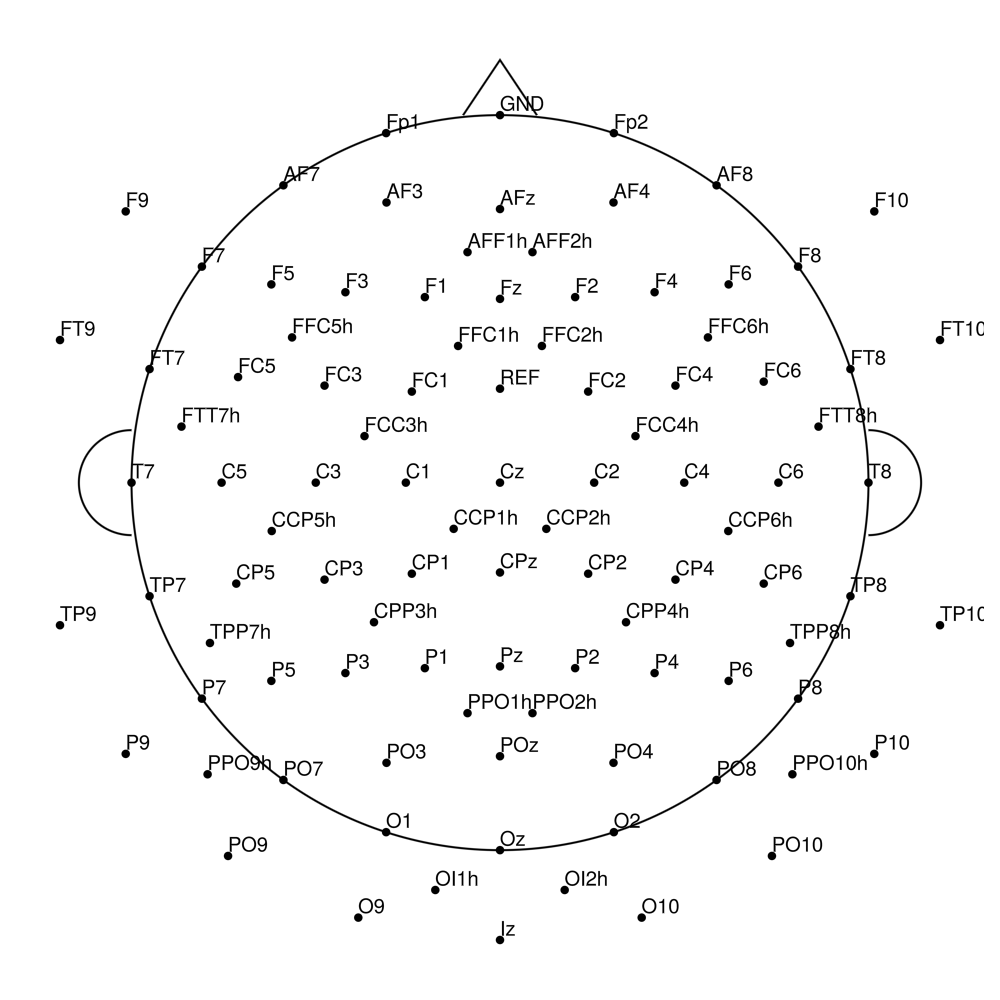
CACS-128 (128 channels) - for BrainAmpDC (with REF)
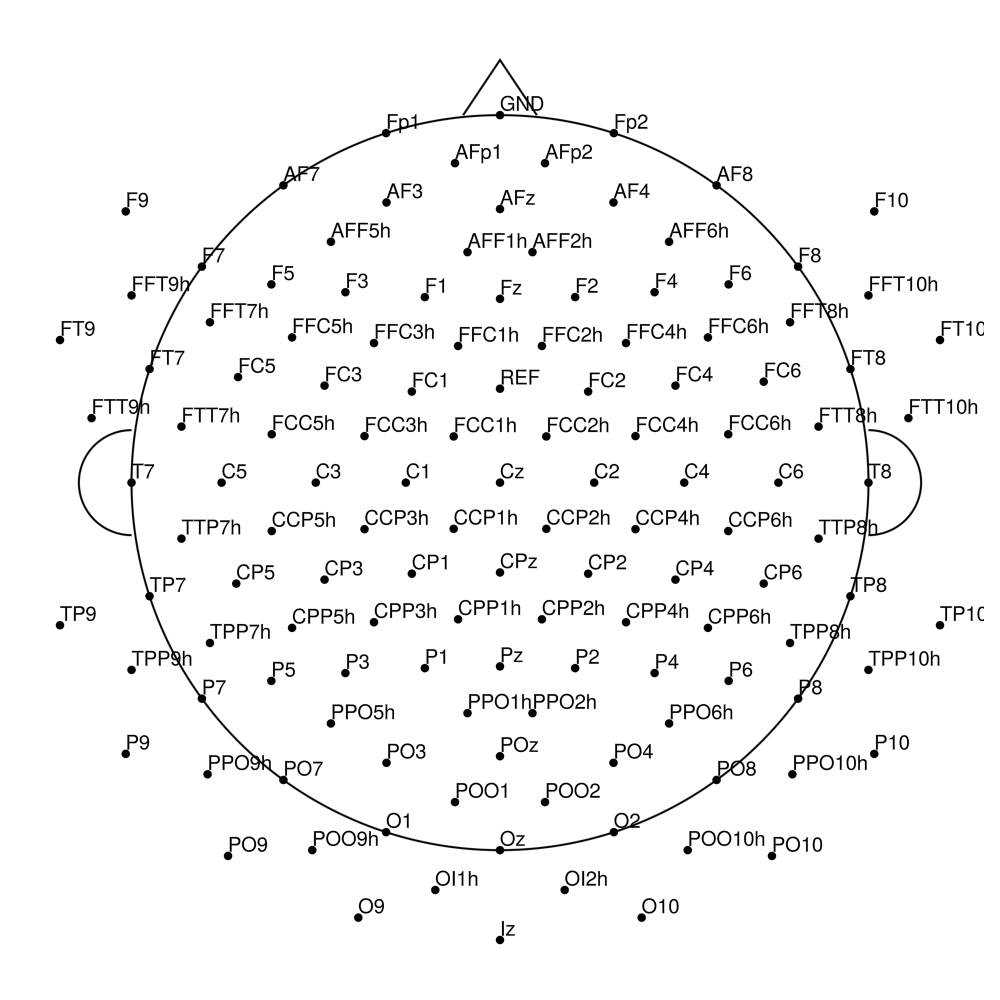
actiCap Slim (ASP) — for LiveAmp
CACS-32 (32 channels) - for LiveAmp (with REF)
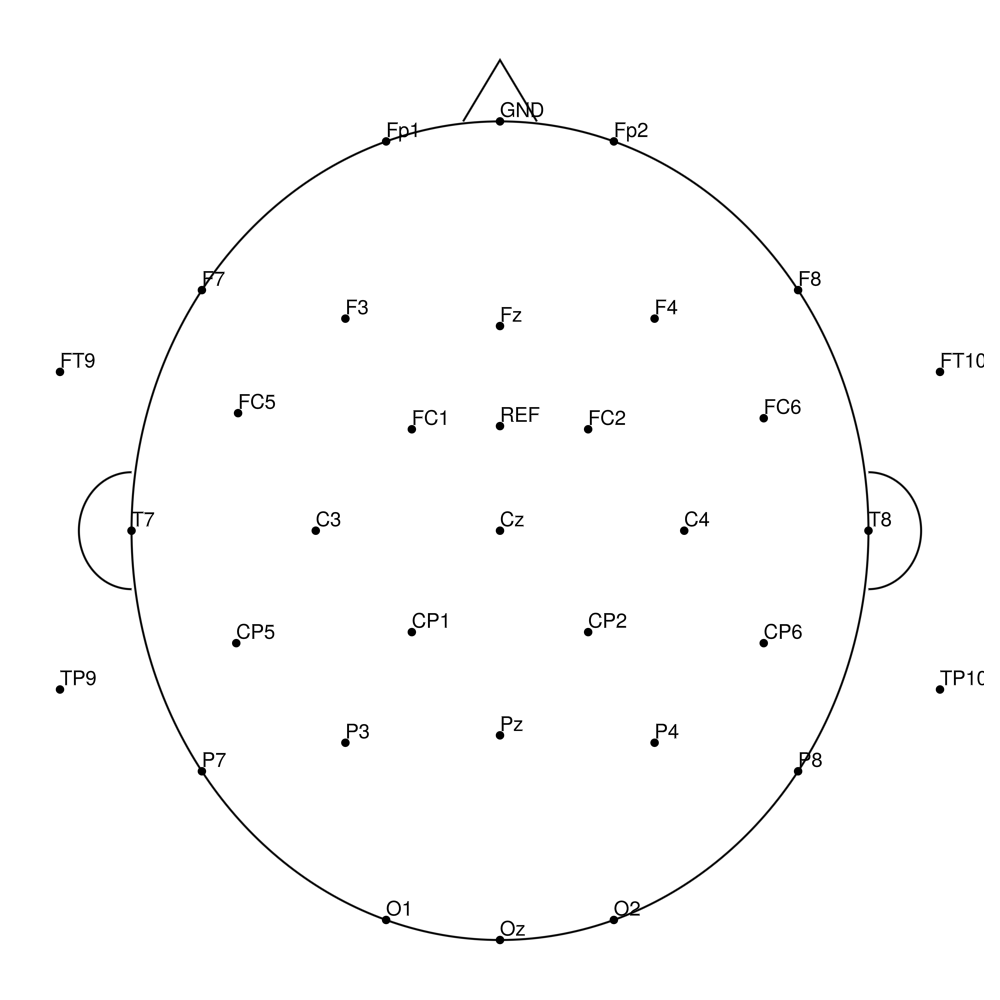
CACS-64 (64 channels) - for LiveAmp (with REF)
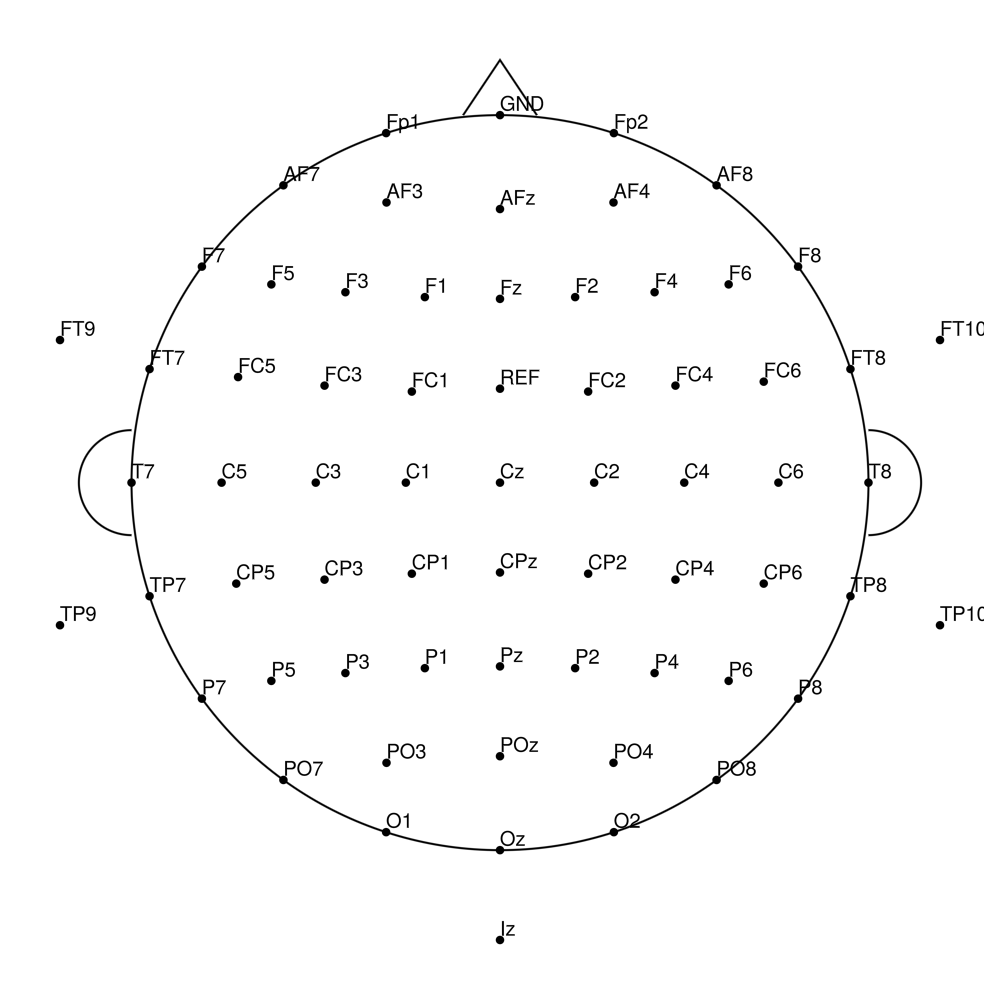
actiCap Slim (ASP) — for actiCHamp Plus
CACS-32 (32 channels) - for actiCHamp Plus (no REF)
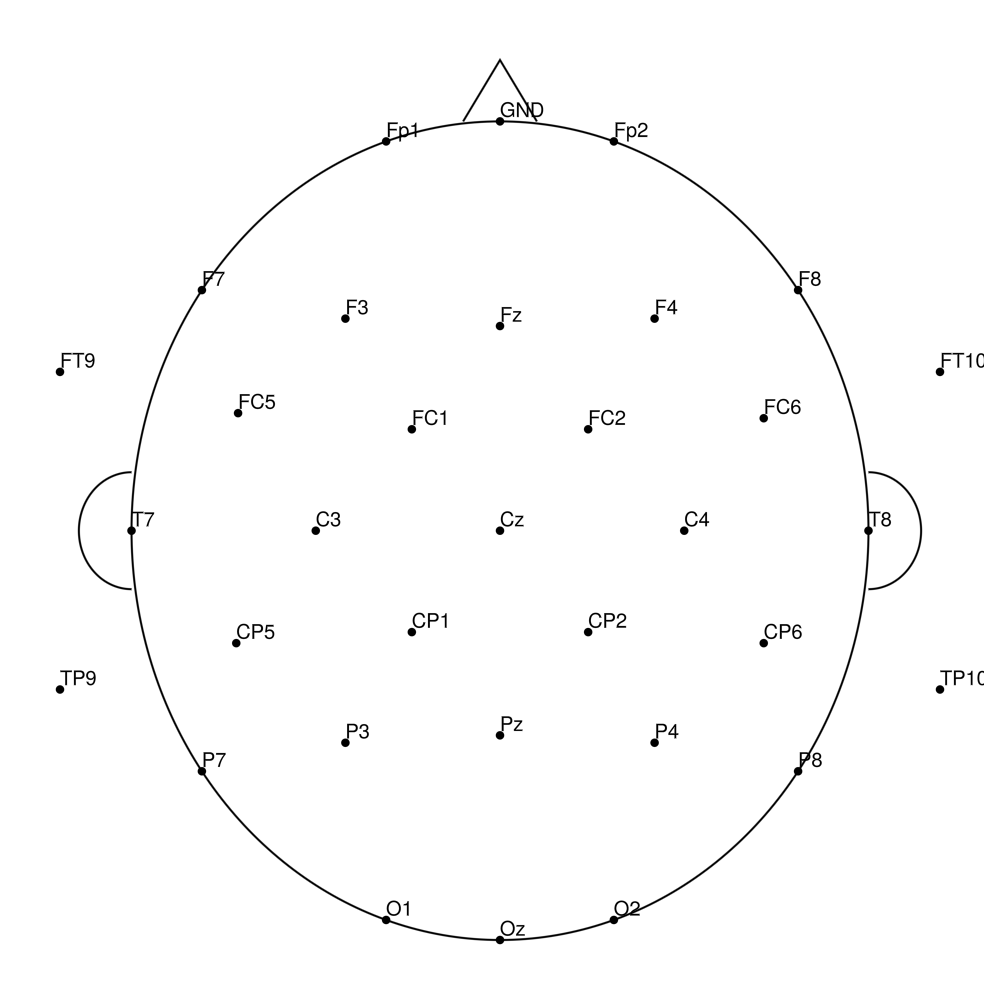
CACS-64 (64 channels) - for actiCHamp Plus (no REF)
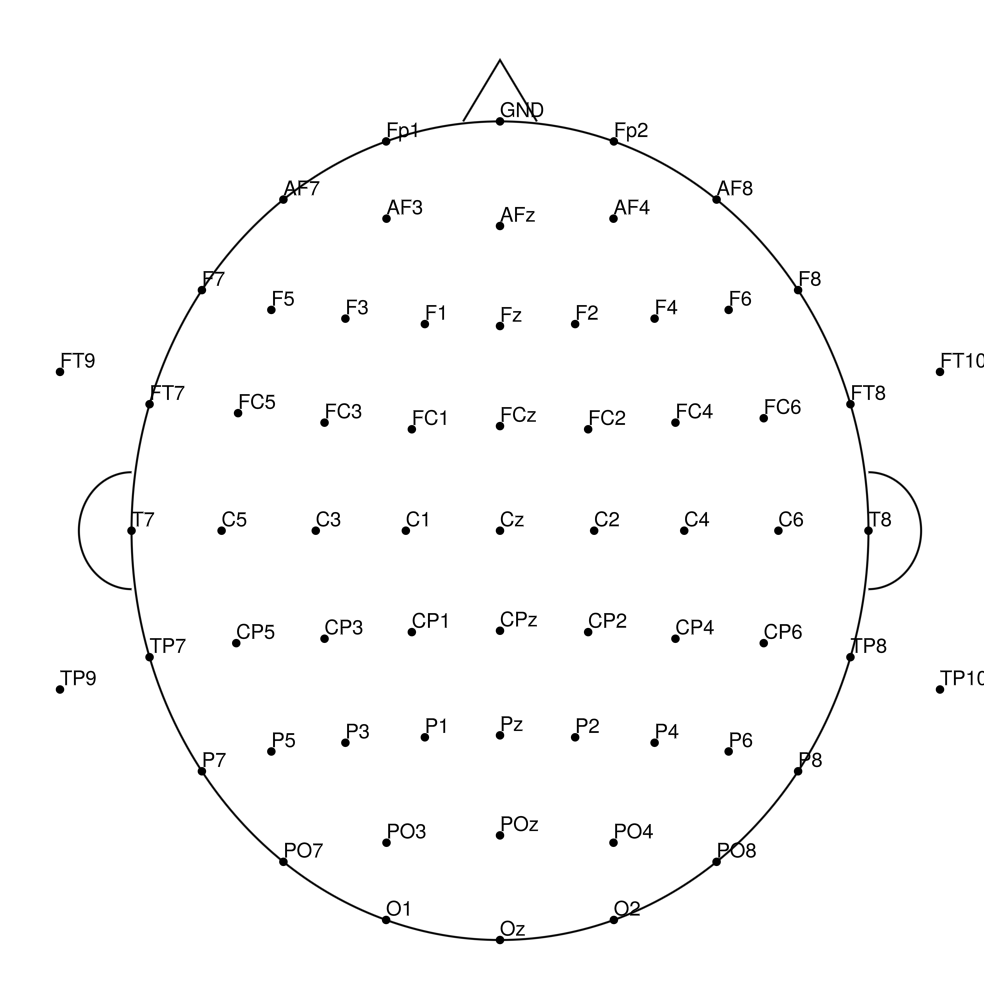
CACS-96 (96 channels) - for actiCHamp Plus (no REF)
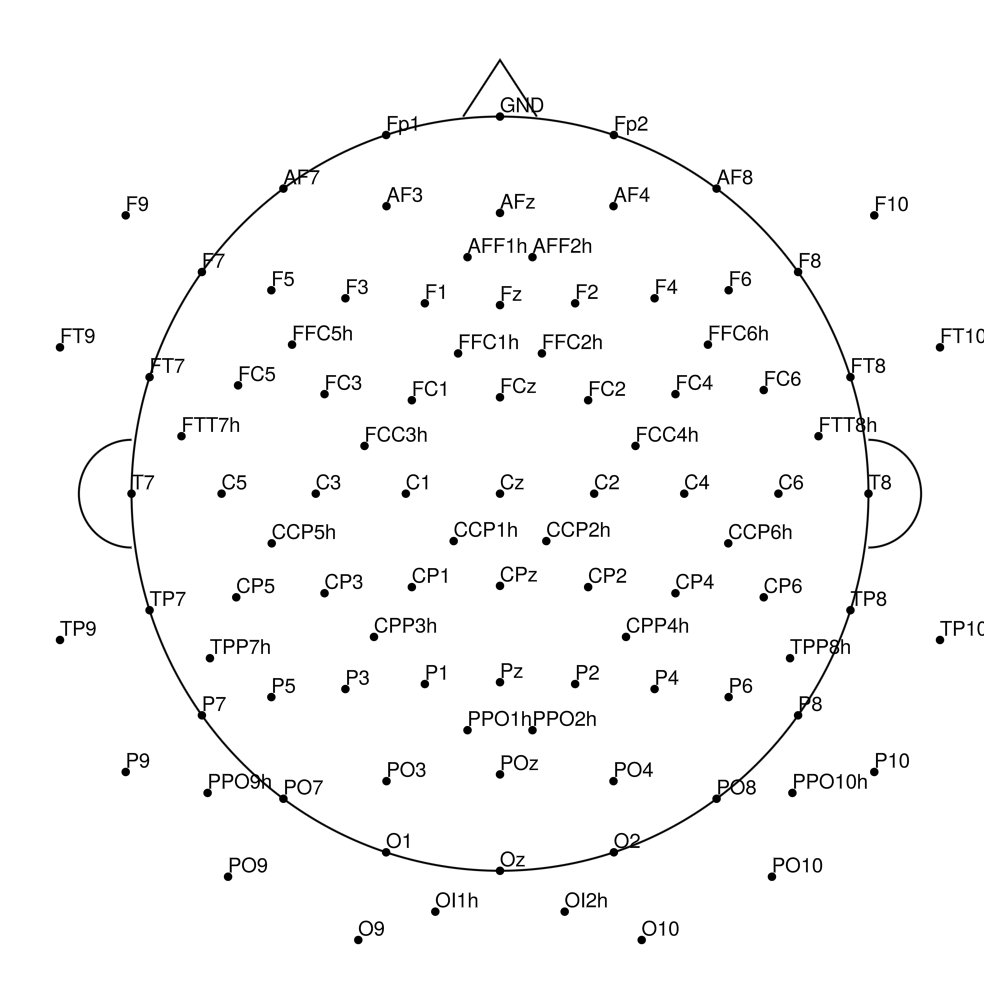
CACS-128 (128 channels) - for actiCHamp Plus (no REF)
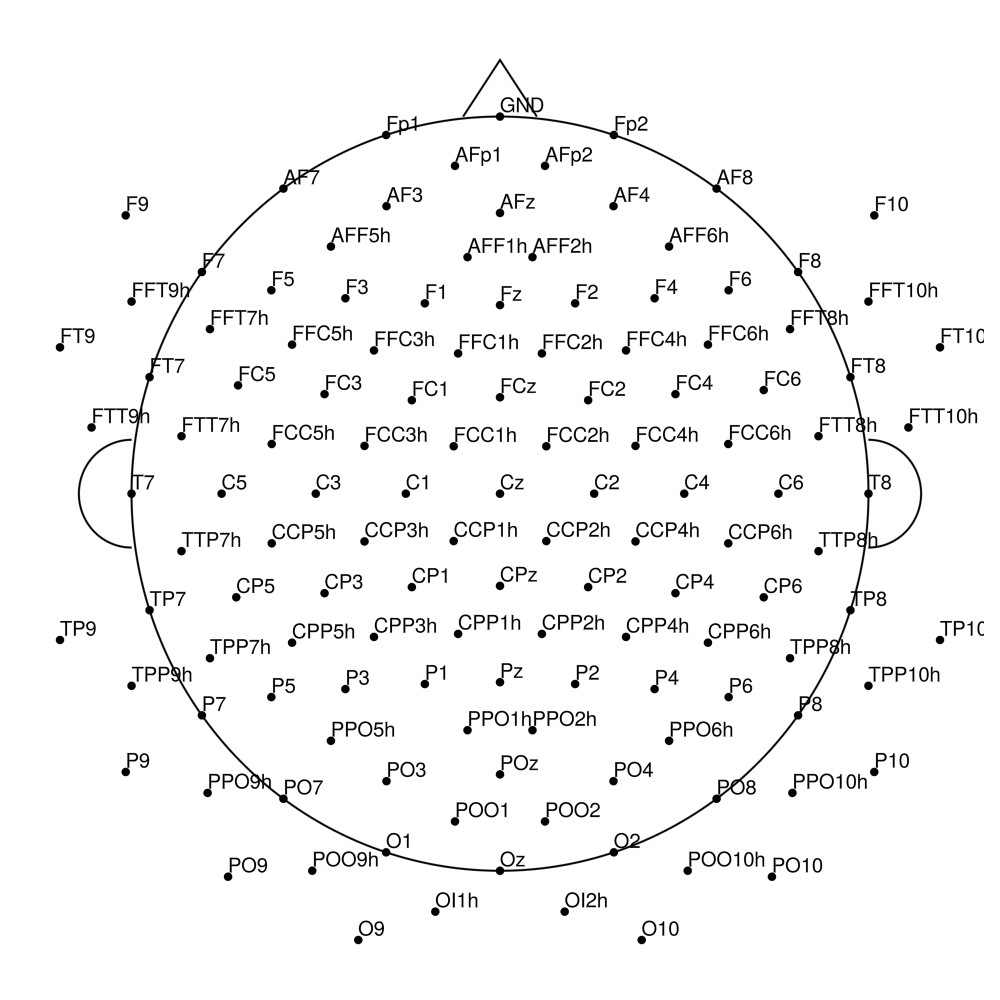
CACS-160 (160 channels) - for actiCHamp Plus (no REF)
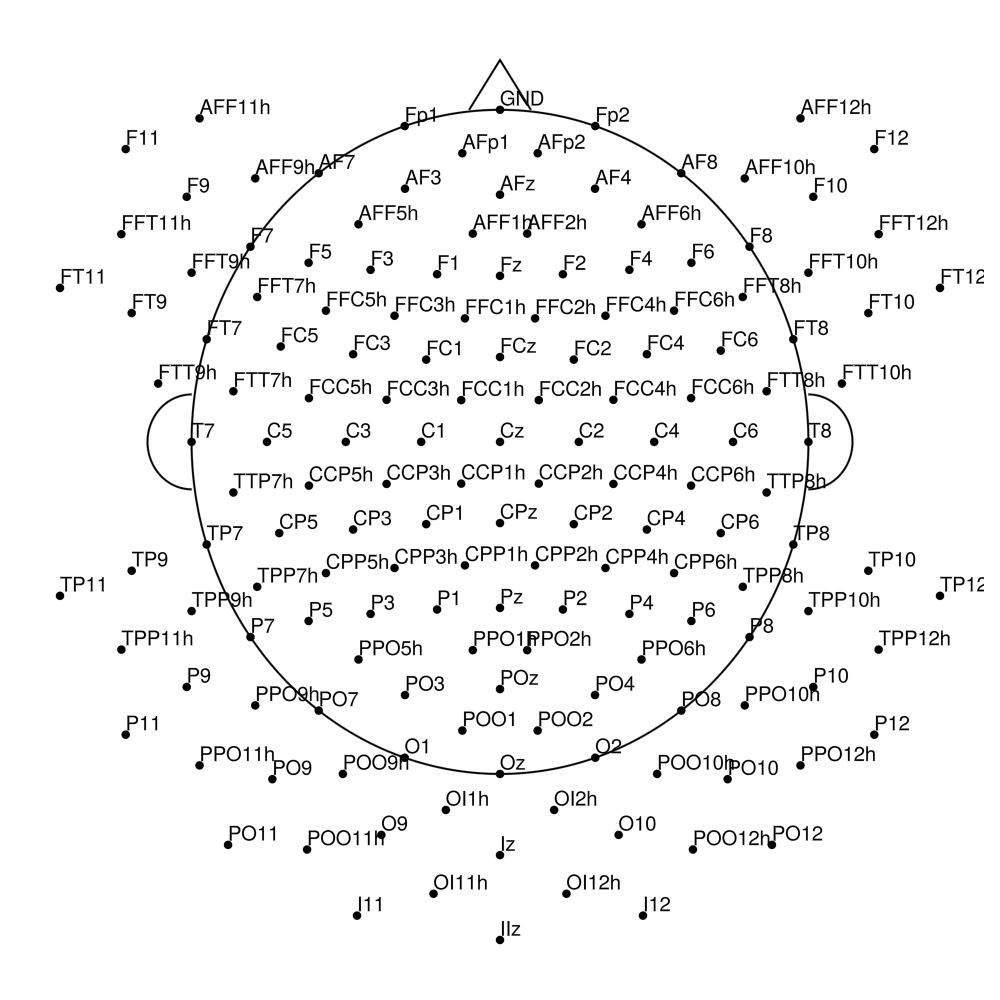
actiCap Snap — for BrainAmpDC
CACS-21 (21 channels) - for BrainAmpDC (with REF)
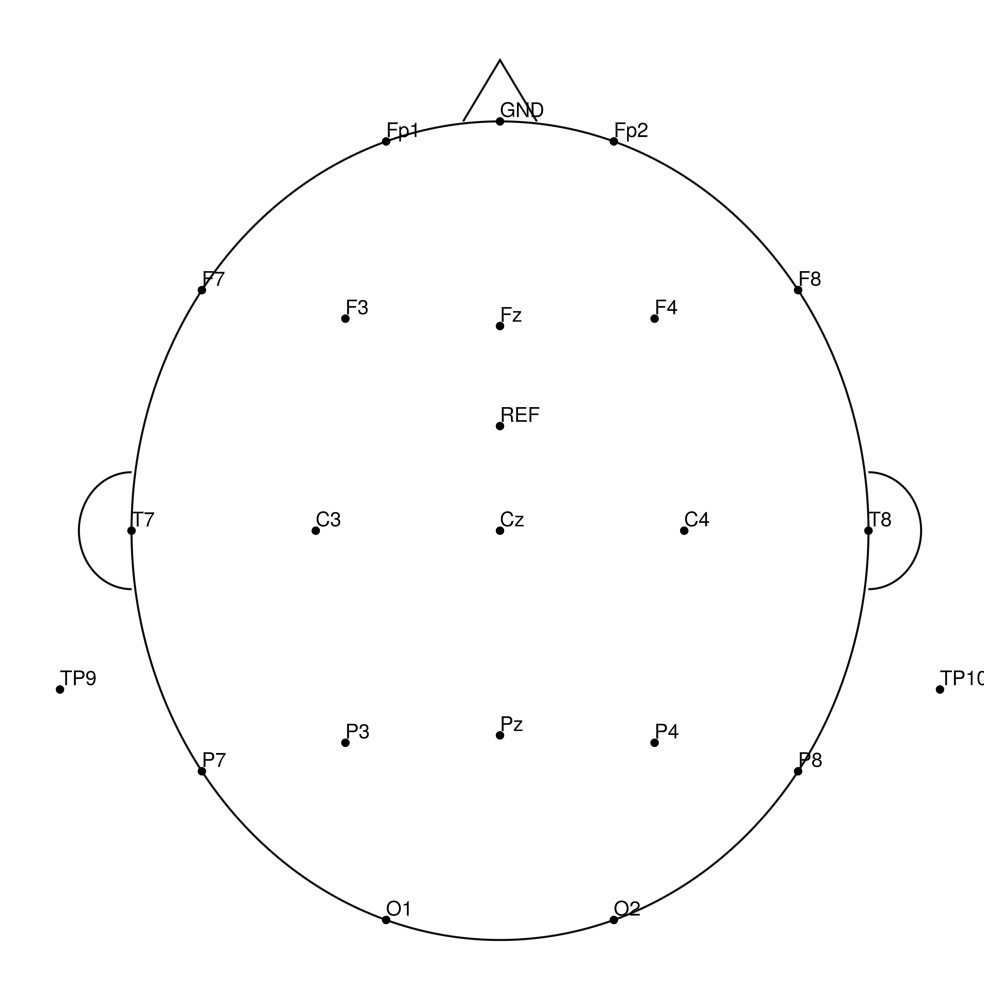
CACS-32 (32 channels) - snap for BrainAmpDC (with REF)
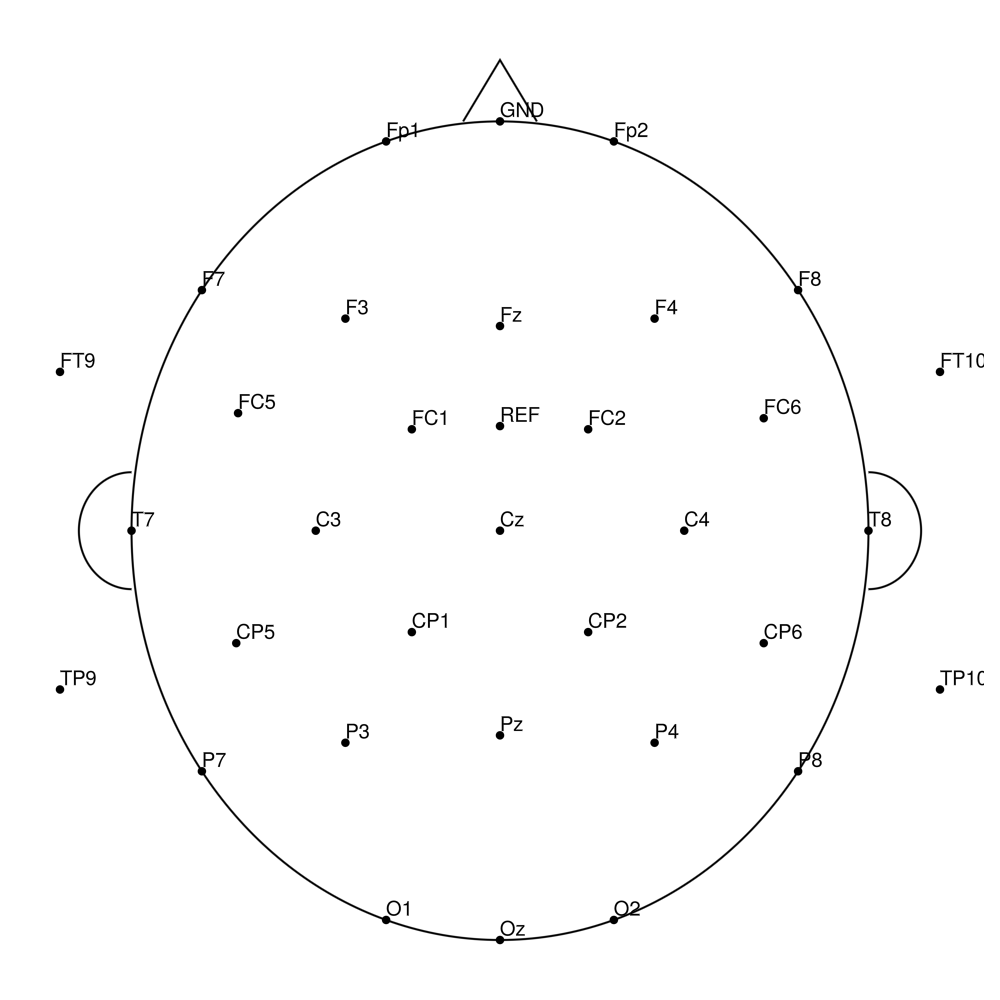
CACS-64 (64 channels) - snap for BrainAmpDC (with REF)
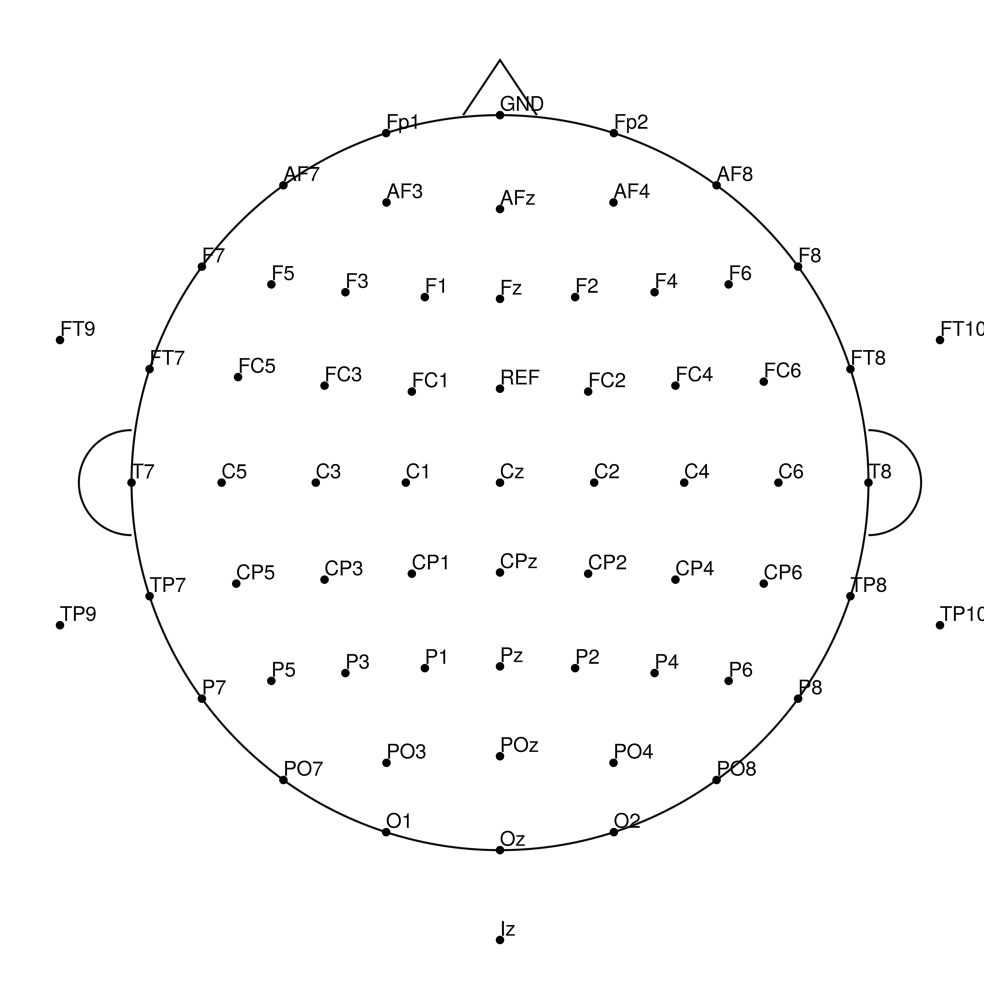
CACS-96 (96 channels) - snap for BrainAmpDC (with REF)
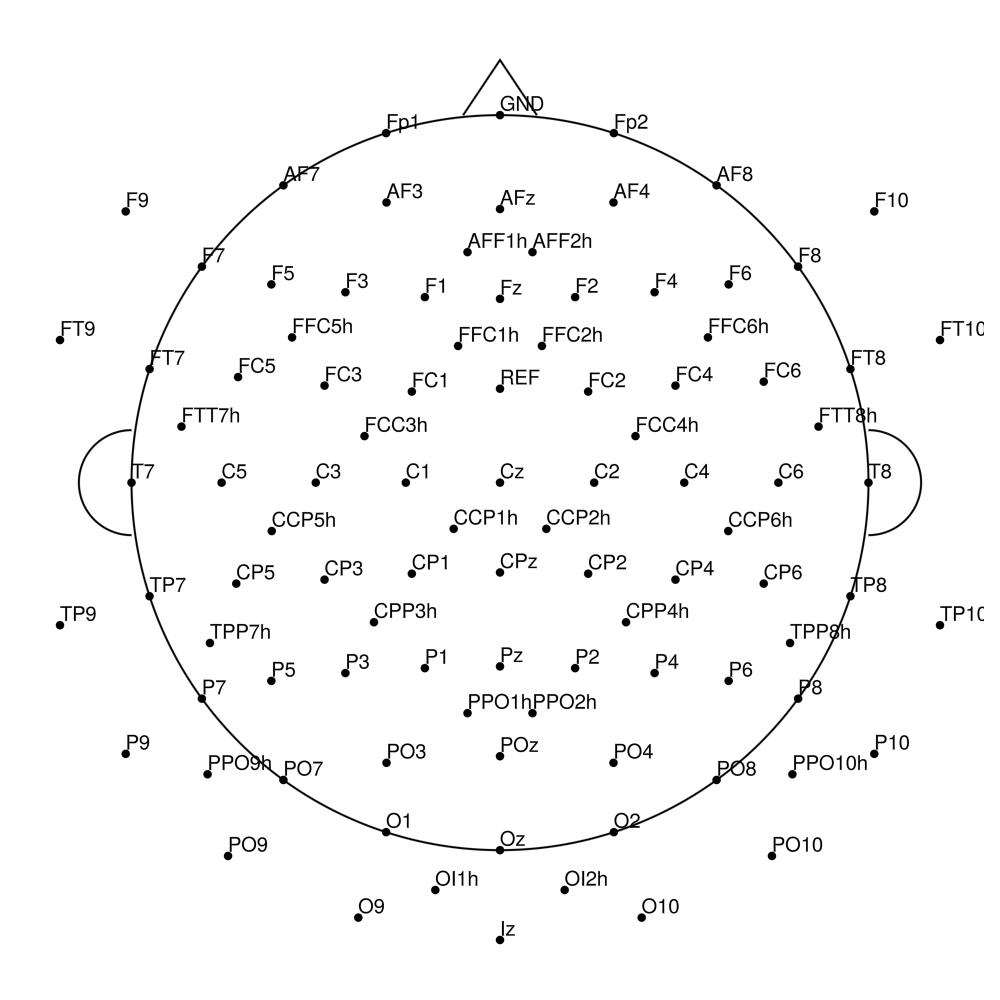
CACS-128 (128 channels) - snap for BrainAmpDC (with REF)
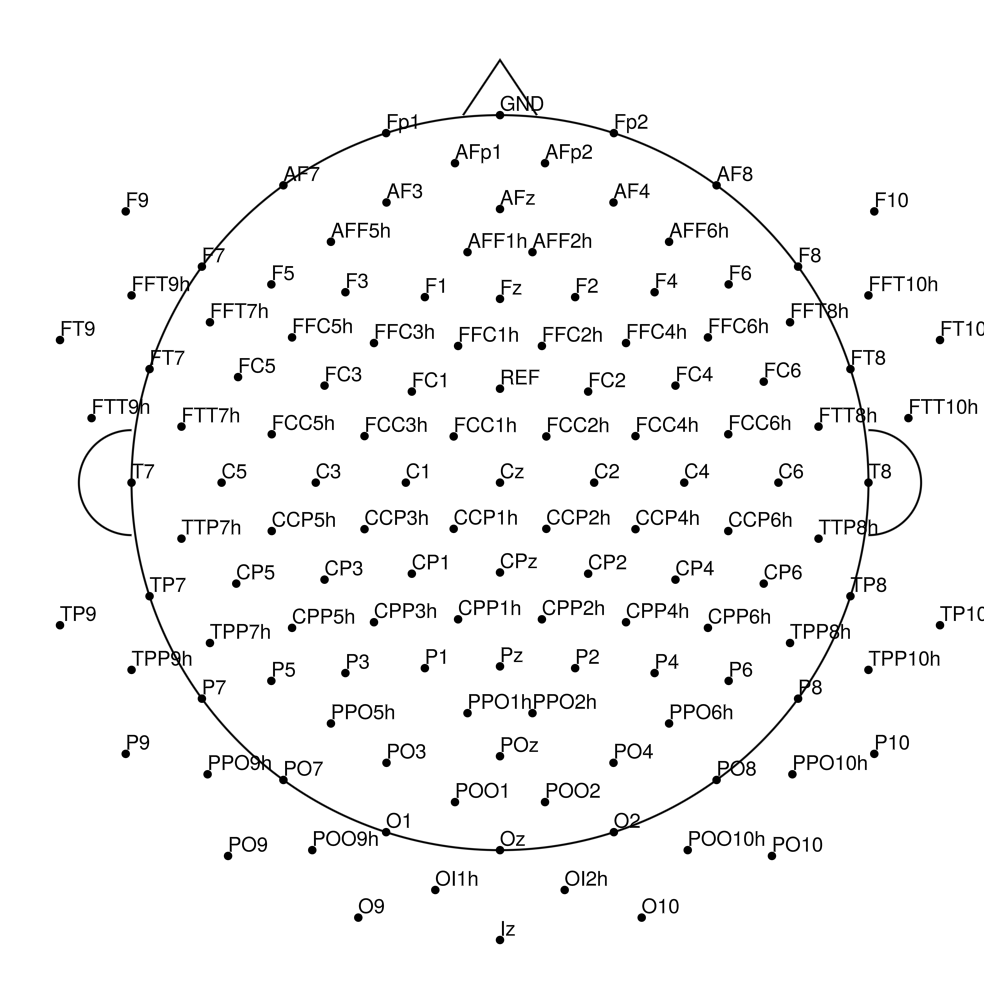
actiCap Snap — for LiveAmp
CACS-21 (21 channels) - snap for LiveAmp (with REF)
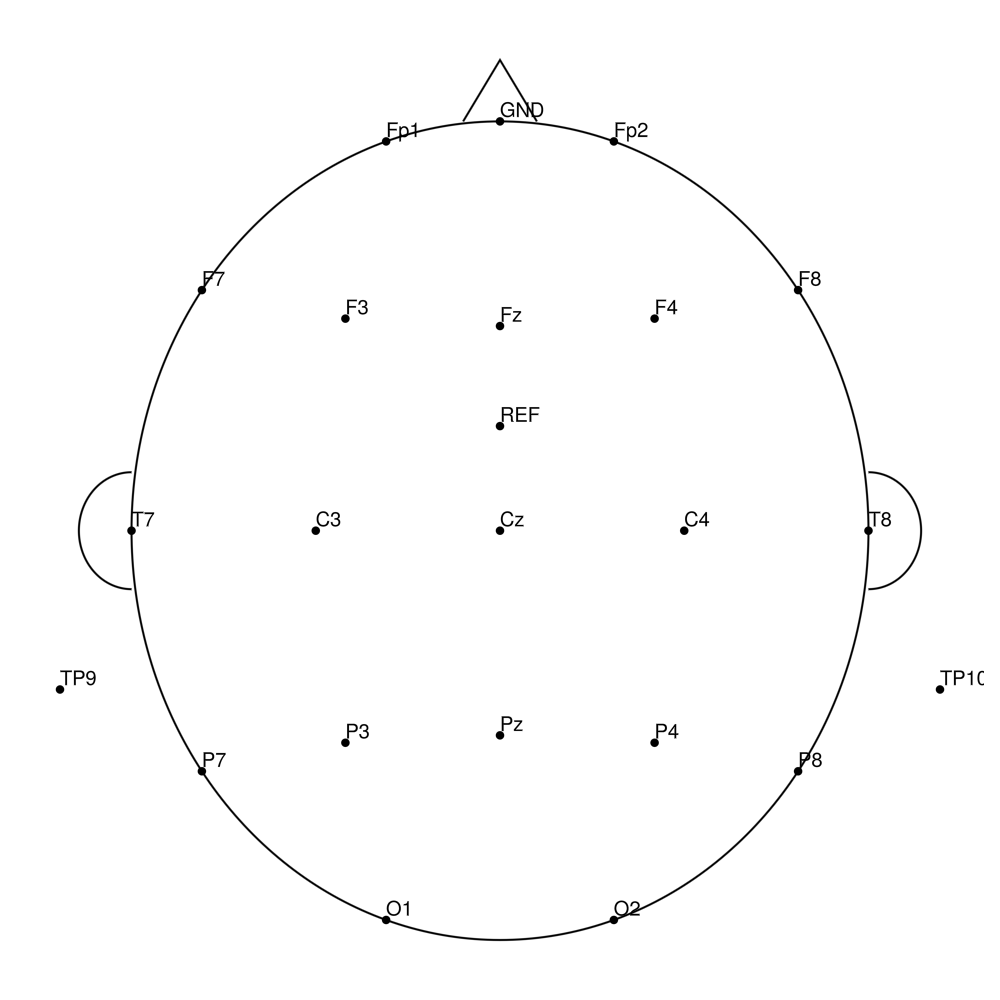
CACS-32 (32 channels) - snap for LiveAmp (with REF)
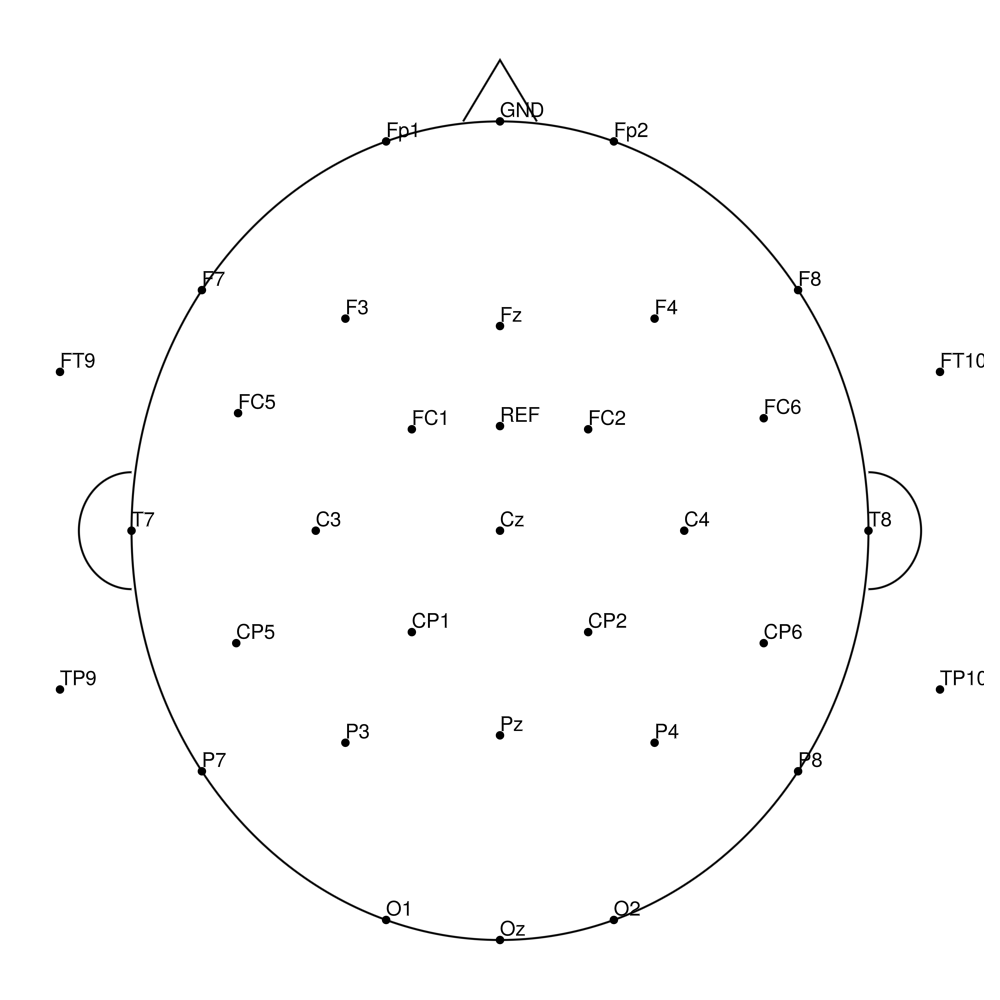
CACS-64 (64 channels) - snap for LiveAmp (with REF)
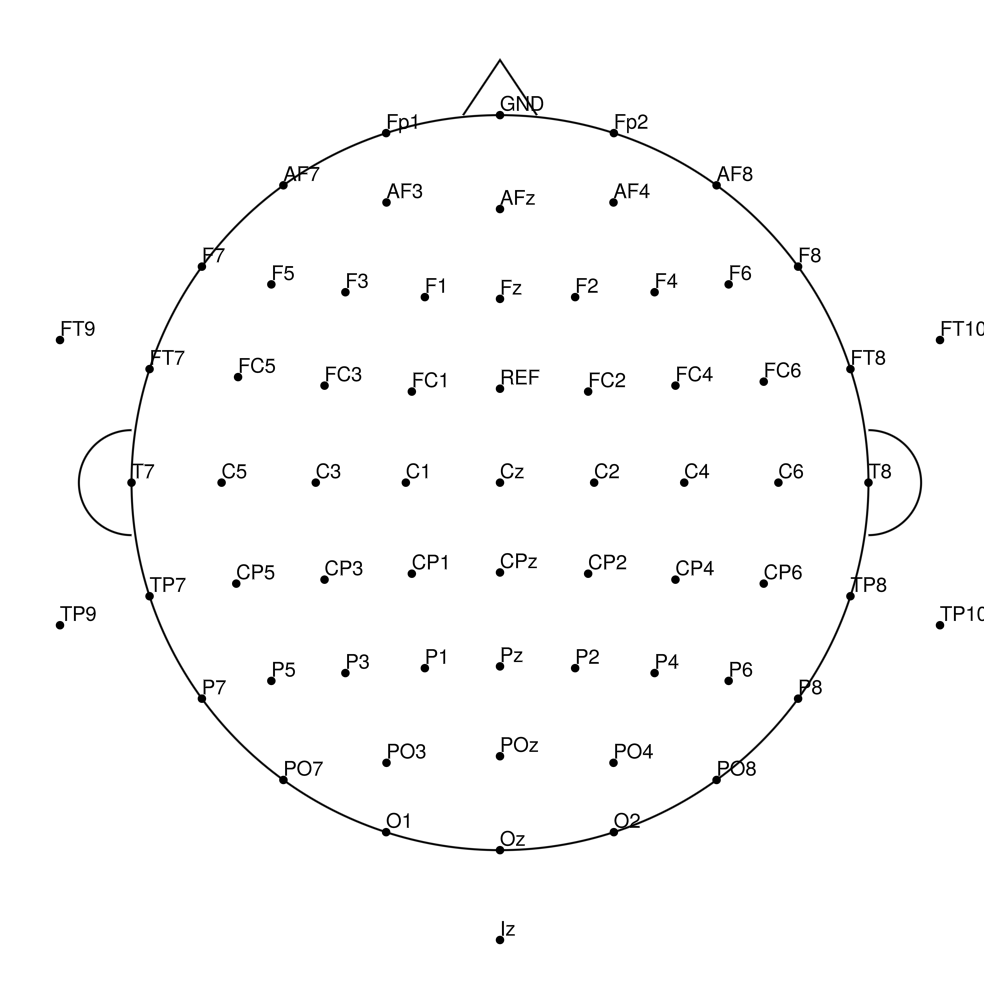
actiCap Snap — for actiCHamp Plus
CACS-21 (21 channels) - snap for actiCHamp Plus (no REF)
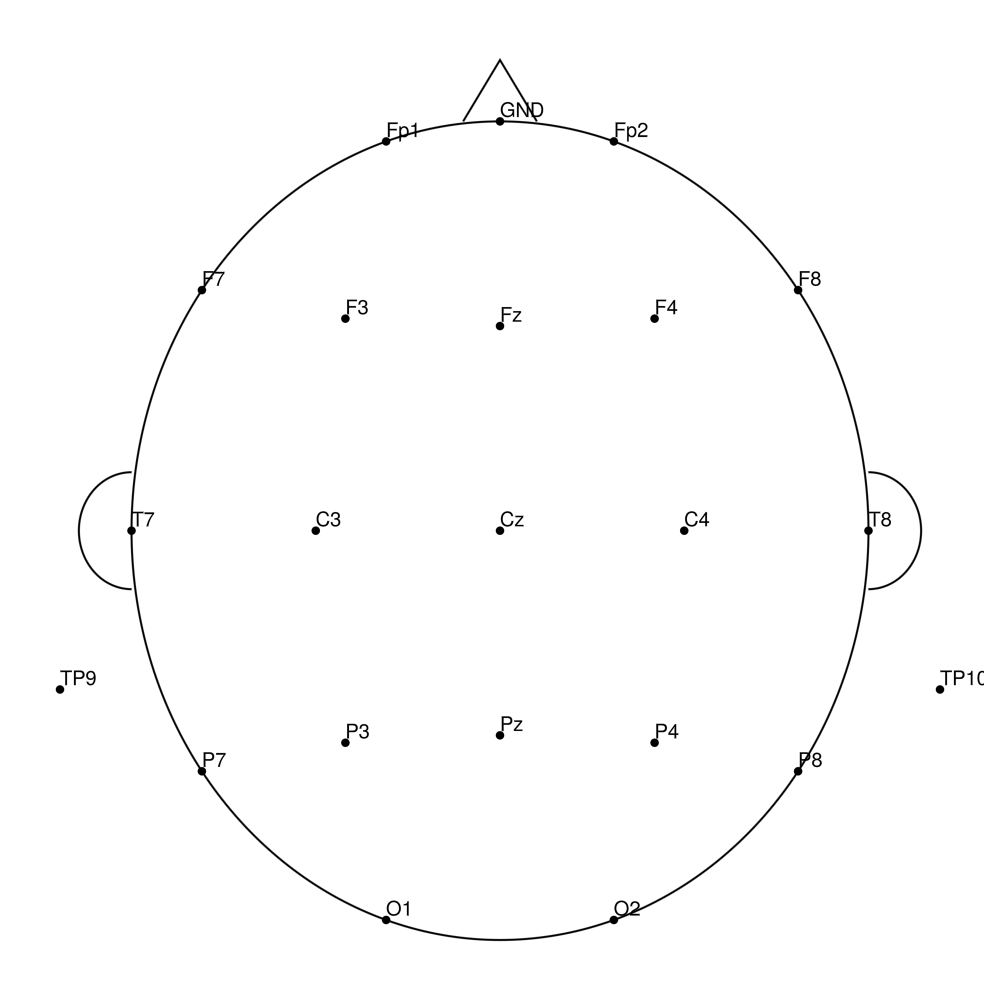
CACS-32 (32 channels) - snap for actiCHamp Plus (no REF)
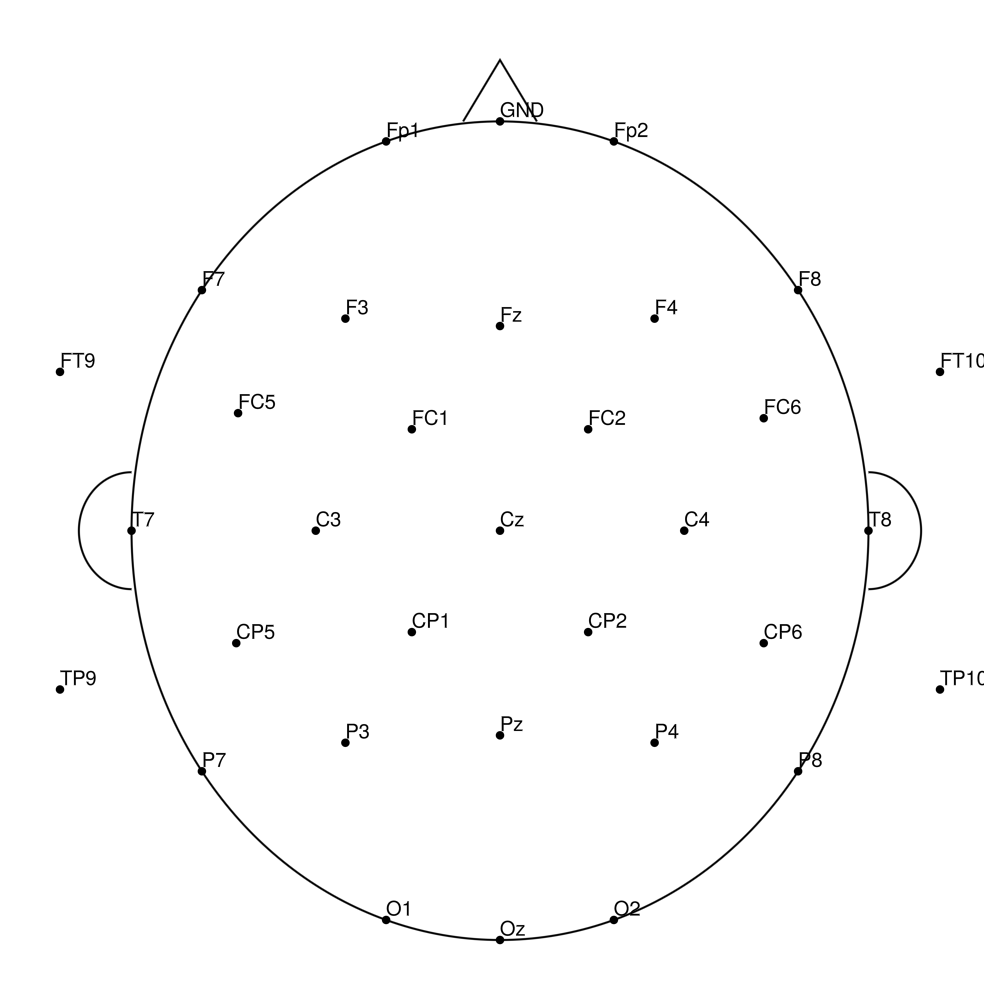
CACS-64 (64 channels) - snap for actiCHamp Plus (no REF)
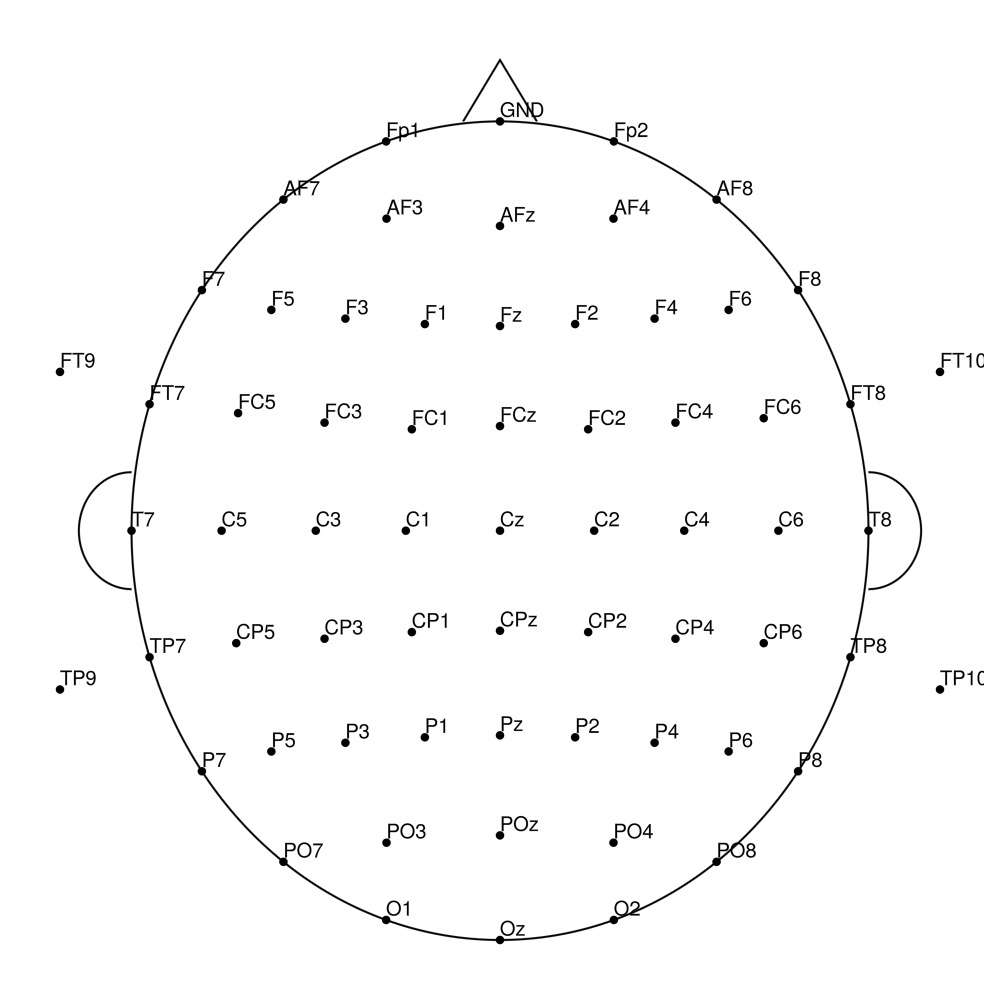
CACS-96 (96 channels) - snap for actiCHamp Plus (no REF)
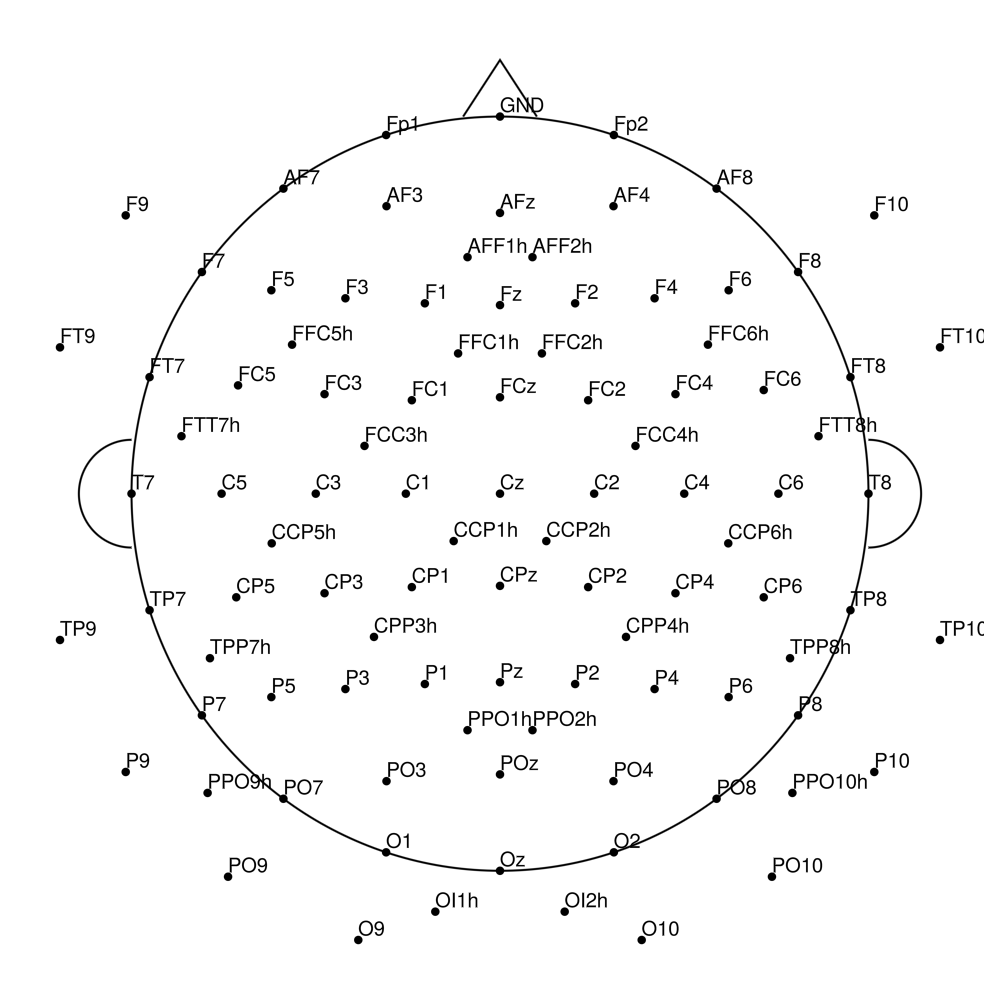
CACS-128 (128 channels) - snap for actiCHamp Plus (no REF)
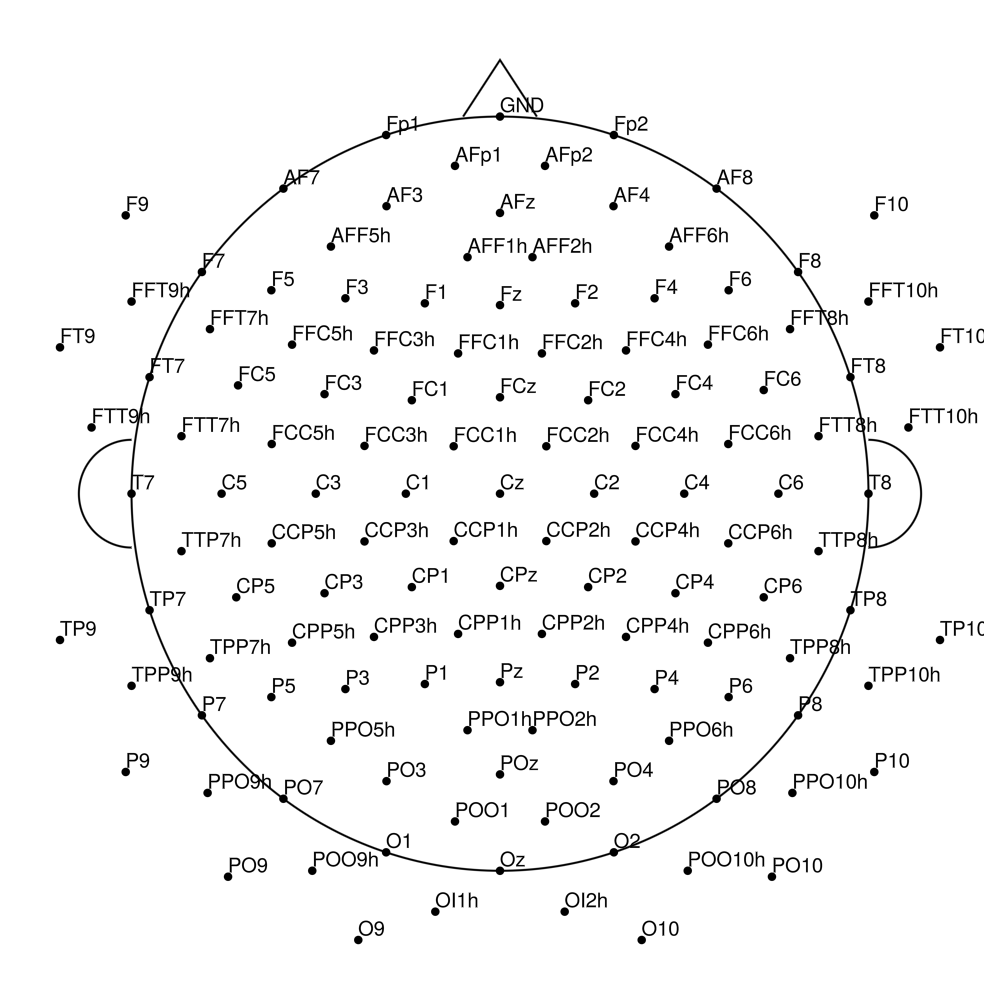
CACS-160 (160 channels) - snap for actiCHamp Plus (no REF)
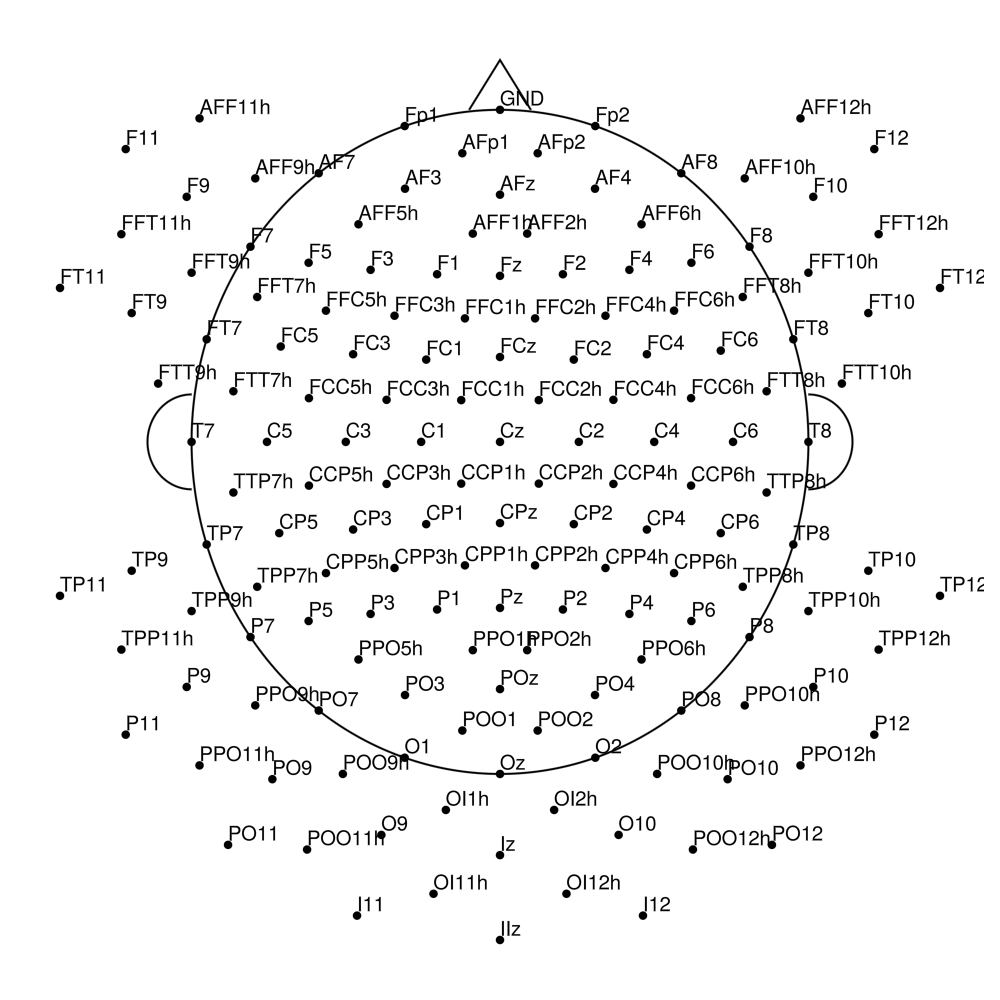
actiCap Snap for LiveAmp (CLACS)
CLACS-32 (32 channels) - actiCap snap for LiveAmp
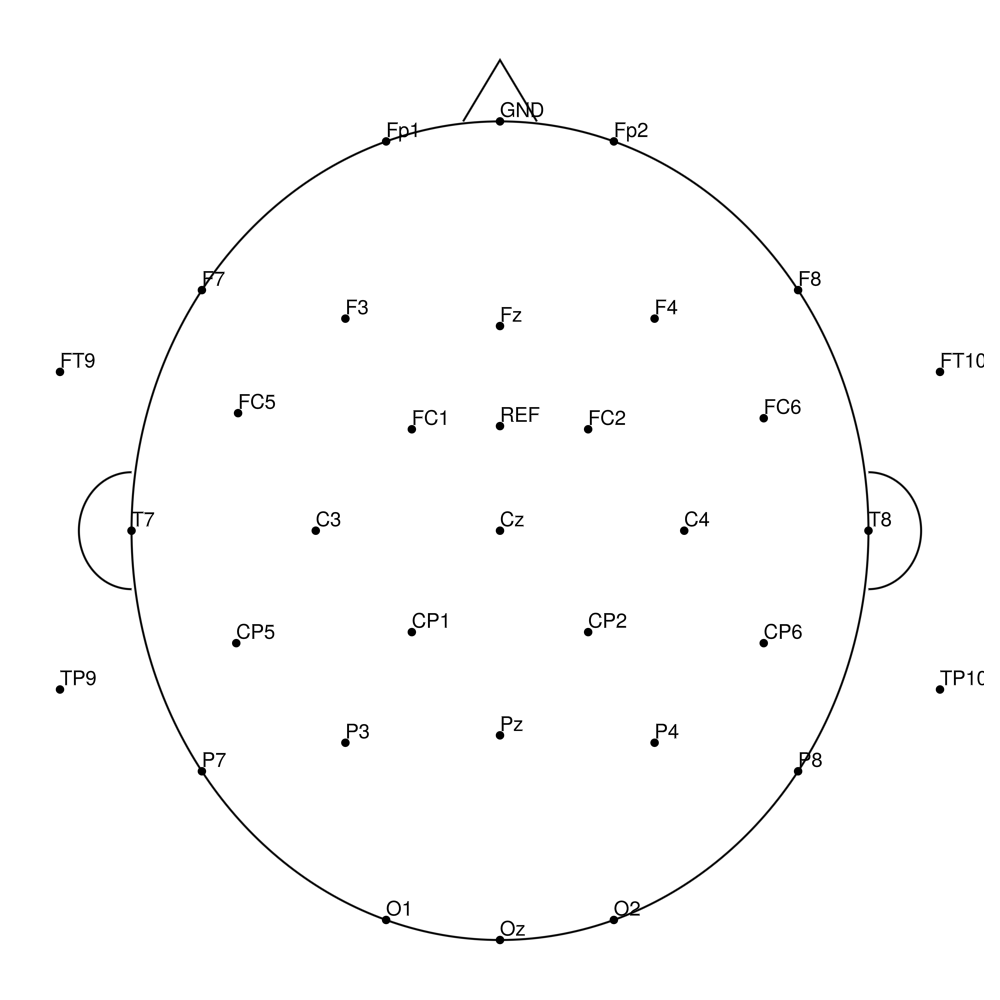
EasyCap
EasyCap montage configurations (M-series):
easycapM1
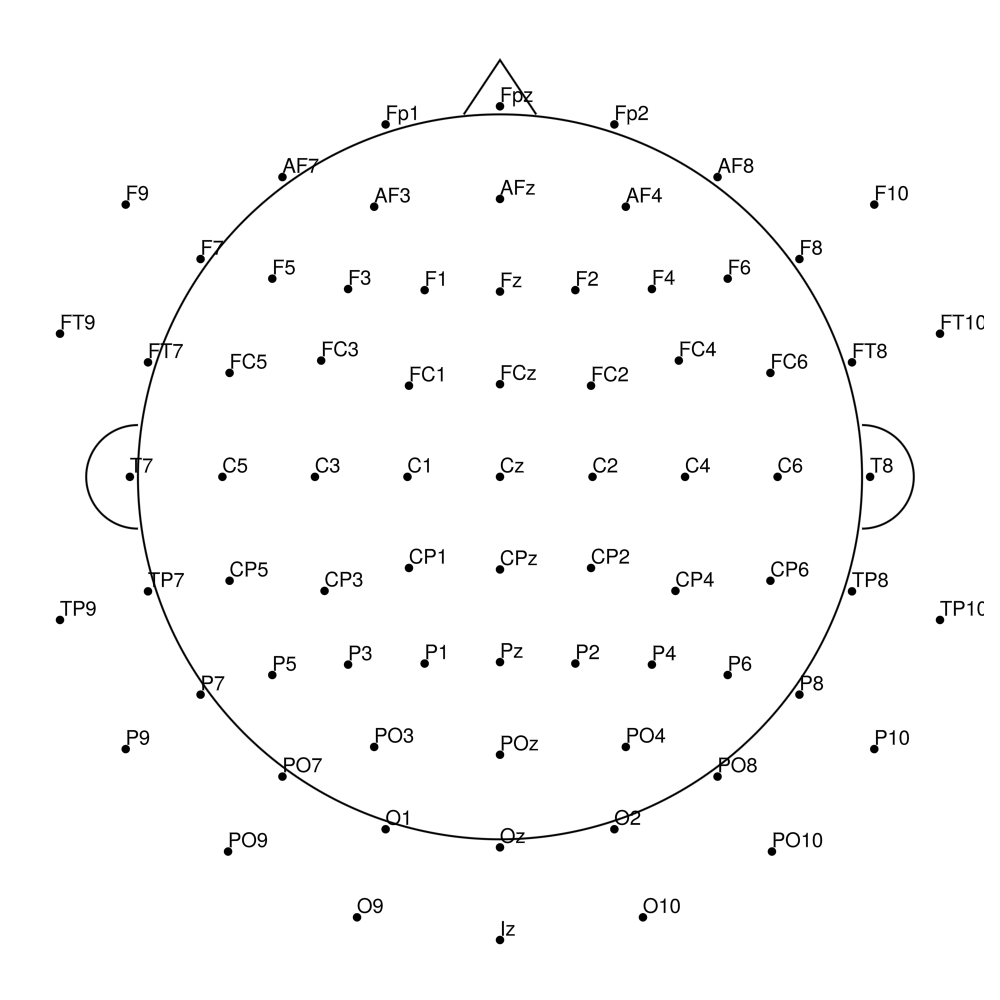
easycapM3
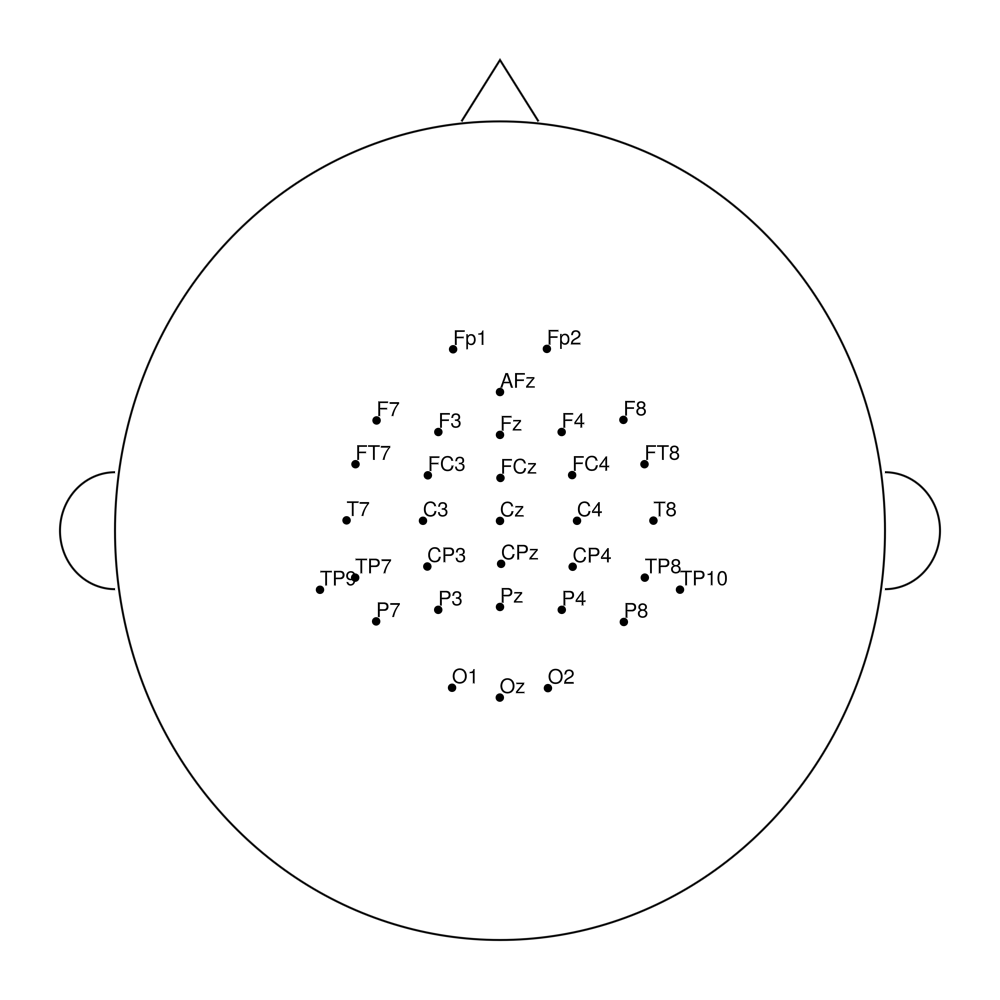
easycapM7
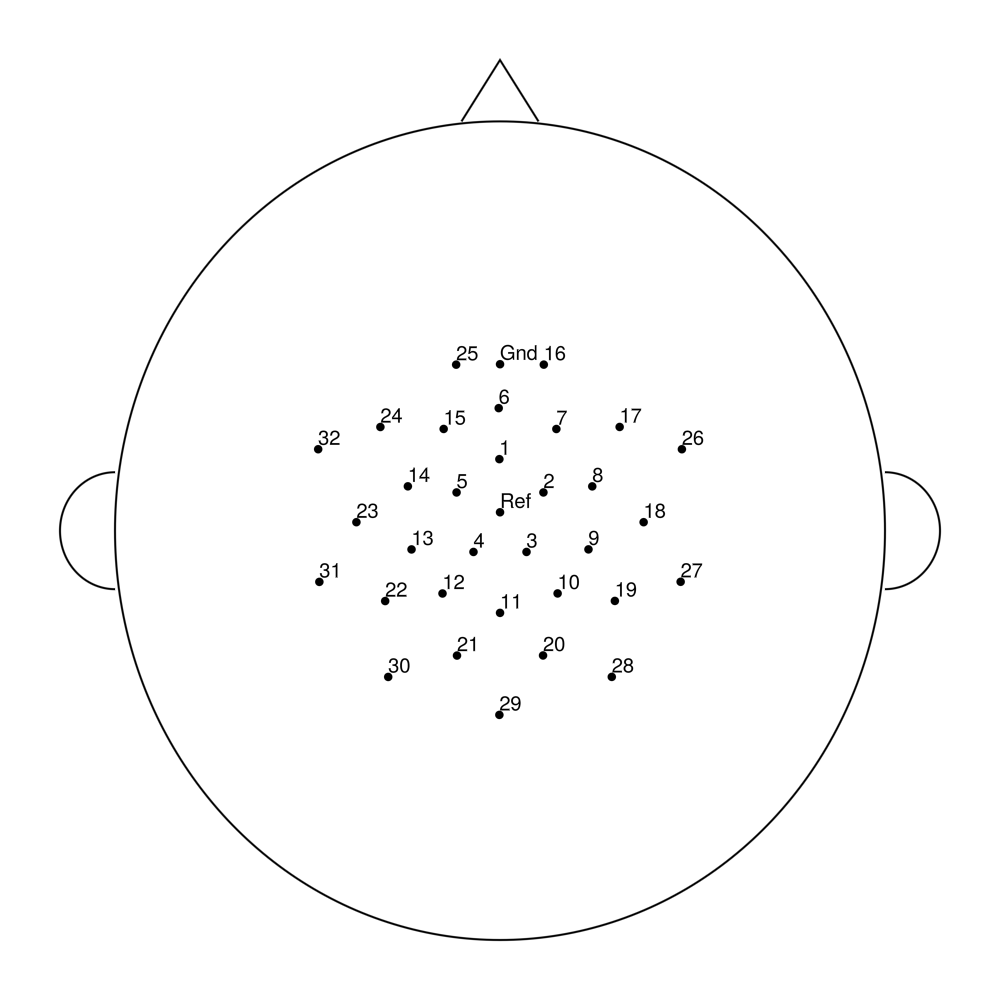
easycapM10
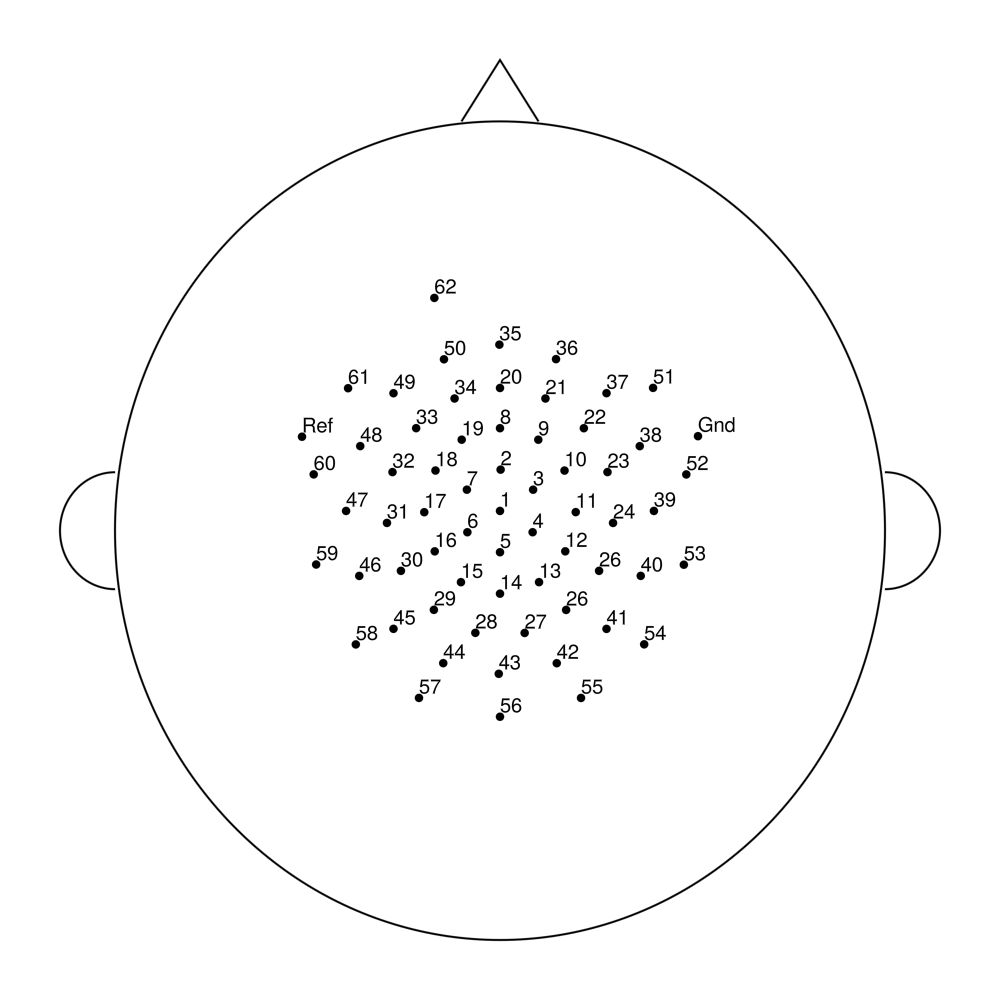
easycapM11
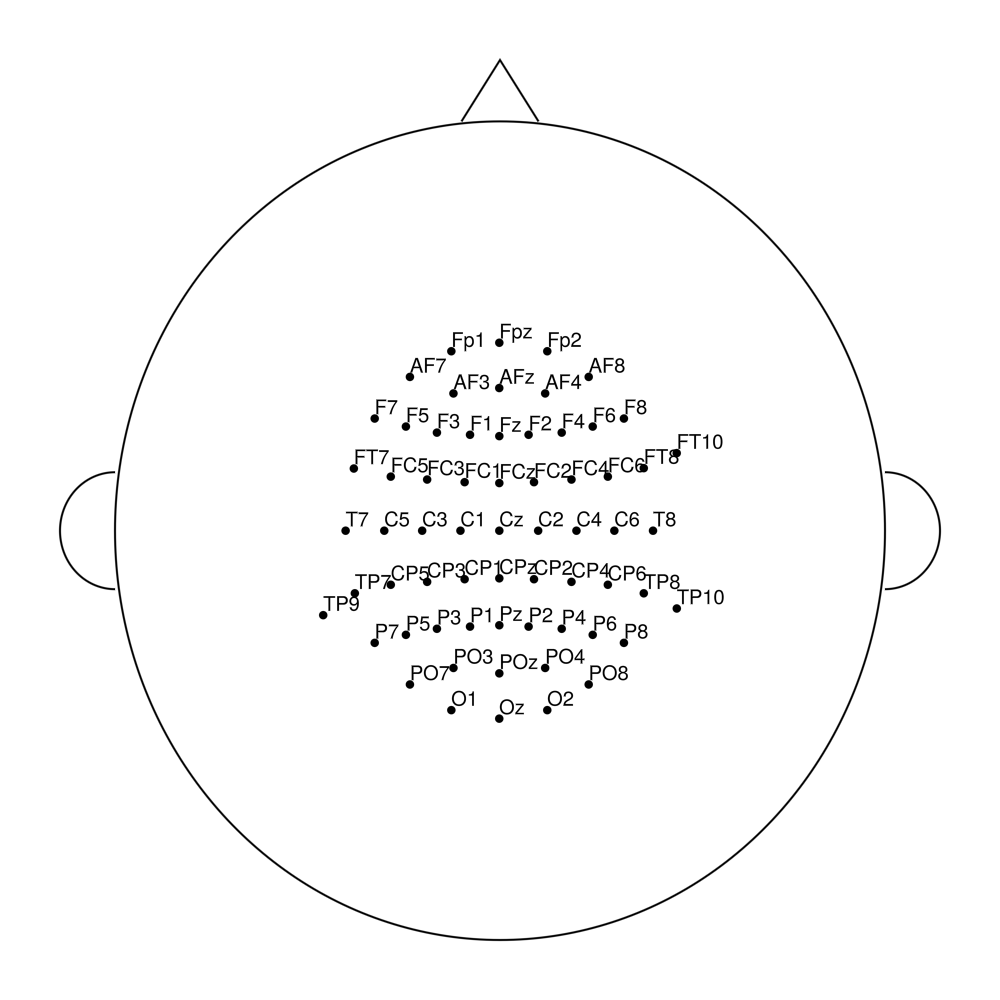
easycapM14
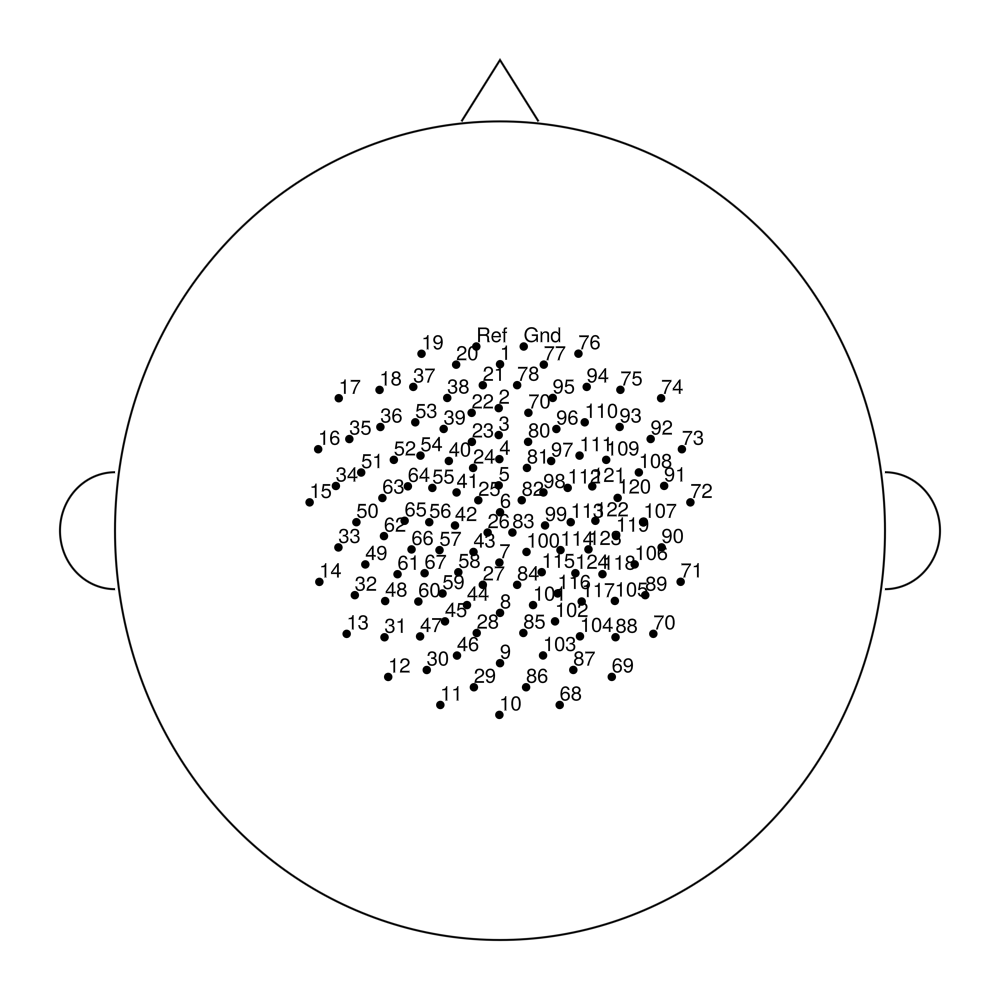
easycapM15
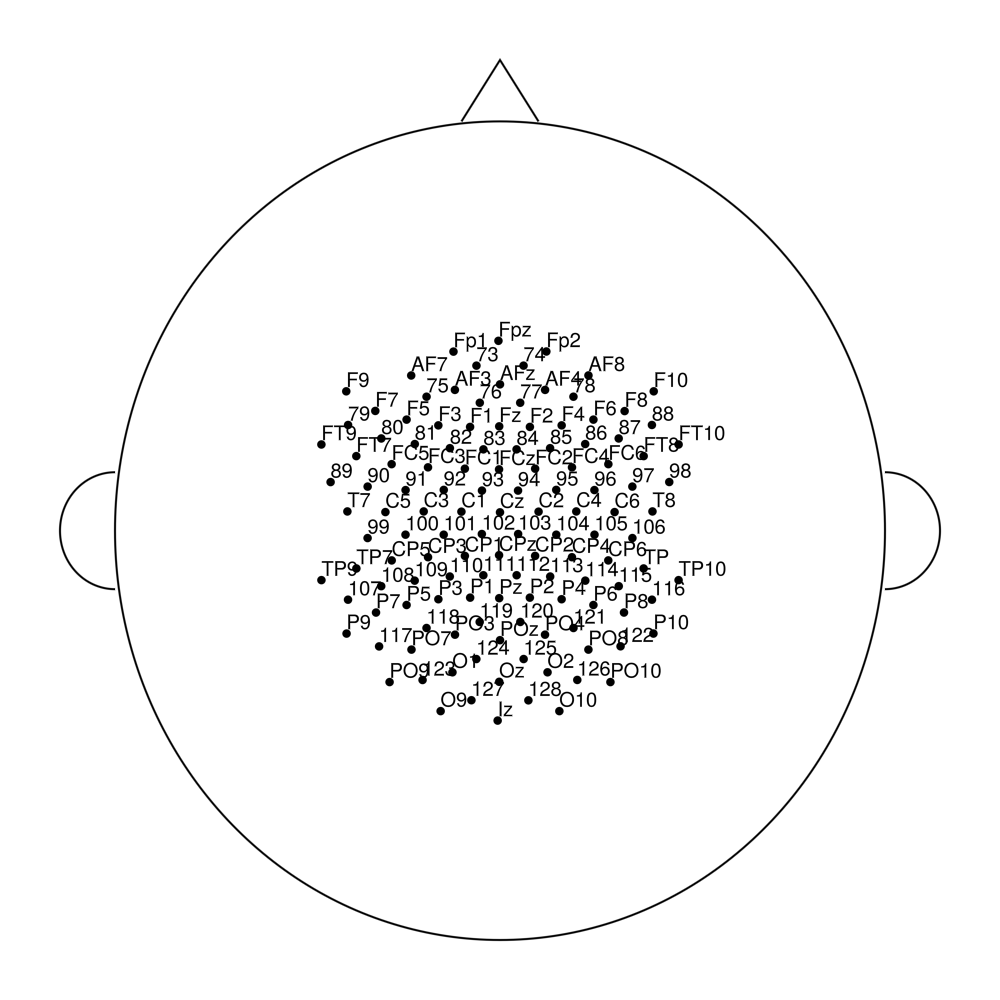
easycapM16
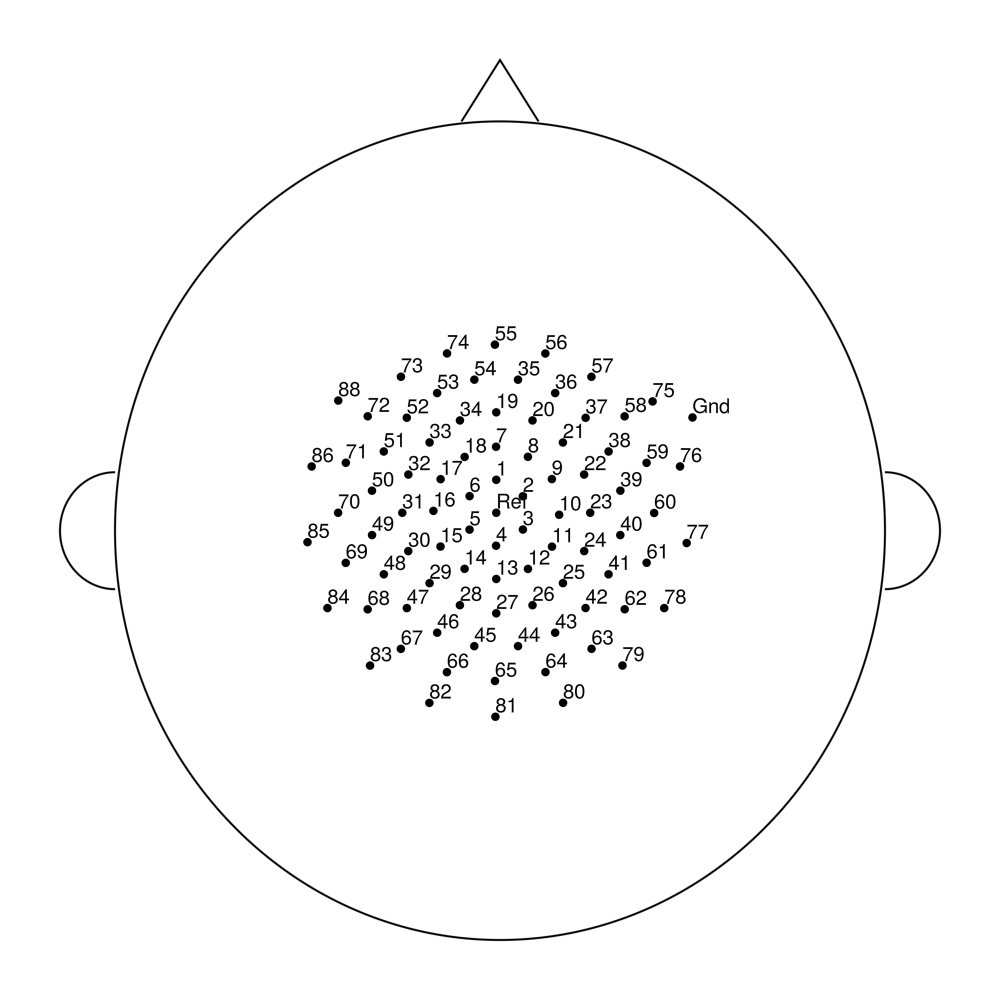
easycapM17
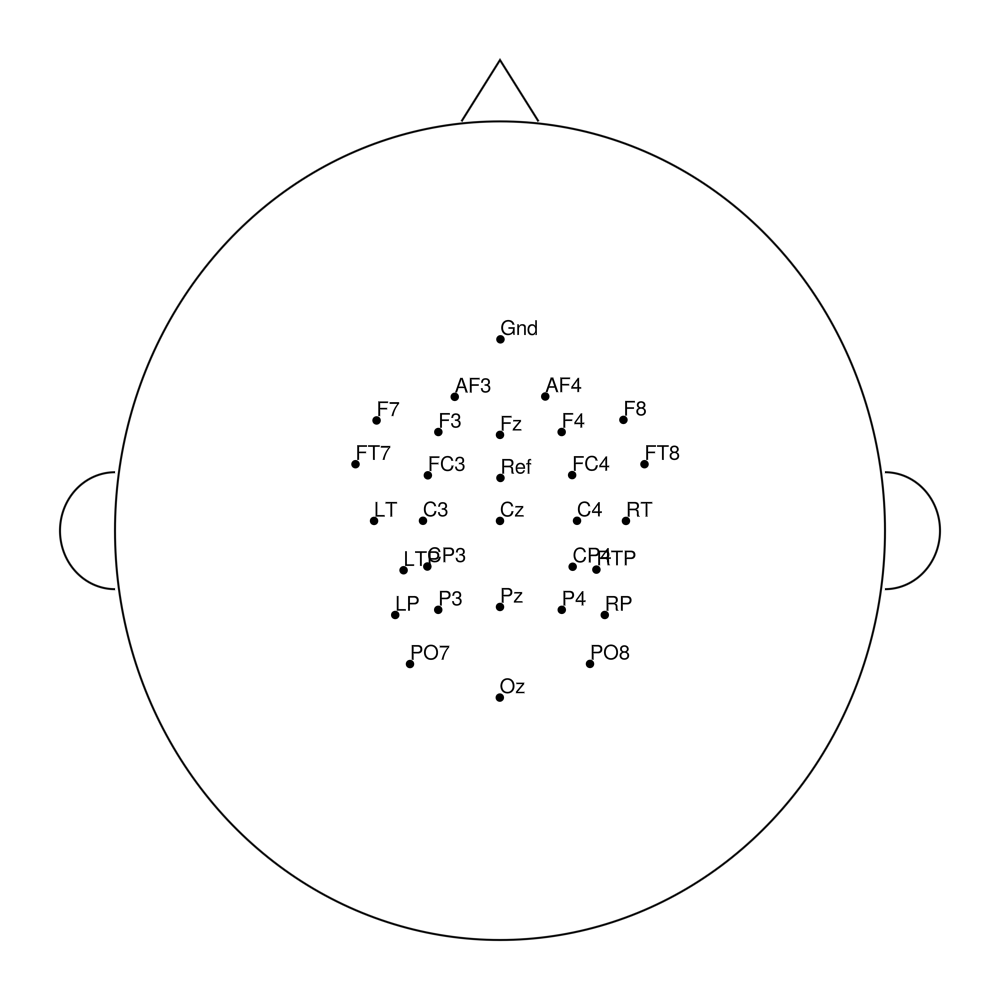
easycapM20
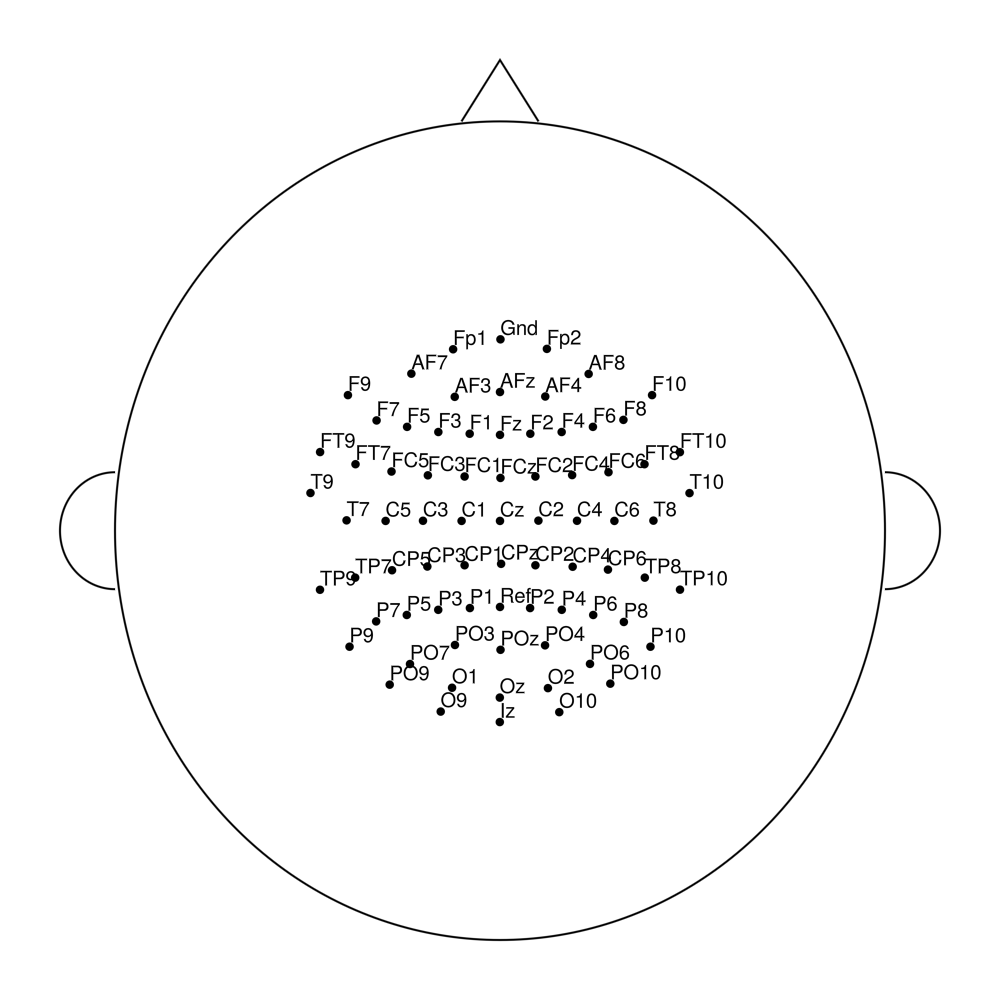
easycapM22
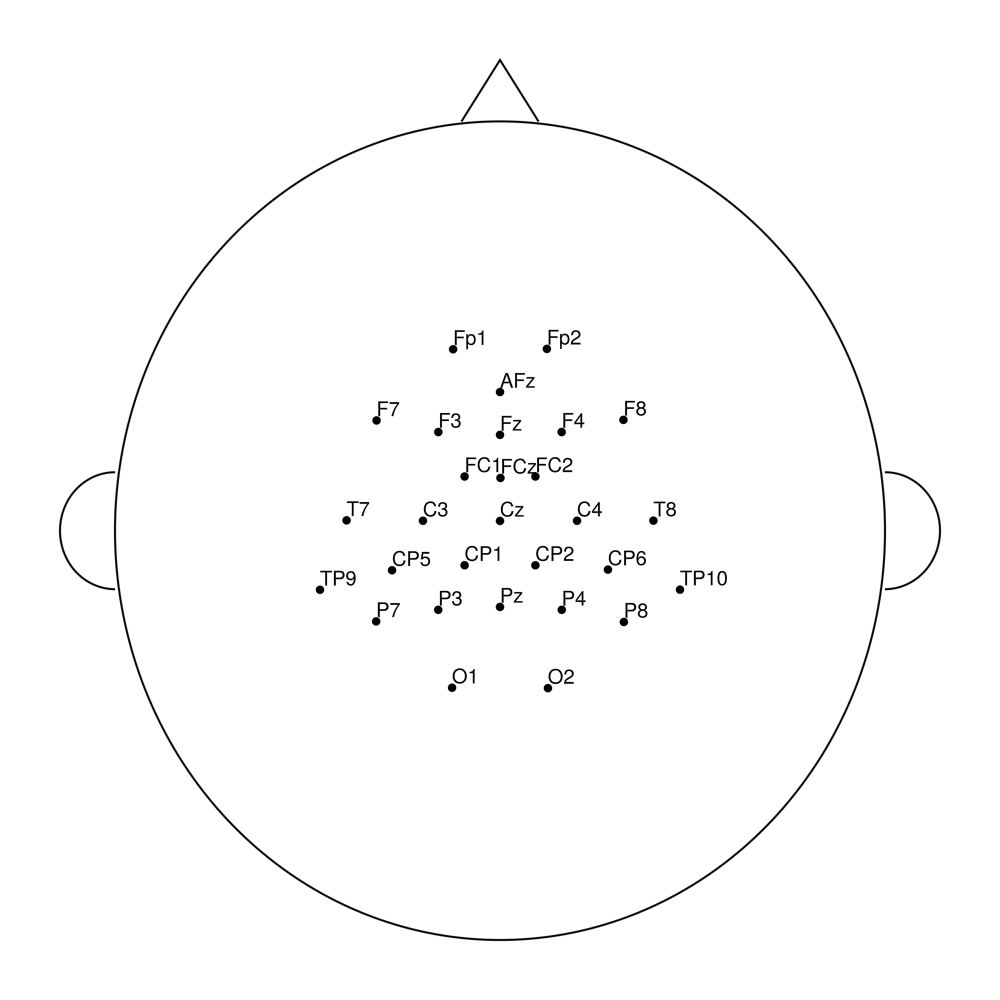
easycapM23
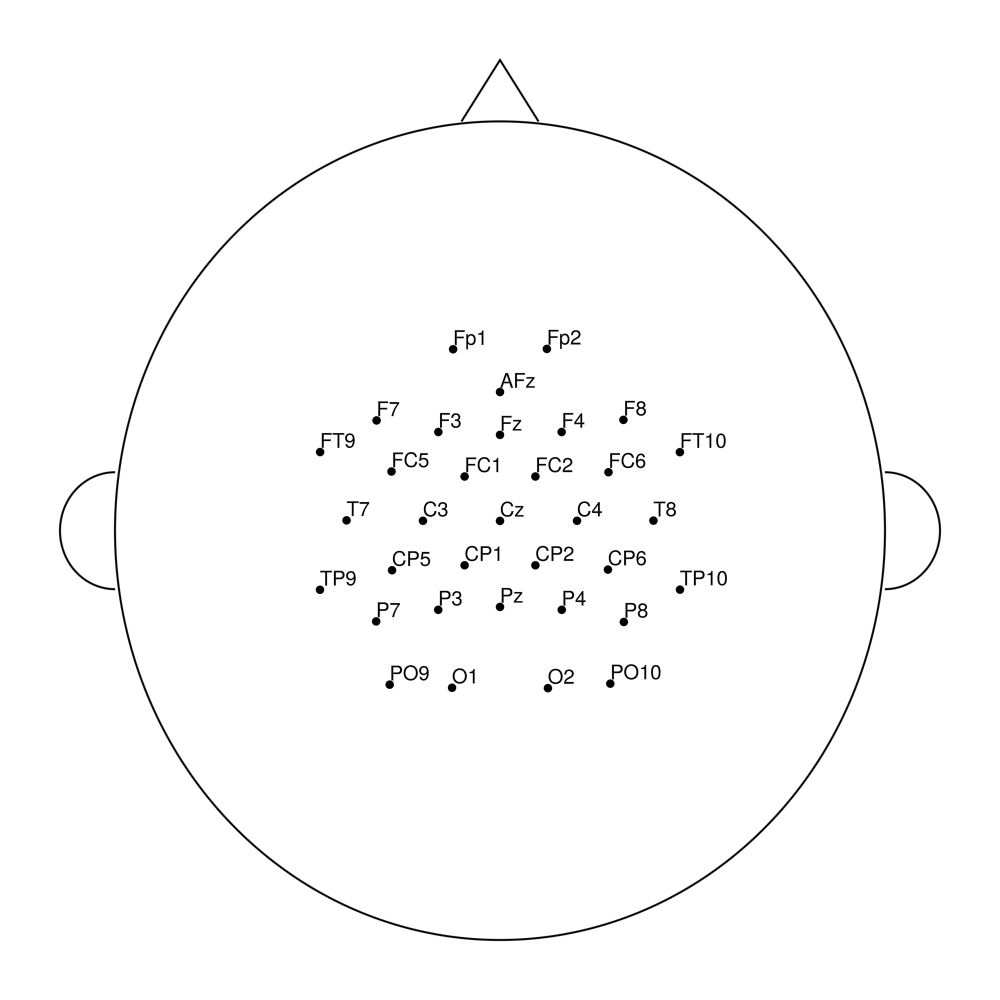
easycapM24
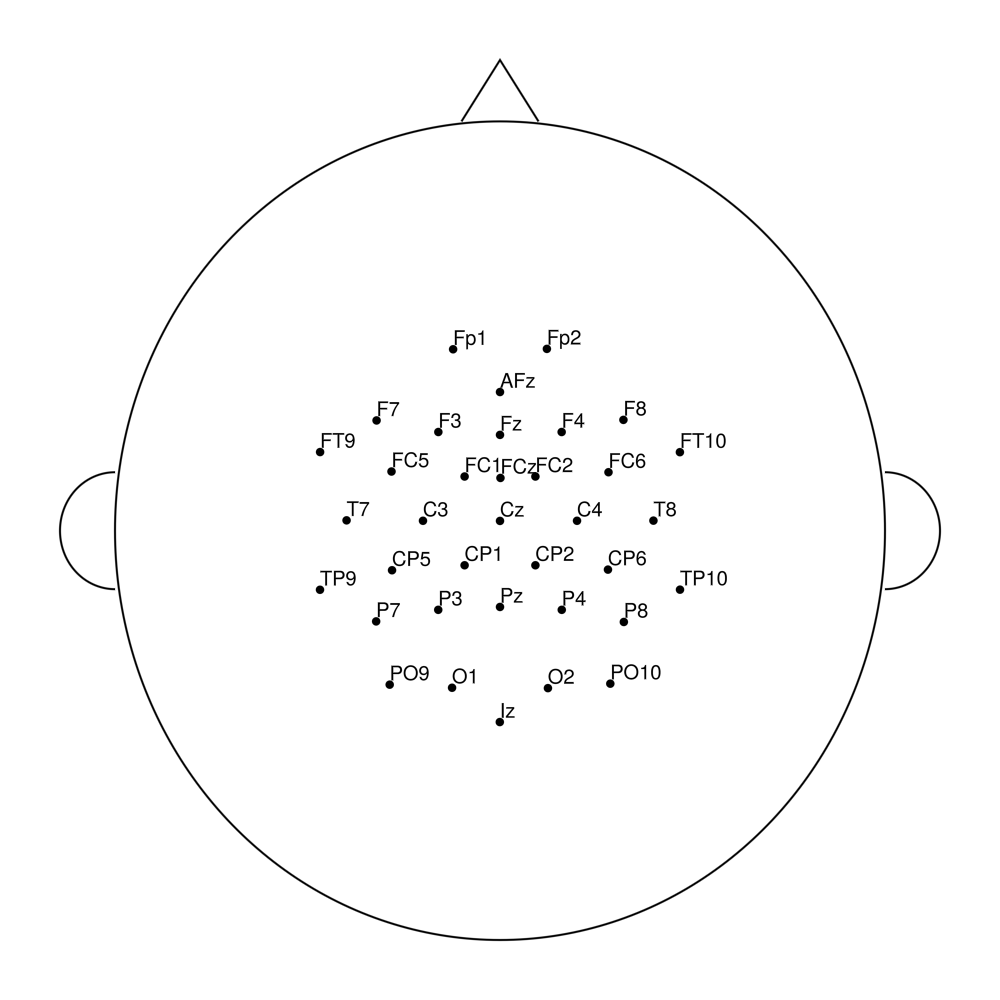
easycapM25
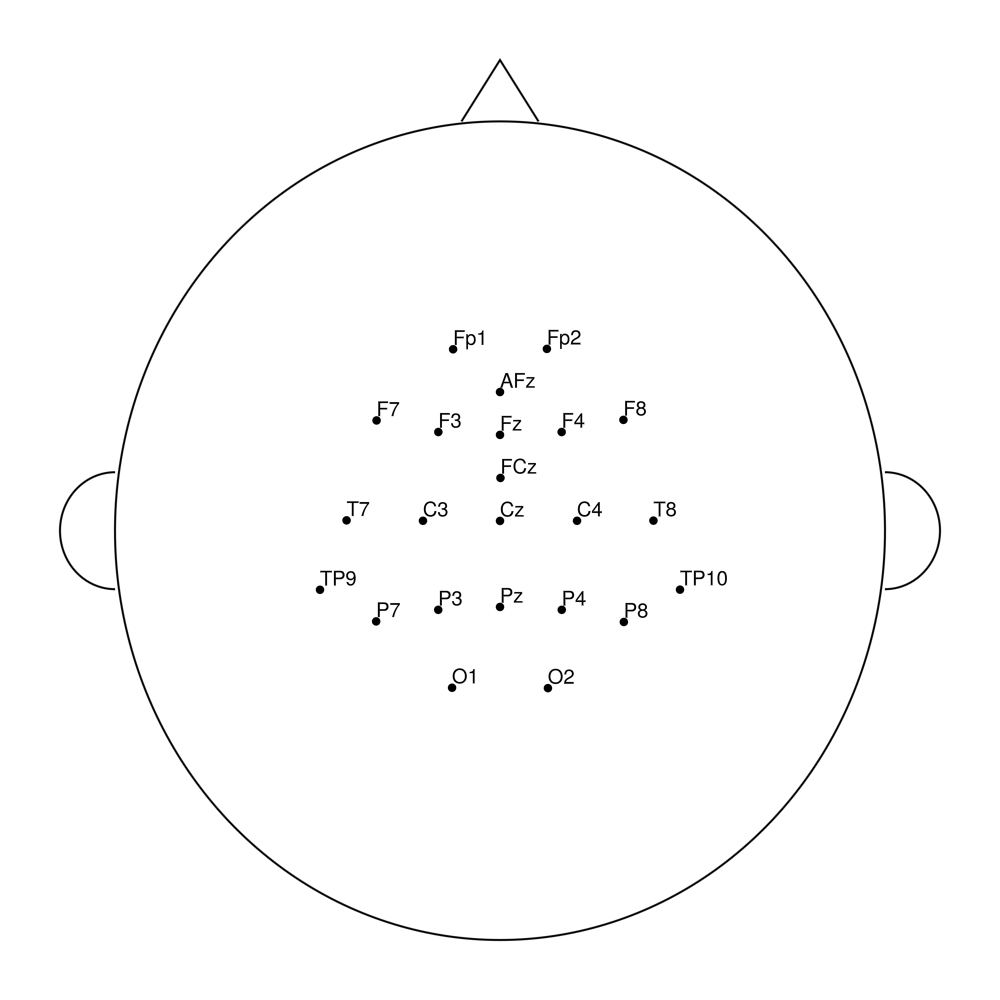
easycapM64
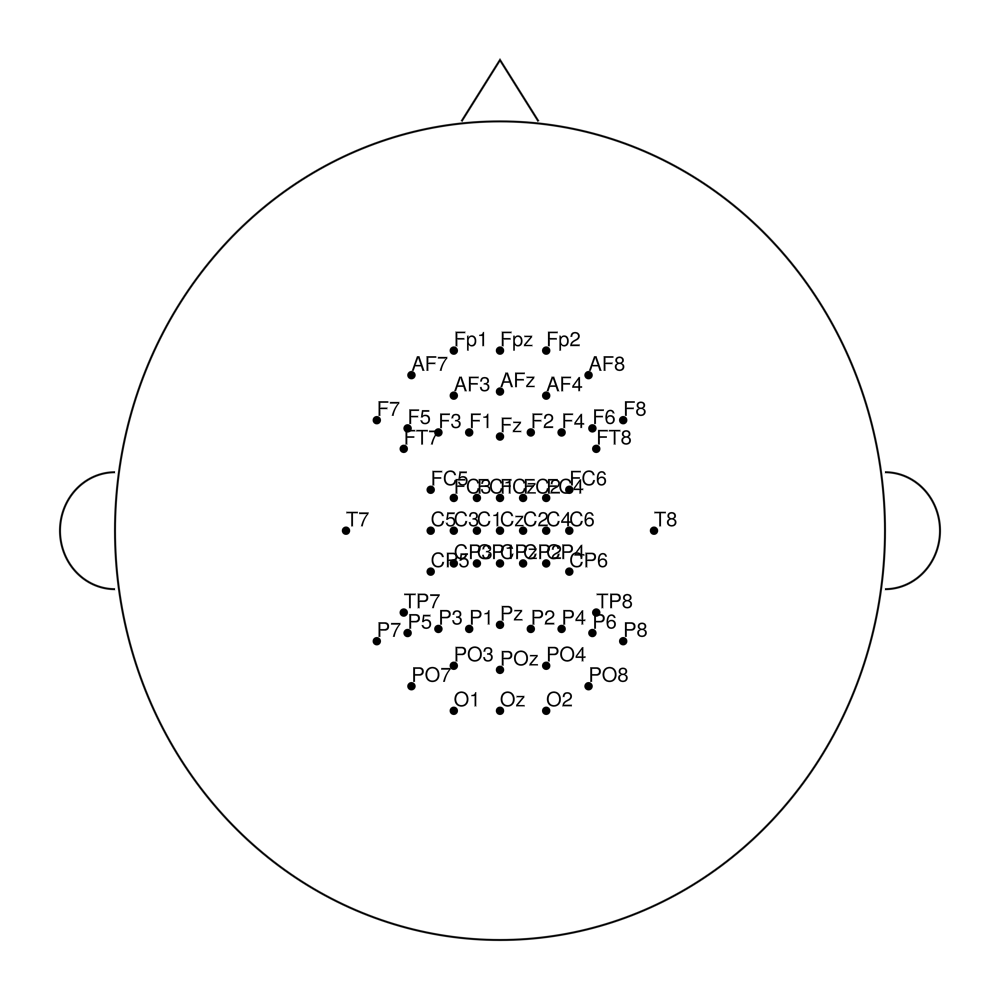
Coordinate Systems
Polar Coordinates (Source of Truth)
Incidence (inc): Vertical angle in degrees from the vertex (Cz = 0°)
0° = vertex (top of head)
90° = equator (ear level)
Values > 90° indicate positions below the equator (e.g., neck electrodes)
Azimuth (azi): Horizontal angle in degrees
0° = right hemisphere
±90° = midline (front/back)
±180° = left hemisphere
Cartesian Coordinates
Cartesian coordinates are computed on-demand from polar coordinates:
2D (x2, y2): Topographic projection for 2D plotting
By default,
preserve_radial_distance=truemaintains anatomical accuracyElectrodes with inc > 90° appear outside the unit circle
3D (x3, y3, z3): Spherical projection for 3D visualization
- Normalized to unit sphere
Related Functions
Layout management and manipulation:
EegFun.read_layout Function
read_layout(file::String)Reads a layout file in CSV format and returns a DataFrame containing the layout information.
Arguments
file::String: The path to the CSV file containing the layout data.
Returns
DataFrame: A DataFrame containing the layout data, with columns for electrode labels, incidence angles, azimuth angles, and calculated Cartesian coordinates (if applicable).
Throws
EegFunError: If the file does not exist or cannot be accessed.
EegFun.validate_layout Function
validate_layout(layout::Layout) -> LayoutValidate that a layout contains all required columns and data.
This function checks that the layout DataFrame contains the minimum required columns for EEG layout operations.
Arguments
layout::Layout: The layout object to validate
Returns
Layout: The validated layout object
Throws
ArgumentError: If required columns are missing
Required Columns
:label: Electrode labels (Symbol type):inc: Incidence angles in degrees:azi: Azimuth angles in degrees
Examples
layout = read_layout("biosemi64.csv")
validated_layout = validate_layout(layout)EegFun.polar_to_cartesian_xy! Function
polar_to_cartesian_xy!(layout::Layout; normalization_radius::Float64=1.0, preserve_radial_distance::Bool=true)Converts polar coordinates (incidence and azimuth angles) from a layout into Cartesian coordinates (x, y).
Arguments
layout::Layout: A Layout containing the layout information with columns for incidence angles (:inc) and azimuth angles (:azi).normalization_radius::Real: Maximum radius for electrode positions (default: 1.0). Only used whenpreserve_radial_distance=false.preserve_radial_distance::Bool: If true, preserves true radial distances from polar coordinates, allowing electrodes with inc>90° to appear outside the unit circle (default: true). When false, normalizes all electrodes to fit within normalization_radius.
Throws
ArgumentError: If required:incand:azicolumns are missing or contain non-numeric values.
Modifies
The input
layoutis modified in place to include new columnsx2andy2, which represent the Cartesian coordinates calculated from the polar coordinates.Clears any existing neighbour information since coordinates have changed.
Returns
- Nothing. The function modifies the
layoutdirectly.
Notes
- When
preserve_radial_distance=true, electrodes imported from EEGLAB with inc>90° (e.g., eye electrodes beyond the scalp) will appear outside the standard head circle, matching EEGLAB's topoplot rendering.
EegFun.polar_to_cartesian_xyz! Function
polar_to_cartesian_xyz!(layout::Layout)Converts polar coordinates (incidence and azimuth angles) from a layout into Cartesian coordinates (x, y, z).
Arguments
layout::Layout: A Layout containing the layout information with columns for incidence angles (:inc) and azimuth angles (:azi).
Throws
ArgumentError: If required:incand:azicolumns are missing or contain non-numeric values.
Modifies
The input
layoutis modified in place to include new columnsx3,y3, andz3, which represent the Cartesian coordinates calculated from the polar coordinates.Clears any existing neighbour information since coordinates have changed.
Returns
- Nothing. The function modifies the
layoutdirectly.
EegFun.get_neighbours_xy! Function
get_neighbours_xy!(layout::Layout, distance_criterion::Real)Get neighbors with automatic computation if needed (mutating version).
This function automatically calculates 2D neighbors if they don't exist or if the distance criterion has changed. It caches the results for efficiency.
Arguments
layout::Layout: The layout objectdistance_criterion::Real: The distance criterion for neighbors in normalized coordinate units (e.g., 0.3–0.5; where 1.0 = the scalp equator at 90° incidence). Think of this as a fraction of the head radius: a value of 0.4 means “neighbours within 40% of the head radius.”
Modifies
layout: Updates neighbor information and criterion
Examples
get_neighbours_xy!(layout, 0.4)EegFun.get_neighbours_xyz! Function
get_neighbours_xyz!(layout::Layout, distance_criterion::Real)Get neighbors with automatic computation if needed (mutating version).
This function automatically calculates 3D neighbors if they don't exist or if the distance criterion has changed. It caches the results for efficiency.
Arguments
layout::Layout: The layout objectdistance_criterion::Real: The distance criterion for neighbors in normalized coordinate units (e.g., 0.3–0.5; where 1.0 = the scalp equator at 90° incidence). Think of this as a fraction of the head radius: a value of 0.4 means “neighbours within 40% of the head radius.”
EegFun.rename_channel! Function
rename_channel!(dat::EegData, rename_dict::Dict{Symbol, Symbol})Rename channels in EEG data using a dictionary mapping old names to new names. Modifies the data in place by updating both the data columns and the layout.
Arguments
dat::EegData: The EEG data object to modifyrename_dict::Dict{Symbol, Symbol}: Dictionary mapping old channel names to new names
Returns
nothing(modifies the data in place)
Examples
# Rename Fp1 to Fpz and Fp2 to Fpz
rename_dict = Dict(:Fp1 => :Fpz, :Fp2 => :Fpz)
rename_channel!(dat, rename_dict)
# Rename a single channel
rename_channel!(dat, Dict(:Cz => :Cz_new))Notes
Only channels that exist in the data will be renamed
If multiple channels would be renamed to the same name, an error is thrown to prevent duplicates
Updates both the data columns and the layout labels
Clears any cached neighbour information in the layout since channel names have changed
Properly handles swaps (e.g., Dict(:A => :B, :B => :A) correctly exchanges the channels)
rename_channel!(layout::Layout, rename_dict::Dict{Symbol, Symbol})Rename channels in a Layout object using a dictionary mapping old names to new names. Modifies the layout in place.
Arguments
layout::Layout: The layout object to modifyrename_dict::Dict{Symbol, Symbol}: Dictionary mapping old channel names to new names
Returns
nothing(modifies the layout in place)
Examples
# Rename Fp1 to Fpz and Fp2 to Fpz
rename_dict = Dict(:Fp1 => :Fpz, :Fp2 => :Fpz)
rename_channel!(layout, rename_dict)
# Rename a single channel
rename_channel!(layout, Dict(:Cz => :Cz_new))Throws
EegFunError: If multiple channels would be renamed to the same name (duplicate prevention)
Notes
Only channels that exist in the layout will be renamed
Clears any cached neighbour information since channel names have changed
Layout visualization:
EegFun.plot_layout_2d Function
plot_layout_2d(layout::Layout; neighbours::Bool=false, correlation_matrix::Union{DataFrame, Nothing}=nothing, display_plot::Bool=true, kwargs...)Create a new figure and plot a 2D EEG electrode layout.
Arguments
layout: Layout containing electrode positionsneighbours: Boolean to show interactive neighbour connections (default: false). Ignored ifcorrelation_matrixis provided.correlation_matrix: Optional DataFrame fromcorrelation_matrix()to show correlation values on hover (default: nothing). When provided,neighboursis ignored.display_plot: Boolean to display the plot (default: true)
Keyword Arguments
All keyword arguments below have sensible defaults. You can override any by passing the corresponding keyword argument.
head_linewidth::Int64=2: Line width of the head shape outline.head_radius::Int64=1: Radius of the head shape (normalized units).head_color::Symbol=black: Color of the head shape outline.
Keyword Arguments
All keyword arguments below have sensible defaults. You can override any by passing the corresponding keyword argument.
point_plot::Bool=true: Whether to plot electrode points.point_markersize::Int64=12: Size of electrode point markers.point_color::Symbol=black: Color of electrode points.point_marker::Symbol=circle: Marker style for electrode points.
Keyword Arguments
All keyword arguments below have sensible defaults. You can override any by passing the corresponding keyword argument.
label_plot::Bool=true: Whether to plot electrode labels.label_fontsize::Int64=20: Font size for electrode labels.label_xoffset::Int64=0: X-axis offset for electrode labels.label_zoffset::Int64=0: Z-axis offset for electrode labels (3D only).label_yoffset::Int64=0: Y-axis offset for electrode labels.label_color::Symbol=black: Color of electrode labels.
Returns
- The figure and axis objects
Example
layout = read_layout("./layouts/biosemi64.csv")
polar_to_cartesian_xy!(layout)
plot_layout_2d(layout)
# With neighbour interactivity
plot_layout_2d(layout, neighbours=true)
# With correlation matrix
cm = correlation_matrix(dat)
plot_layout_2d(layout, correlation_matrix=cm)EegFun.plot_layout_3d Function
plot_layout_3d(layout::Layout; neighbours::Bool=false, display_plot::Bool=true, kwargs...)Create a new figure and plot a 3D EEG electrode layout.
Arguments
layout: Layout containing electrode positionsneighbours: Boolean to show interactive neighbour connections (default: false)display_plot: Boolean to display the plot (default: true)
Keyword Arguments
All keyword arguments below have sensible defaults. You can override any by passing the corresponding keyword argument.
head_linewidth::Int64=2: Line width of the head shape outline.head_radius::Int64=1: Radius of the head shape (normalized units).head_color::Symbol=black: Color of the head shape outline.
Keyword Arguments
All keyword arguments below have sensible defaults. You can override any by passing the corresponding keyword argument.
point_plot::Bool=true: Whether to plot electrode points.point_markersize::Int64=12: Size of electrode point markers.point_color::Symbol=black: Color of electrode points.point_marker::Symbol=circle: Marker style for electrode points.
Keyword Arguments
All keyword arguments below have sensible defaults. You can override any by passing the corresponding keyword argument.
label_plot::Bool=true: Whether to plot electrode labels.label_fontsize::Int64=20: Font size for electrode labels.label_xoffset::Int64=0: X-axis offset for electrode labels.label_zoffset::Int64=0: Z-axis offset for electrode labels (3D only).label_yoffset::Int64=0: Y-axis offset for electrode labels.label_color::Symbol=black: Color of electrode labels.
Returns
- The figure and axis objects
Example
layout = read_layout("./layouts/biosemi64.csv")
polar_to_cartesian_xyz!(layout)
plot_layout_3d(layout)
# With neighbour interactivity
plot_layout_3d(layout, neighbours=true)